Mostafa Rahimi Azghadi
A lightweight Transformer-based model for fish landmark detection
Sep 13, 2022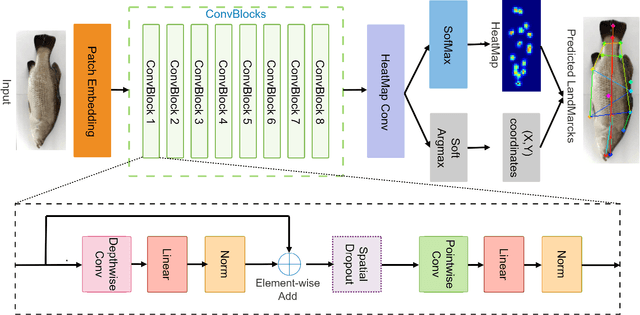
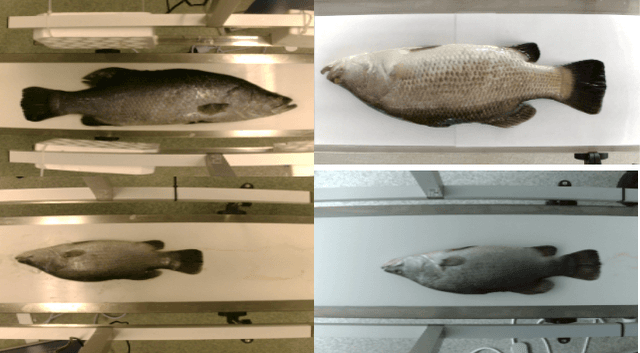
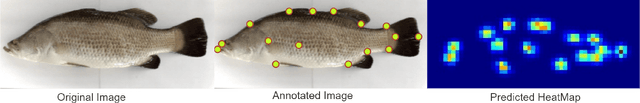
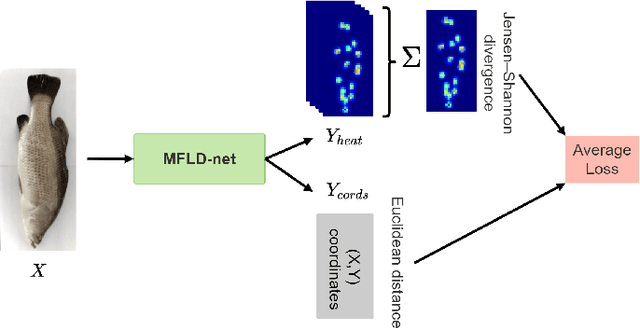


Abstract:Transformer-based models, such as the Vision Transformer (ViT), can outperform onvolutional Neural Networks (CNNs) in some vision tasks when there is sufficient training data. However, (CNNs) have a strong and useful inductive bias for vision tasks (i.e. translation equivariance and locality). In this work, we developed a novel model architecture that we call a Mobile fish landmark detection network (MFLD-net). We have made this model using convolution operations based on ViT (i.e. Patch embeddings, Multi-Layer Perceptrons). MFLD-net can achieve competitive or better results in low data regimes while being lightweight and therefore suitable for embedded and mobile devices. Furthermore, we show that MFLD-net can achieve keypoint (landmark) estimation accuracies on-par or even better than some of the state-of-the-art (CNNs) on a fish image dataset. Additionally, unlike ViT, MFLD-net does not need a pre-trained model and can generalise well when trained on a small dataset. We provide quantitative and qualitative results that demonstrate the model's generalisation capabilities. This work will provide a foundation for future efforts in developing mobile, but efficient fish monitoring systems and devices.
Unsupervised Fish Trajectory Tracking and Segmentation
Aug 23, 2022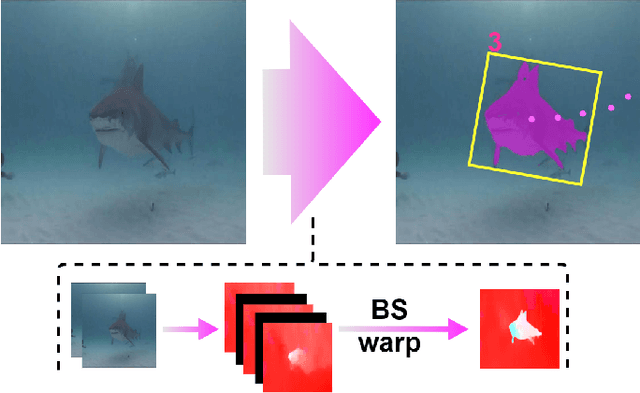
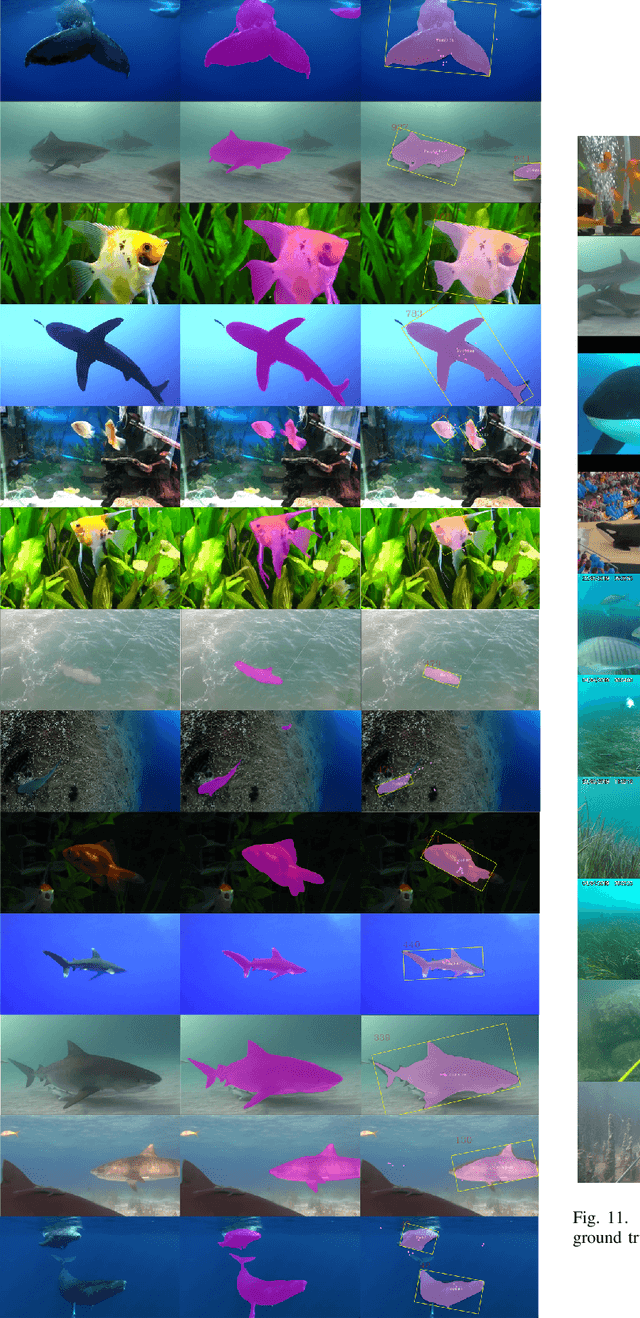
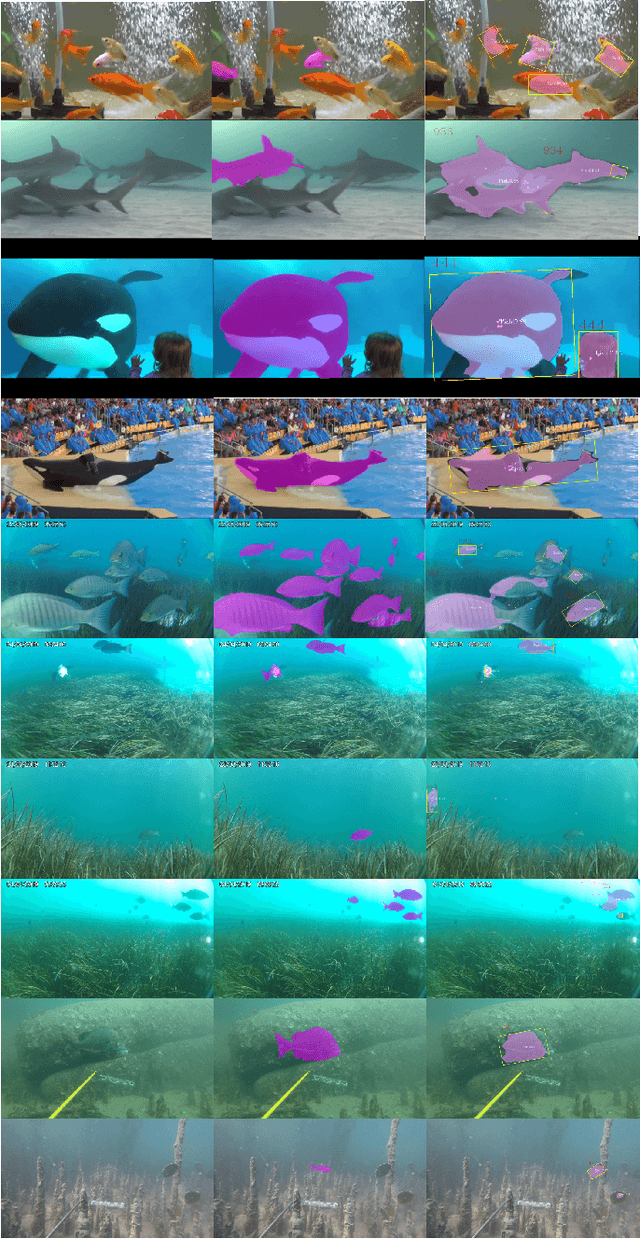


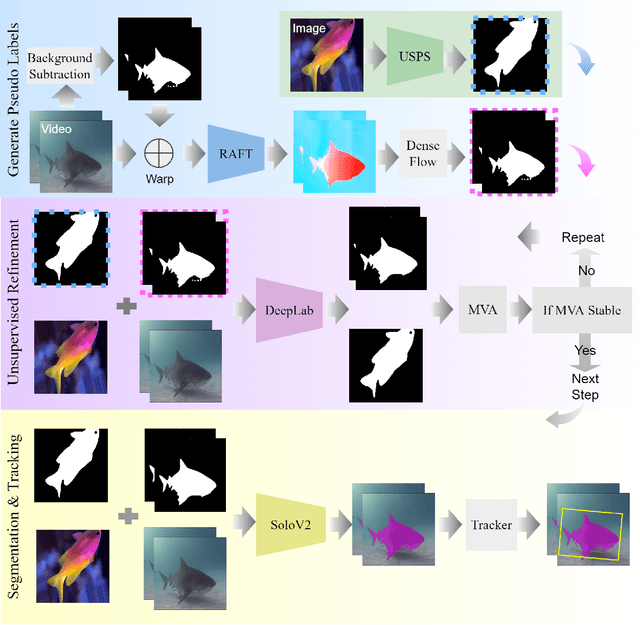
Abstract:DNN for fish tracking and segmentation based on high-quality labels is expensive. Alternative unsupervised approaches rely on spatial and temporal variations that naturally occur in video data to generate noisy pseudo-ground-truth labels. These pseudo-labels are used to train a multi-task deep neural network. In this paper, we propose a three-stage framework for robust fish tracking and segmentation, where the first stage is an optical flow model, which generates the pseudo labels using spatial and temporal consistency between frames. In the second stage, a self-supervised model refines the pseudo-labels incrementally. In the third stage, the refined labels are used to train a segmentation network. No human annotations are used during the training or inference. Extensive experiments are performed to validate our method on three public underwater video datasets and to demonstrate that it is highly effective for video annotation and segmentation. We also evaluate the robustness of our framework to different imaging conditions and discuss the limitations of our current implementation.
Seizure Detection and Prediction by Parallel Memristive Convolutional Neural Networks
Jun 20, 2022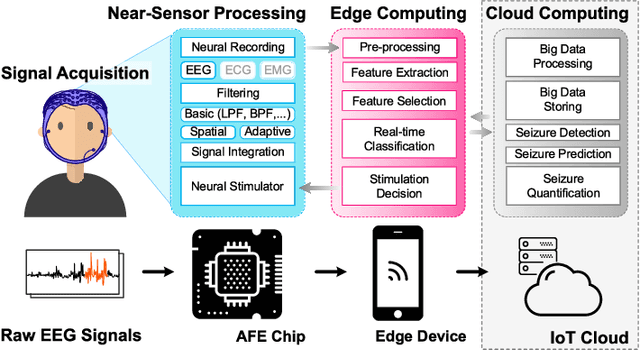
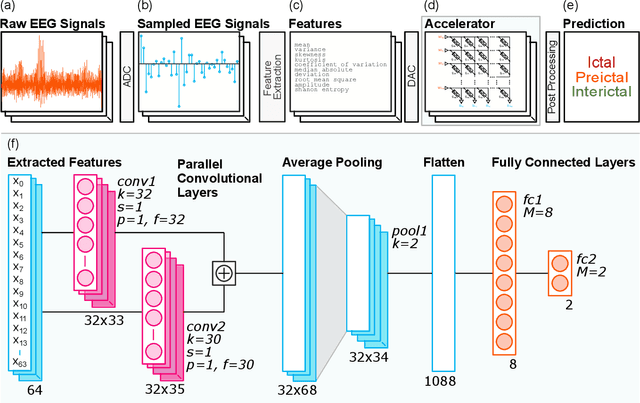
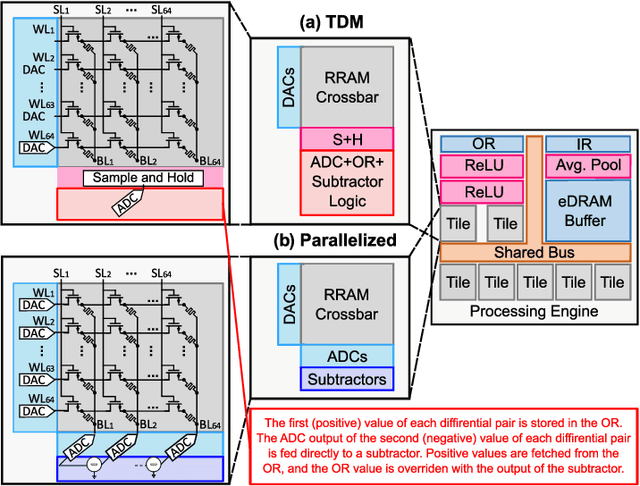
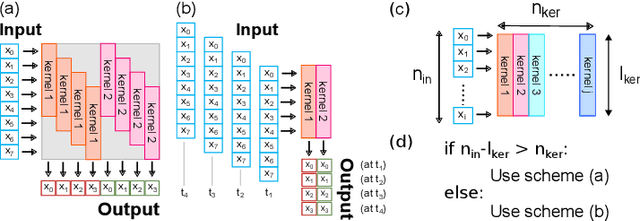
Abstract:During the past two decades, epileptic seizure detection and prediction algorithms have evolved rapidly. However, despite significant performance improvements, their hardware implementation using conventional technologies, such as Complementary Metal-Oxide-Semiconductor (CMOS), in power and area-constrained settings remains a challenging task; especially when many recording channels are used. In this paper, we propose a novel low-latency parallel Convolutional Neural Network (CNN) architecture that has between 2-2,800x fewer network parameters compared to SOTA CNN architectures and achieves 5-fold cross validation accuracy of 99.84% for epileptic seizure detection, and 99.01% and 97.54% for epileptic seizure prediction, when evaluated using the University of Bonn Electroencephalogram (EEG), CHB-MIT and SWEC-ETHZ seizure datasets, respectively. We subsequently implement our network onto analog crossbar arrays comprising Resistive Random-Access Memory (RRAM) devices, and provide a comprehensive benchmark by simulating, laying out, and determining hardware requirements of the CNN component of our system. To the best of our knowledge, we are the first to parallelize the execution of convolution layer kernels on separate analog crossbars to enable 2 orders of magnitude reduction in latency compared to SOTA hybrid Memristive-CMOS DL accelerators. Furthermore, we investigate the effects of non-idealities on our system and investigate Quantization Aware Training (QAT) to mitigate the performance degradation due to low ADC/DAC resolution. Finally, we propose a stuck weight offsetting methodology to mitigate performance degradation due to stuck RON/ROFF memristor weights, recovering up to 32% accuracy, without requiring retraining. The CNN component of our platform is estimated to consume approximately 2.791W of power while occupying an area of 31.255mm$^2$ in a 22nm FDSOI CMOS process.
* Accepted by IEEE Transactions on Biomedical Circuits and Systems
Applications of Deep Learning in Fish Habitat Monitoring: A Tutorial and Survey
Jun 11, 2022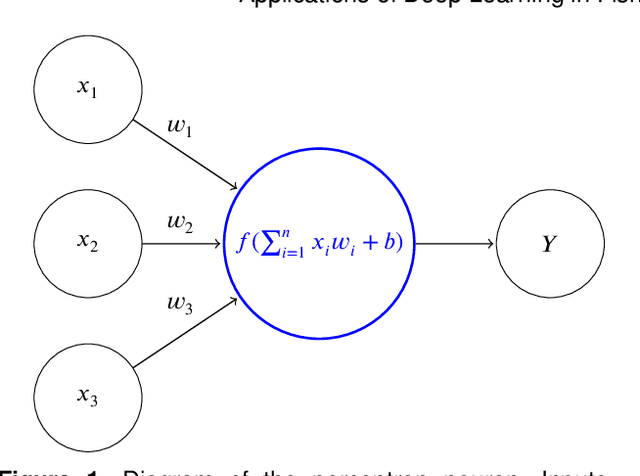
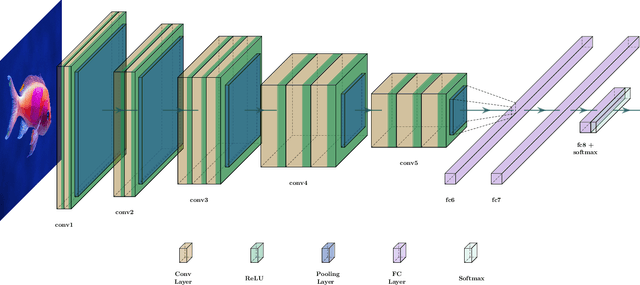

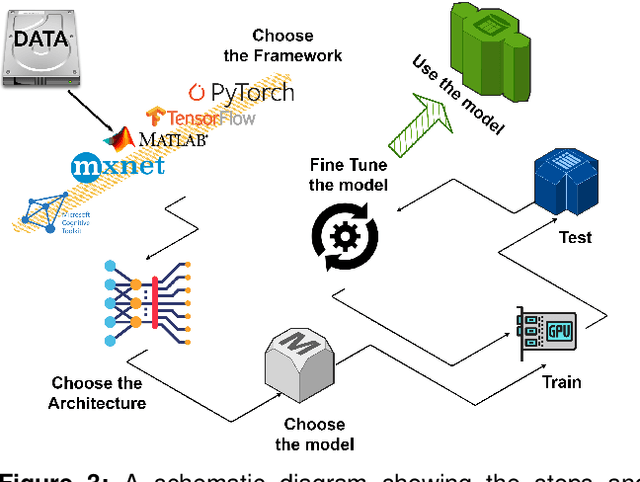
Abstract:Marine ecosystems and their fish habitats are becoming increasingly important due to their integral role in providing a valuable food source and conservation outcomes. Due to their remote and difficult to access nature, marine environments and fish habitats are often monitored using underwater cameras. These cameras generate a massive volume of digital data, which cannot be efficiently analysed by current manual processing methods, which involve a human observer. DL is a cutting-edge AI technology that has demonstrated unprecedented performance in analysing visual data. Despite its application to a myriad of domains, its use in underwater fish habitat monitoring remains under explored. In this paper, we provide a tutorial that covers the key concepts of DL, which help the reader grasp a high-level understanding of how DL works. The tutorial also explains a step-by-step procedure on how DL algorithms should be developed for challenging applications such as underwater fish monitoring. In addition, we provide a comprehensive survey of key deep learning techniques for fish habitat monitoring including classification, counting, localization, and segmentation. Furthermore, we survey publicly available underwater fish datasets, and compare various DL techniques in the underwater fish monitoring domains. We also discuss some challenges and opportunities in the emerging field of deep learning for fish habitat processing. This paper is written to serve as a tutorial for marine scientists who would like to grasp a high-level understanding of DL, develop it for their applications by following our step-by-step tutorial, and see how it is evolving to facilitate their research efforts. At the same time, it is suitable for computer scientists who would like to survey state-of-the-art DL-based methodologies for fish habitat monitoring.
Transformer-based Self-Supervised Fish Segmentation in Underwater Videos
Jun 11, 2022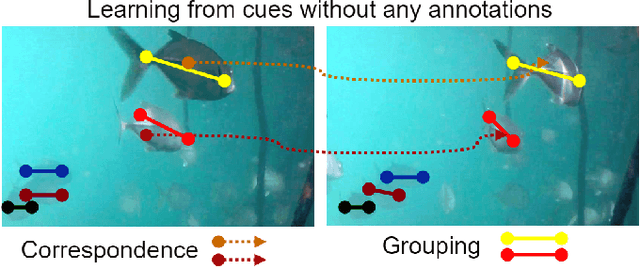
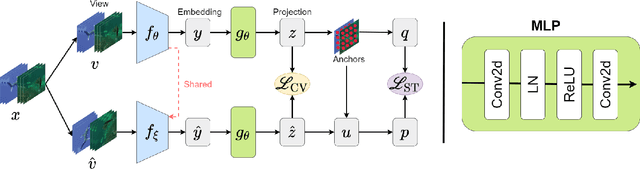
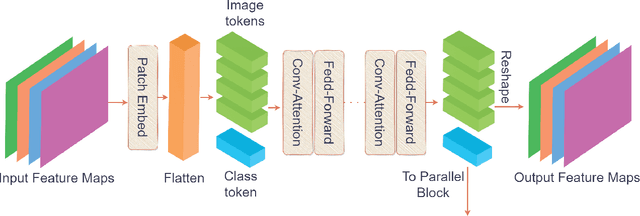
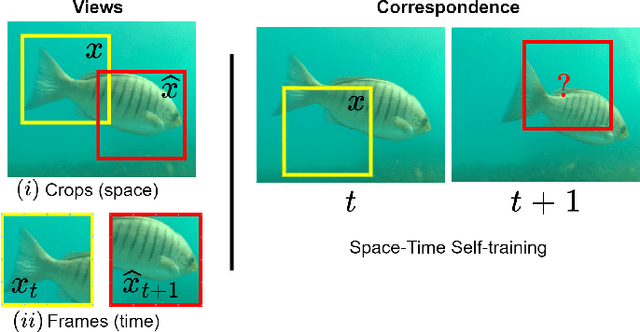
Abstract:Underwater fish segmentation to estimate fish body measurements is still largely unsolved due to the complex underwater environment. Relying on fully-supervised segmentation models requires collecting per-pixel labels, which is time-consuming and prone to overfitting. Self-supervised learning methods can help avoid the requirement of large annotated training datasets, however, to be useful in real-world applications, they should achieve good segmentation quality. In this paper, we introduce a Transformer-based method that uses self-supervision for high-quality fish segmentation. Our proposed model is trained on videos -- without any annotations -- to perform fish segmentation in underwater videos taken in situ in the wild. We show that when trained on a set of underwater videos from one dataset, the proposed model surpasses previous CNN-based and Transformer-based self-supervised methods and achieves performance relatively close to supervised methods on two new unseen underwater video datasets. This demonstrates the great generalisability of our model and the fact that it does not need a pre-trained model. In addition, we show that, due to its dense representation learning, our model is compute-efficient. We provide quantitative and qualitative results that demonstrate our model's significant capabilities.
Toward A Formalized Approach for Spike Sorting Algorithms and Hardware Evaluation
May 13, 2022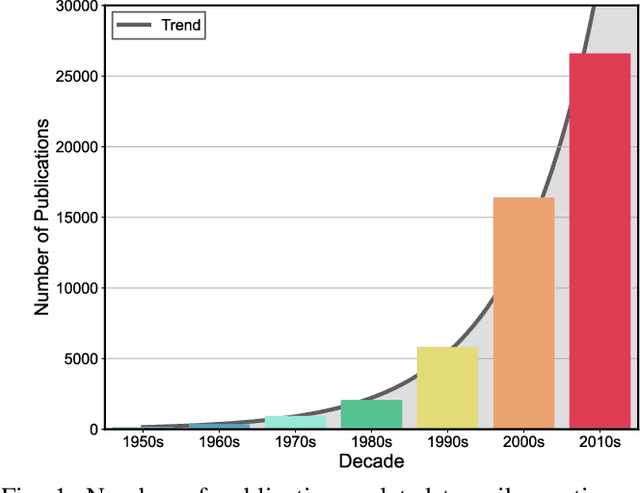
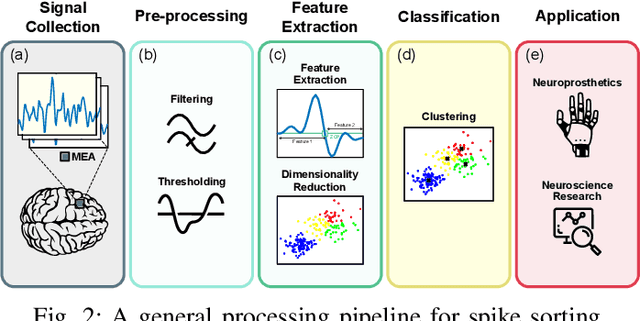
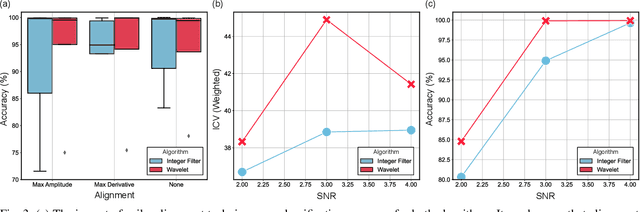
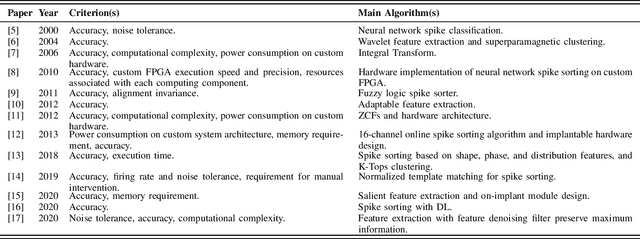
Abstract:Spike sorting algorithms are used to separate extracellular recordings of neuronal populations into single-unit spike activities. The development of customized hardware implementing spike sorting algorithms is burgeoning. However, there is a lack of a systematic approach and a set of standardized evaluation criteria to facilitate direct comparison of both software and hardware implementations. In this paper, we formalize a set of standardized criteria and a publicly available synthetic dataset entitled Synthetic Simulations Of Extracellular Recordings (SSOER), which was constructed by aggregating existing synthetic datasets with varying Signal-To-Noise Ratios (SNRs). Furthermore, we present a benchmark for future comparison, and use our criteria to evaluate a simulated Resistive Random-Access Memory (RRAM) In-Memory Computing (IMC) system using the Discrete Wavelet Transform (DWT) for feature extraction. Our system consumes approximately (per channel) 10.72mW and occupies an area of 0.66mm$^2$ in a 22nm FDSOI Complementary Metal-Oxide-Semiconductor (CMOS) process.
Computer Vision and Deep Learning for Fish Classification in Underwater Habitats: A Survey
Mar 15, 2022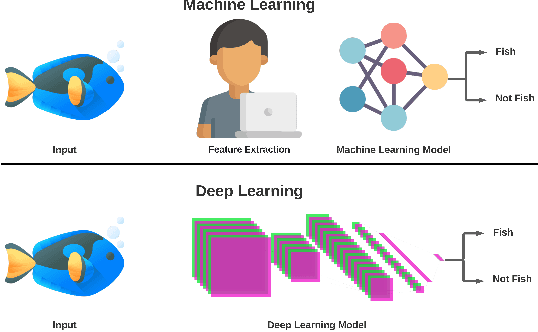
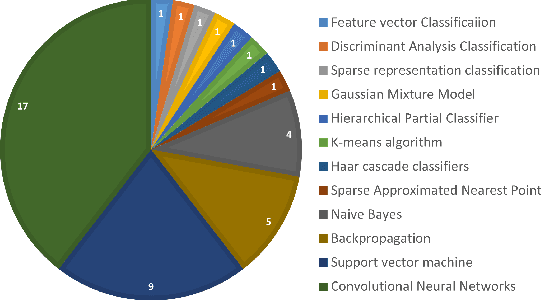
Abstract:Marine scientists use remote underwater video recording to survey fish species in their natural habitats. This helps them understand and predict how fish respond to climate change, habitat degradation, and fishing pressure. This information is essential for developing sustainable fisheries for human consumption, and for preserving the environment. However, the enormous volume of collected videos makes extracting useful information a daunting and time-consuming task for a human. A promising method to address this problem is the cutting-edge Deep Learning (DL) technology.DL can help marine scientists parse large volumes of video promptly and efficiently, unlocking niche information that cannot be obtained using conventional manual monitoring methods. In this paper, we provide an overview of the key concepts of DL, while presenting a survey of literature on fish habitat monitoring with a focus on underwater fish classification. We also discuss the main challenges faced when developing DL for underwater image processing and propose approaches to address them. Finally, we provide insights into the marine habitat monitoring research domain and shed light on what the future of DL for underwater image processing may hold. This paper aims to inform a wide range of readers from marine scientists who would like to apply DL in their research to computer scientists who would like to survey state-of-the-art DL-based underwater fish habitat monitoring literature.
Navigating Local Minima in Quantized Spiking Neural Networks
Feb 15, 2022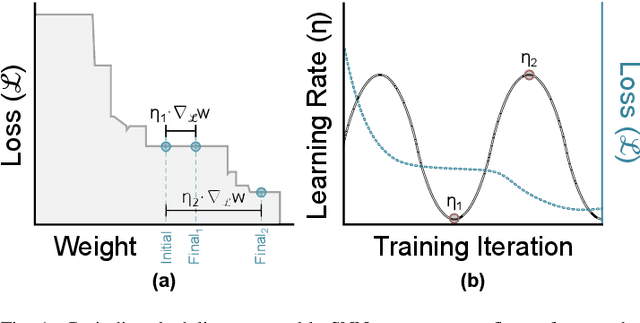
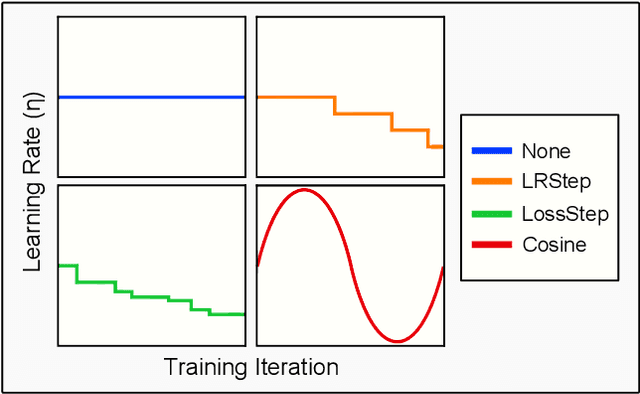
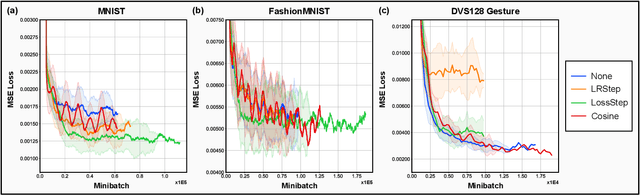
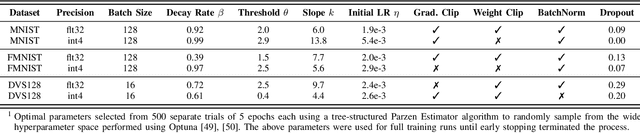
Abstract:Spiking and Quantized Neural Networks (NNs) are becoming exceedingly important for hyper-efficient implementations of Deep Learning (DL) algorithms. However, these networks face challenges when trained using error backpropagation, due to the absence of gradient signals when applying hard thresholds. The broadly accepted trick to overcoming this is through the use of biased gradient estimators: surrogate gradients which approximate thresholding in Spiking Neural Networks (SNNs), and Straight-Through Estimators (STEs), which completely bypass thresholding in Quantized Neural Networks (QNNs). While noisy gradient feedback has enabled reasonable performance on simple supervised learning tasks, it is thought that such noise increases the difficulty of finding optima in loss landscapes, especially during the later stages of optimization. By periodically boosting the Learning Rate (LR) during training, we expect the network can navigate unexplored solution spaces that would otherwise be difficult to reach due to local minima, barriers, or flat surfaces. This paper presents a systematic evaluation of a cosine-annealed LR schedule coupled with weight-independent adaptive moment estimation as applied to Quantized SNNs (QSNNs). We provide a rigorous empirical evaluation of this technique on high precision and 4-bit quantized SNNs across three datasets, demonstrating (close to) state-of-the-art performance on the more complex datasets. Our source code is available at this link: https://github.com/jeshraghian/QSNNs.
Design Space Exploration of Dense and Sparse Mapping Schemes for RRAM Architectures
Jan 25, 2022Abstract:The impact of device and circuit-level effects in mixed-signal Resistive Random Access Memory (RRAM) accelerators typically manifest as performance degradation of Deep Learning (DL) algorithms, but the degree of impact varies based on algorithmic features. These include network architecture, capacity, weight distribution, and the type of inter-layer connections. Techniques are continuously emerging to efficiently train sparse neural networks, which may have activation sparsity, quantization, and memristive noise. In this paper, we present an extended Design Space Exploration (DSE) methodology to quantify the benefits and limitations of dense and sparse mapping schemes for a variety of network architectures. While sparsity of connectivity promotes less power consumption and is often optimized for extracting localized features, its performance on tiled RRAM arrays may be more susceptible to noise due to under-parameterization, when compared to dense mapping schemes. Moreover, we present a case study quantifying and formalizing the trade-offs of typical non-idealities introduced into 1-Transistor-1-Resistor (1T1R) tiled memristive architectures and the size of modular crossbar tiles using the CIFAR-10 dataset.
A Deep Learning Localization Method for Measuring Abdominal Muscle Dimensions in Ultrasound Images
Sep 30, 2021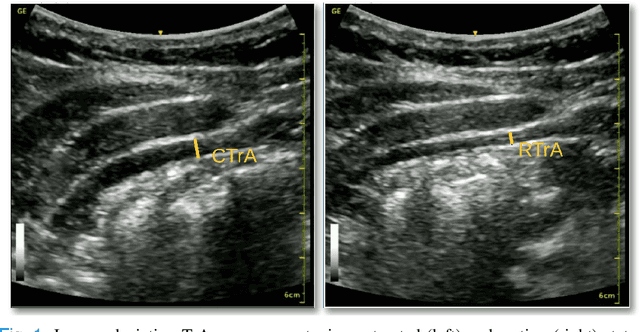
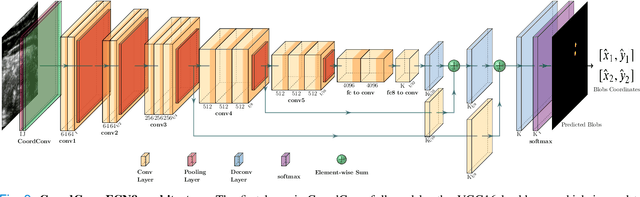
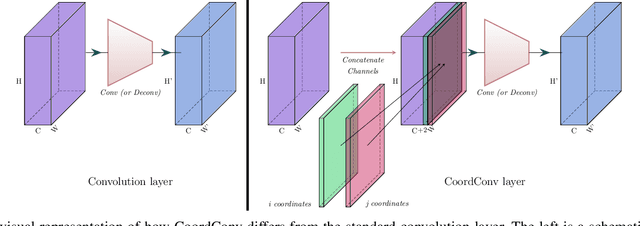

Abstract:Health professionals extensively use Two- Dimensional (2D) Ultrasound (US) videos and images to visualize and measure internal organs for various purposes including evaluation of muscle architectural changes. US images can be used to measure abdominal muscles dimensions for the diagnosis and creation of customized treatment plans for patients with Low Back Pain (LBP), however, they are difficult to interpret. Due to high variability, skilled professionals with specialized training are required to take measurements to avoid low intra-observer reliability. This variability stems from the challenging nature of accurately finding the correct spatial location of measurement endpoints in abdominal US images. In this paper, we use a Deep Learning (DL) approach to automate the measurement of the abdominal muscle thickness in 2D US images. By treating the problem as a localization task, we develop a modified Fully Convolutional Network (FCN) architecture to generate blobs of coordinate locations of measurement endpoints, similar to what a human operator does. We demonstrate that using the TrA400 US image dataset, our network achieves a Mean Absolute Error (MAE) of 0.3125 on the test set, which almost matches the performance of skilled ultrasound technicians. Our approach can facilitate next steps for automating the process of measurements in 2D US images, while reducing inter-observer as well as intra-observer variability for more effective clinical outcomes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge