Michael G. Schimek
Robust Node Affinities via Jaccard-Biased Random Walks and Rank Aggregation
Mar 05, 2026Abstract:Estimating node similarity is a fundamental task in network analysis and graph-based machine learning, with applications in clustering, community detection, classification, and recommendation. We propose TopKGraphs, a method based on start-node-anchored random walks that bias transitions toward nodes with structurally similar neighborhoods, measured via Jaccard similarity. Rather than computing stationary distributions, walks are treated as stochastic neighborhood samplers, producing partial node rankings that are aggregated using robust rank aggregation to construct interpretable node-to-node affinity matrices. TopKGraphs provides a non-parametric, interpretable, and general-purpose representation of node similarity that can be applied in both network analysis and machine learning workflows. We evaluate the method on synthetic graphs (stochastic block models, Lancichinetti-Fortunato-Radicchi benchmark graphs), k-nearest-neighbor graphs from tabular datasets, and a curated high-confidence protein-protein interaction network. Across all scenarios, TopKGraphs achieves competitive or superior performance compared to standard similarity measures (Jaccard, Dice), a diffusion-based method (personalized PageRank), and an embedding-based approach (Node2Vec), demonstrating robustness in sparse, noisy, or heterogeneous networks. These results suggest that TopKGraphs is a versatile and interpretable tool for bridging simple local similarity measures with more complex embedding-based approaches, facilitating both data mining and network analysis applications.
Parea: multi-view ensemble clustering for cancer subtype discovery
Sep 30, 2022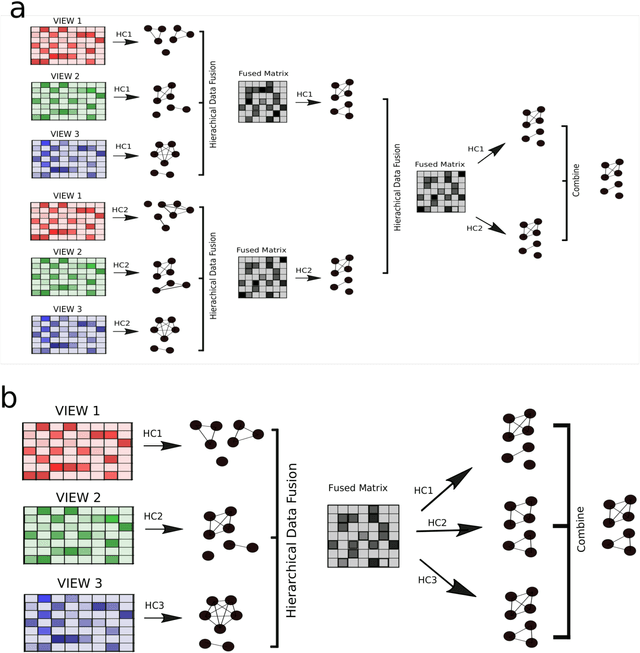
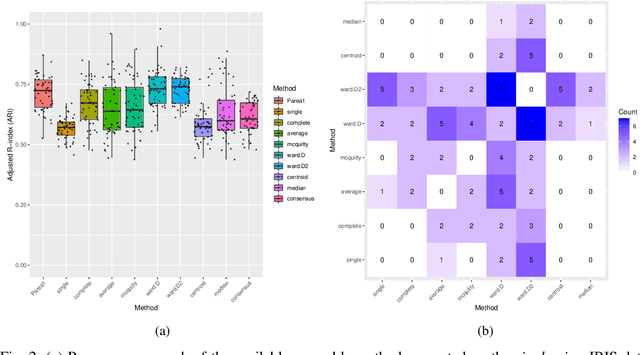
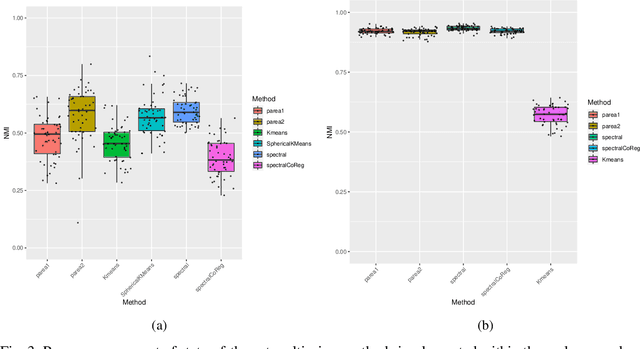
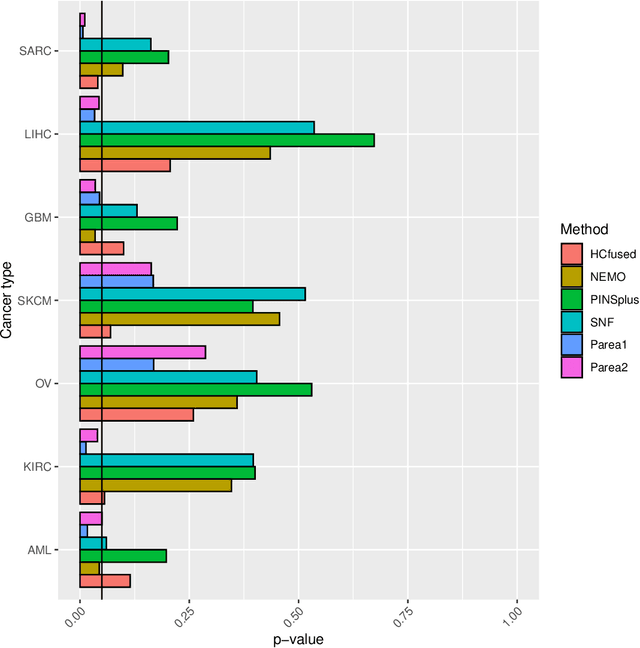
Abstract:Multi-view clustering methods are essential for the stratification of patients into sub-groups of similar molecular characteristics. In recent years, a wide range of methods has been developed for this purpose. However, due to the high diversity of cancer-related data, a single method may not perform sufficiently well in all cases. We present Parea, a multi-view hierarchical ensemble clustering approach for disease subtype discovery. We demonstrate its performance on several machine learning benchmark datasets. We apply and validate our methodology on real-world multi-view cancer patient data. Parea outperforms the current state-of-the-art on six out of seven analysed cancer types. We have integrated the Parea method into our developed Python package Pyrea (https://github.com/mdbloice/Pyrea), which enables the effortless and flexible design of ensemble workflows while incorporating a wide range of fusion and clustering algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge