Nadjia Kachenoura
Progressive Learning with Anatomical Priors for Reliable Left Atrial Scar Segmentation from Late Gadolinium Enhancement MRI
Mar 27, 2026Abstract:Cardiac MRI late gadolinium enhancement (LGE) enables non-invasive identification of left atrial (LA) scar, whose spatial distribution is strongly associated with atrial fibrillation (AF) severity and recurrence. However, automatic LA scar segmentation remains challenging due to low contrast, annotation variability, and the lack of anatomical constraints, often leading to non-reliable predictions. Accordingly, our aim was to propose a progressive learning strategy to segment LA scar from LGE images inspired from a clinical workflow. A 3-stage framework based on SwinUNETR was implemented, comprising: 1) a first LA cavity pre-learning model, 2) dual-task model which further learns spatial relationship between LA geometry and scar patterns, and 3) fine-tuning on precise segmentation of the scar. Furthermore, we introduced an anatomy-aware spatially weighted loss that incorporates prior clinical knowledge by constraining scar predictions to anatomically plausible LA wall regions while mitigating annotation bias. Our preliminary results obtained on validation LGE volumes from LASCARQS public dataset after 5-fold cross validation, LA segmentation had Dice score of 0.94, LA scar segmentation achieved Dice score of 0.50, Hausdorff Distance of 11.84 mm, Average Surface Distance of 1.80 mm, outperforming only a one-stage scar segmentation with 0.49, 13.02 mm, 1.96 mm, repectively. By explicitly embedding clinical anatomical priors and diagnostic reasoning into deep learning, the proposed approach improved the accuracy and reliability of LA scar segmentation from LGE, revealing the importance of clinically informed model design.
SFB-net for cardiac segmentation: Bridging the semantic gap with attention
Oct 24, 2024



Abstract:In the past few years, deep learning algorithms have been widely used for cardiac image segmentation. However, most of these architectures rely on convolutions that hardly model long-range dependencies, limiting their ability to extract contextual information. In order to tackle this issue, this article introduces the Swin Filtering Block network (SFB-net) which takes advantage of both conventional and swin transformer layers. The former are used to introduce spatial attention at the bottom of the network, while the latter are applied to focus on high level semantically rich features between the encoder and decoder. An average Dice score of 92.4 was achieved on the ACDC dataset. To the best of our knowledge, this result outperforms any other work on this dataset. The average Dice score of 87.99 obtained on the M\&M's dataset demonstrates that the proposed method generalizes well to data from different vendors and centres.
Deep Learning-based Automated Aortic Area and Distensibility Assessment: The Multi-Ethnic Study of Atherosclerosis
Mar 03, 2021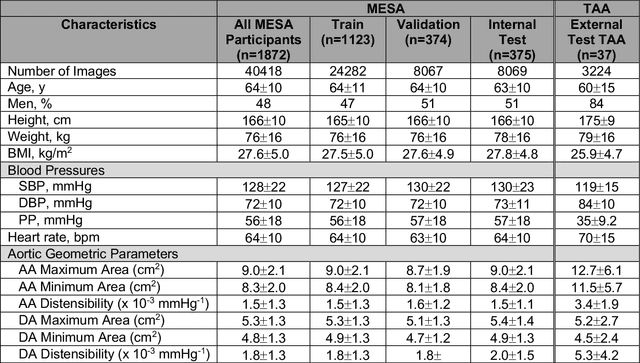
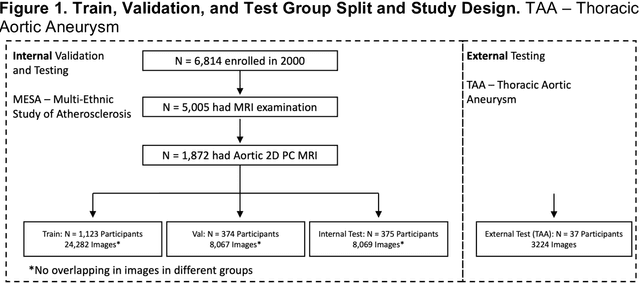

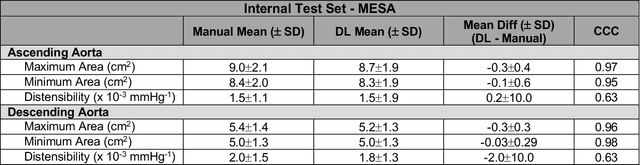
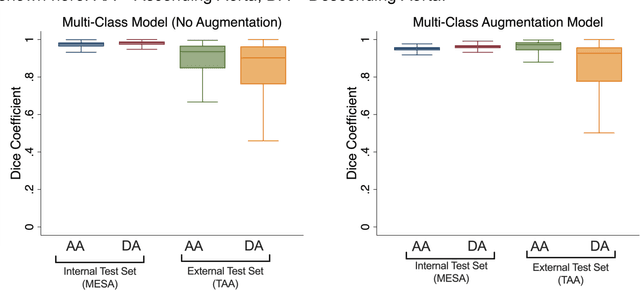
Abstract:This study applies convolutional neural network (CNN)-based automatic segmentation and distensibility measurement of the ascending and descending aorta from 2D phase-contrast cine magnetic resonance imaging (PC-cine MRI) within the large MESA cohort with subsequent assessment on an external cohort of thoracic aortic aneurysm (TAA) patients. 2D PC-cine MRI images of the ascending and descending aorta at the pulmonary artery bifurcation from the MESA study were included. Train, validation, and internal test sets consisted of 1123 studies (24282 images), 374 studies (8067 images), and 375 studies (8069 images), respectively. An external test set of TAAs consisted of 37 studies (3224 images). A U-Net based CNN was constructed, and performance was evaluated utilizing dice coefficient (for segmentation) and concordance correlation coefficients (CCC) of aortic geometric parameters by comparing to manual segmentation and parameter estimation. Dice coefficients for aorta segmentation were 97.6% (CI: 97.5%-97.6%) and 93.6% (84.6%-96.7%) on the internal and external test of TAAs, respectively. CCC for comparison of manual and CNN maximum and minimum ascending aortic areas were 0.97 and 0.95, respectively, on the internal test set and 0.997 and 0.995, respectively, for the external test. CCCs for maximum and minimum descending aortic areas were 0.96 and 0. 98, respectively, on the internal test set and 0.93 and 0.93, respectively, on the external test set. We successfully developed and validated a U-Net based ascending and descending aortic segmentation and distensibility quantification model in a large multi-ethnic database and in an external cohort of TAA patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge