Myeonghun Lee
AmorProt: Amino Acid Molecular Fingerprints Repurposing based Protein Fingerprint
Mar 27, 2023Abstract:As protein therapeutics play an important role in almost all medical fields, numerous studies have been conducted on proteins using artificial intelligence. Artificial intelligence has enabled data driven predictions without the need for expensive experiments. Nevertheless, unlike the various molecular fingerprint algorithms that have been developed, protein fingerprint algorithms have rarely been studied. In this study, we proposed the amino acid molecular fingerprints repurposing based protein (AmorProt) fingerprint, a protein sequence representation method that effectively uses the molecular fingerprints corresponding to 20 amino acids. Subsequently, the performances of the tree based machine learning and artificial neural network models were compared using (1) amyloid classification and (2) isoelectric point regression. Finally, the applicability and advantages of the developed platform were demonstrated through a case study and the following experiments: (3) comparison of dataset dependence with feature based methods; (4) feature importance analysis; and (5) protein space analysis. Consequently, the significantly improved model performance and data set independent versatility of the AmorProt fingerprint were verified. The results revealed that the current protein representation method can be applied to various fields related to proteins, such as predicting their fundamental properties or interaction with ligands.
MGCVAE: Multi-objective Inverse Design via Molecular Graph Conditional Variational Autoencoder
Feb 14, 2022
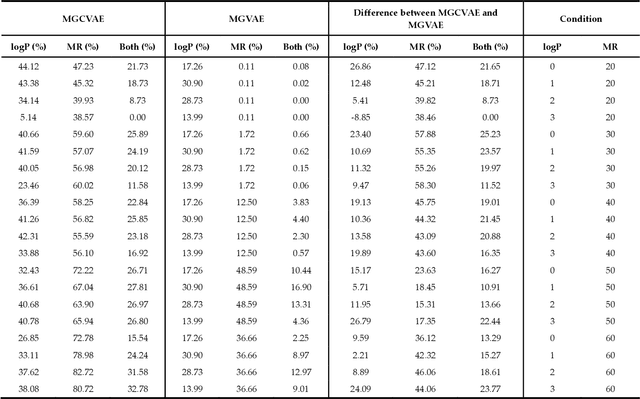
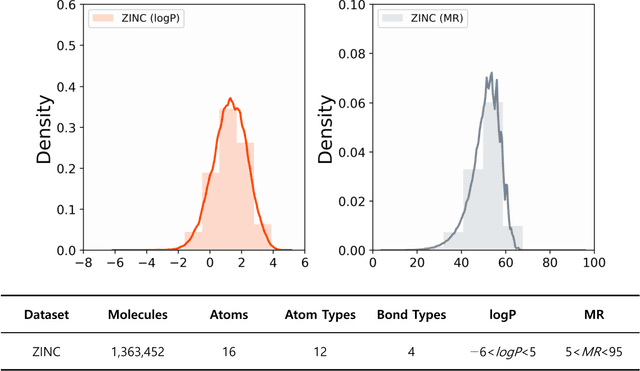

Abstract:The ultimate goal of various fields is to directly generate molecules with desired properties, such as finding water-soluble molecules in drug development and finding molecules suitable for organic light-emitting diode (OLED) or photosensitizers in the field of development of new organic materials. In this respect, this study proposes a molecular graph generative model based on the autoencoder for de novo design. The performance of molecular graph conditional variational autoencoder (MGCVAE) for generating molecules having specific desired properties is investigated by comparing it to molecular graph variational autoencoder (MGVAE). Furthermore, multi-objective optimization for MGCVAE was applied to satisfy two selected properties simultaneously. In this study, two physical properties -- logP and molar refractivity -- were used as optimization targets for the purpose of designing de novo molecules, especially in drug discovery. As a result, it was confirmed that among generated molecules, 25.89% optimized molecules were generated in MGCVAE compared to 0.66% in MGVAE. Hence, it demonstrates that MGCVAE effectively produced drug-like molecules with two target properties. The results of this study suggest that these graph-based data-driven models are one of the effective methods of designing new molecules that fulfill various physical properties, such as drug discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge