Michael Escobar
Comparing Variation in Tokenizer Outputs Using a Series of Problematic and Challenging Biomedical Sentences
May 15, 2023


Abstract:Background & Objective: Biomedical text data are increasingly available for research. Tokenization is an initial step in many biomedical text mining pipelines. Tokenization is the process of parsing an input biomedical sentence (represented as a digital character sequence) into a discrete set of word/token symbols, which convey focused semantic/syntactic meaning. The objective of this study is to explore variation in tokenizer outputs when applied across a series of challenging biomedical sentences. Method: Diaz [2015] introduce 24 challenging example biomedical sentences for comparing tokenizer performance. In this study, we descriptively explore variation in outputs of eight tokenizers applied to each example biomedical sentence. The tokenizers compared in this study are the NLTK white space tokenizer, the NLTK Penn Tree Bank tokenizer, Spacy and SciSpacy tokenizers, Stanza/Stanza-Craft tokenizers, the UDPipe tokenizer, and R-tokenizers. Results: For many examples, tokenizers performed similarly effectively; however, for certain examples, there were meaningful variation in returned outputs. The white space tokenizer often performed differently than other tokenizers. We observed performance similarities for tokenizers implementing rule-based systems (e.g. pattern matching and regular expressions) and tokenizers implementing neural architectures for token classification. Oftentimes, the challenging tokens resulting in the greatest variation in outputs, are those words which convey substantive and focused biomedical/clinical meaning (e.g. x-ray, IL-10, TCR/CD3, CD4+ CD8+, and (Ca2+)-regulated). Conclusion: When state-of-the-art, open-source tokenizers from Python and R were applied to a series of challenging biomedical example sentences, we observed subtle variation in the returned outputs.
Doubly Robust Estimation with Machine Learning Predictions
Aug 03, 2021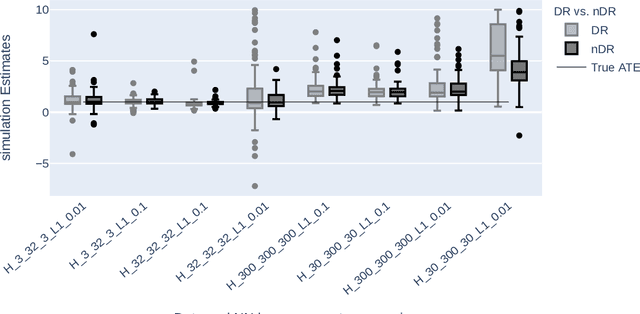
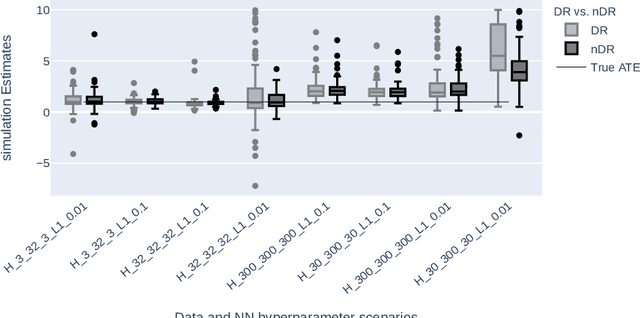
Abstract:The estimation of Average Treatment Effect (ATE) as a causal parameter is carried out in two steps, wherein the first step, the treatment, and outcome are modeled to incorporate the potential confounders, and in the second step, the predictions are inserted into the ATE estimators such as the Augmented Inverse Probability Weighting (AIPW) estimator. Due to the concerns regarding the nonlinear or unknown relationships between confounders and the treatment and outcome, there has been an interest in applying non-parametric methods such as Machine Learning (ML) algorithms instead. \cite{farrell2018deep} proposed to use two separate Neural Networks (NNs) where there's no regularization on the network's parameters except the Stochastic Gradient Descent (SGD) in the NN's optimization. Our simulations indicate that the AIPW estimator suffers extensively if no regularization is utilized. We propose the normalization of AIPW (referred to as nAIPW) which can be helpful in some scenarios. nAIPW, provably, has the same properties as AIPW, that is double-robustness and orthogonality \citep{chernozhukov2018double}. Further, if the first step algorithms converge fast enough, under regulatory conditions \citep{chernozhukov2018double}, nAIPW will be asymptotically normal.
The Bias-Variance Tradeoff of Doubly Robust Estimator with Targeted $L_1$ regularized Neural Networks Predictions
Aug 02, 2021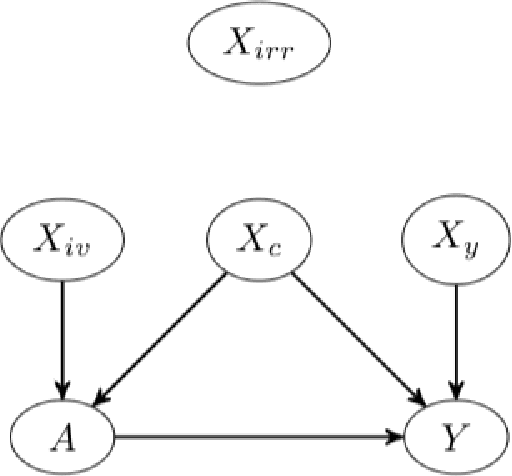
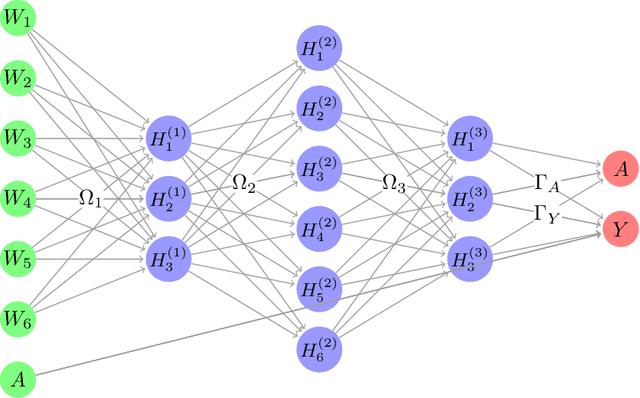
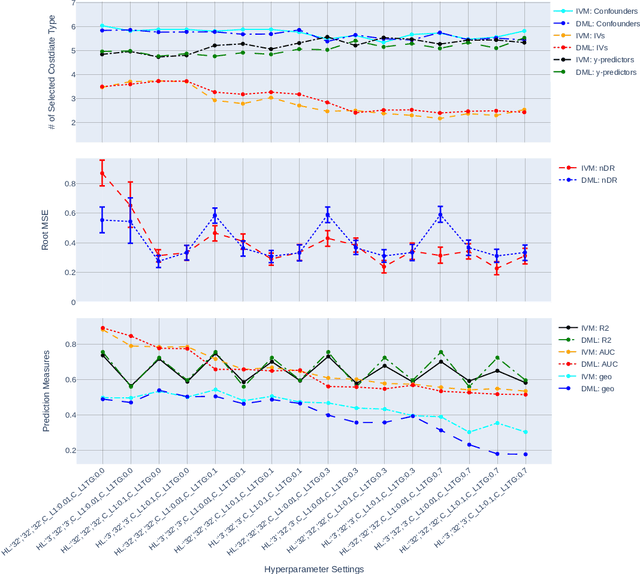
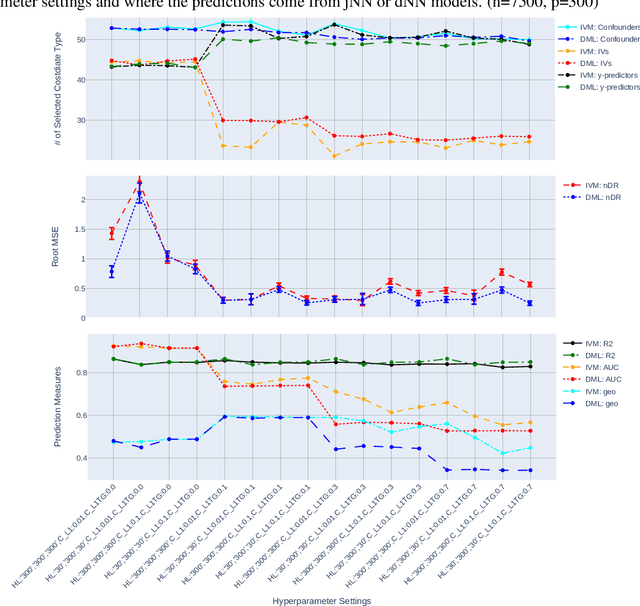
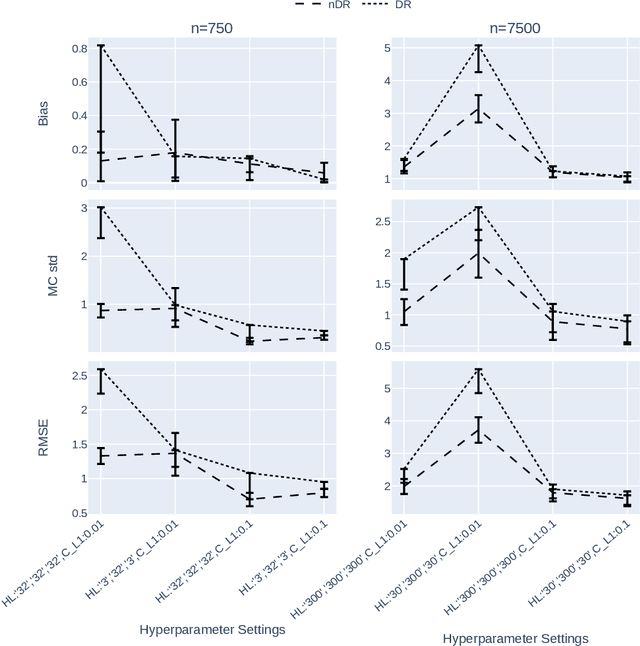
Abstract:The Doubly Robust (DR) estimation of ATE can be carried out in 2 steps, where in the first step, the treatment and outcome are modeled, and in the second step the predictions are inserted into the DR estimator. The model misspecification in the first step has led researchers to utilize Machine Learning algorithms instead of parametric algorithms. However, existence of strong confounders and/or Instrumental Variables (IVs) can lead the complex ML algorithms to provide perfect predictions for the treatment model which can violate the positivity assumption and elevate the variance of DR estimators. Thus the ML algorithms must be controlled to avoid perfect predictions for the treatment model while still learn the relationship between the confounders and the treatment and outcome. We use two Neural network architectures and investigate how their hyperparameters should be tuned in the presence of confounders and IVs to achieve a low bias-variance tradeoff for ATE estimators such as DR estimator. Through simulation results, we will provide recommendations as to how NNs can be employed for ATE estimation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge