Muhammad Muneeb
G2DR: A Genotype-First Framework for Genetics-Informed Target Prioritization and Drug Repurposing
Mar 20, 2026Abstract:Human genetics offers a promising route to therapeutic discovery, yet practical frameworks translating genotype-derived signal into ranked target and drug hypotheses remain limited, particularly when matched disease transcriptomics are unavailable. Here we present G2DR, a genotype-first prioritization framework propagating inherited variation through genetically predicted expression, multi-method gene-level testing, pathway enrichment, network context, druggability, and multi-source drug--target evidence integration. In a migraine case study with 733 UK Biobank participants under stratified five-fold cross-validation, we imputed expression across seven transcriptome-weight resources and ranked genes using a reproducibility-aware discovery score from training and validation data, followed by a balanced integrated score for target selection. Discovery-based prioritization generalized to held-out data, achieving gene-level ROC-AUC of 0.775 and PR-AUC of 0.475, while retaining enrichment for curated migraine biology. Mapping prioritized genes to compounds via Open Targets, DGIdb, and ChEMBL yielded drug sets enriched for migraine-linked compounds relative to a global background, though recovery favoured broader mechanism-linked and off-label space over migraine-specific approved therapies. Directionality filtering separated broadly recovered compounds from mechanistically compatible candidates. G2DR is a modular framework for genetics-informed hypothesis generation, not a clinically actionable recommendation system. All outputs require independent experimental, pharmacological, and clinical validation.
Deep learning pipeline for image classification on mobile phones
May 31, 2022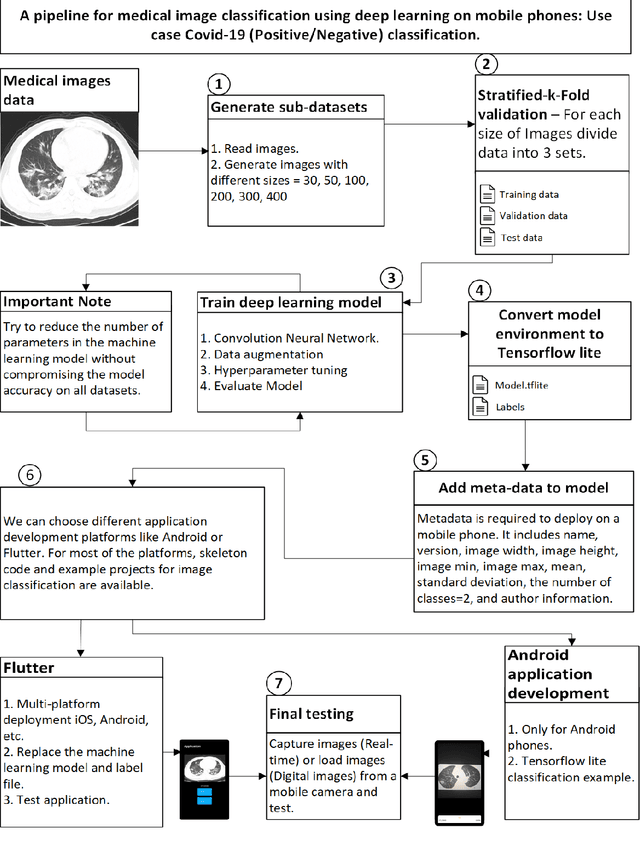
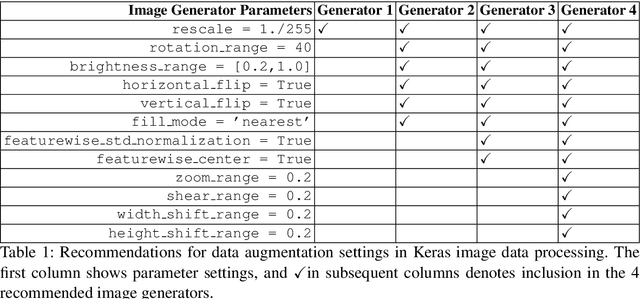
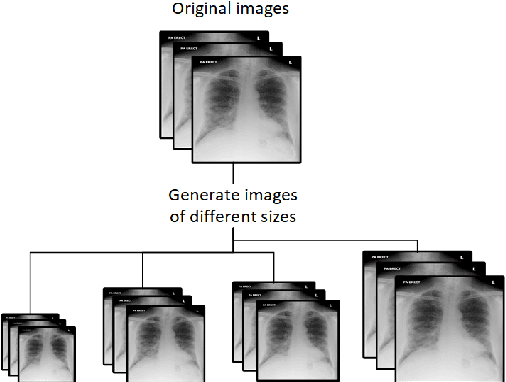
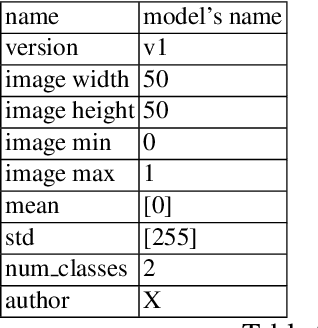
Abstract:This article proposes and documents a machine-learning framework and tutorial for classifying images using mobile phones. Compared to computers, the performance of deep learning model performance degrades when deployed on a mobile phone and requires a systematic approach to find a model that performs optimally on both computers and mobile phones. By following the proposed pipeline, which consists of various computational tools, simple procedural recipes, and technical considerations, one can bring the power of deep learning medical image classification to mobile devices, potentially unlocking new domains of applications. The pipeline is demonstrated on four different publicly available datasets: COVID X-rays, COVID CT scans, leaves, and colorectal cancer. We used two application development frameworks: TensorFlow Lite (real-time testing) and Flutter (digital image testing) to test the proposed pipeline. We found that transferring deep learning models to a mobile phone is limited by hardware and classification accuracy drops. To address this issue, we proposed this pipeline to find an optimized model for mobile phones. Finally, we discuss additional applications and computational concerns related to deploying deep-learning models on phones, including real-time analysis and image preprocessing. We believe the associated documentation and code can help physicians and medical experts develop medical image classification applications for distribution.
* 20 pages
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge