Nanda Horeweg
A protocol for evaluating robustness to H&E staining variation in computational pathology models
Mar 13, 2026Abstract:Sensitivity to staining variation remains a major barrier to deploying computational pathology (CPath) models as hematoxylin and eosin (H&E) staining varies across laboratories, requiring systematic assessment of how this variability affects model prediction. In this work, we developed a three-step protocol for evaluating robustness to H&E staining variation in CPath models. Step 1: Select reference staining conditions, Step 2: Characterize test set staining properties, Step 3: Apply CPath model(s) under simulated reference staining conditions. Here, we first created a new reference staining library based on the PLISM dataset. As an exemplary use case, we applied the protocol to assess the robustness properties of 306 microsatellite instability (MSI) classification models on the unseen SurGen colorectal cancer dataset (n=738), including 300 attention-based multiple instance learning models trained on the TCGA-COAD/READ datasets across three feature extractors (UNI2-h, H-Optimus-1, Virchow2), alongside six public MSI classification models. Classification performance was measured as AUC, and robustness as the min-max AUC range across four simulated staining conditions (low/high H&E intensity, low/high H&E color similarity). Across models and staining conditions, classification performance ranged from AUC 0.769-0.911 ($Δ$ = 0.142). Robustness ranged from 0.007-0.079 ($Δ$ = 0.072), and showed a weak inverse correlation with classification performance (Pearson r=-0.22, 95% CI [-0.34, -0.11]). Thus, we show that the proposed evaluation protocol enables robustness-informed CPath model selection and provides insight into performance shifts across H&E staining conditions, supporting the identification of operational ranges for reliable model deployment. Code is available at https://github.com/CTPLab/staining-robustness-evaluation .
Automated causal inference in application to randomized controlled clinical trials
Jan 20, 2022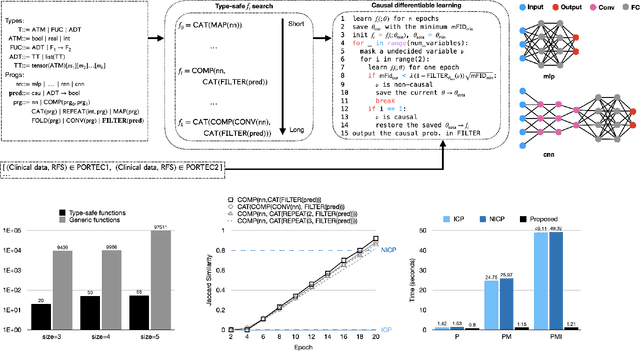
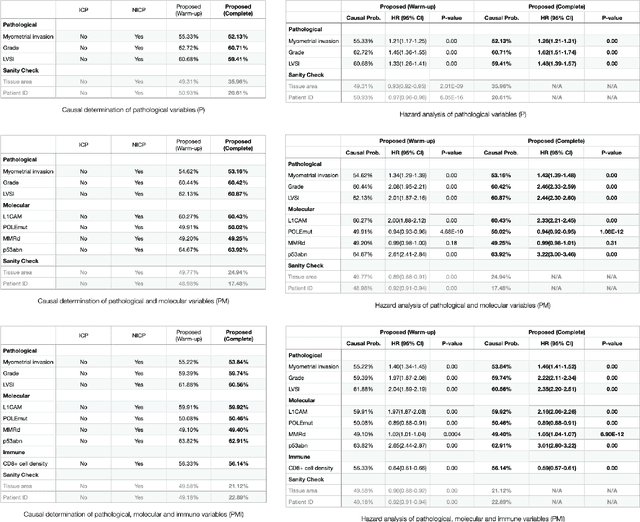
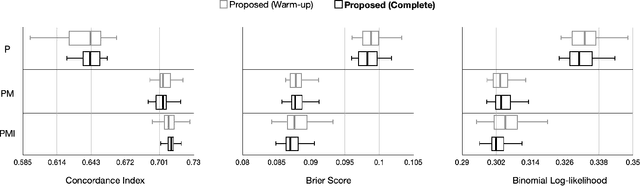
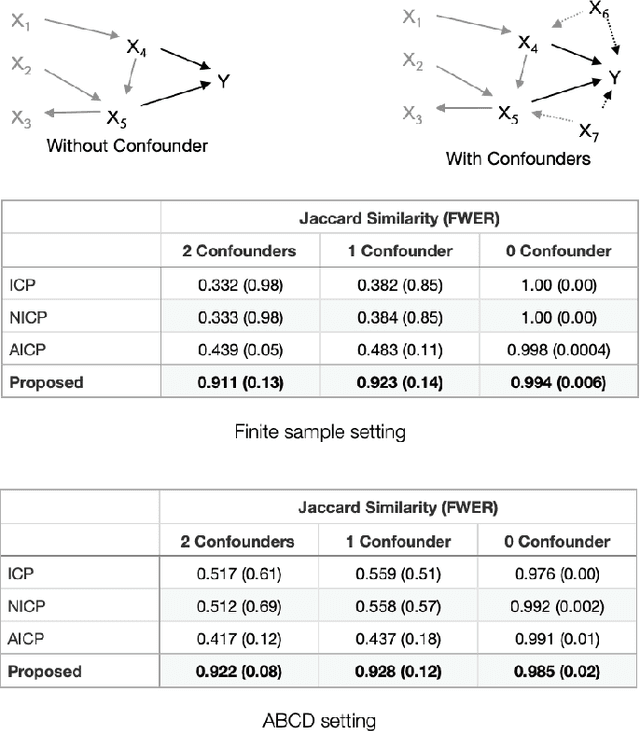
Abstract:Randomized controlled trials (RCTs) are considered as the gold standard for testing causal hypotheses in the clinical domain. However, the investigation of prognostic variables of patient outcome in a hypothesized cause-effect route is not feasible using standard statistical methods. Here, we propose a new automated causal inference method (AutoCI) built upon the invariant causal prediction (ICP) framework for the causal re-interpretation of clinical trial data. Compared to existing methods, we show that the proposed AutoCI allows to efficiently determine the causal variables with a clear differentiation on two real-world RCTs of endometrial cancer patients with mature outcome and extensive clinicopathological and molecular data. This is achieved via suppressing the causal probability of non-causal variables by a wide margin. In ablation studies, we further demonstrate that the assignment of causal probabilities by AutoCI remain consistent in the presence of confounders. In conclusion, these results confirm the robustness and feasibility of AutoCI for future applications in real-world clinical analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge