Matthias Nickles
Drug Similarity and Link Prediction Using Graph Embeddings on Medical Knowledge Graphs
Oct 29, 2021
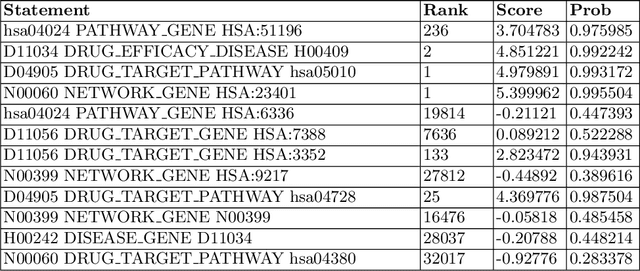
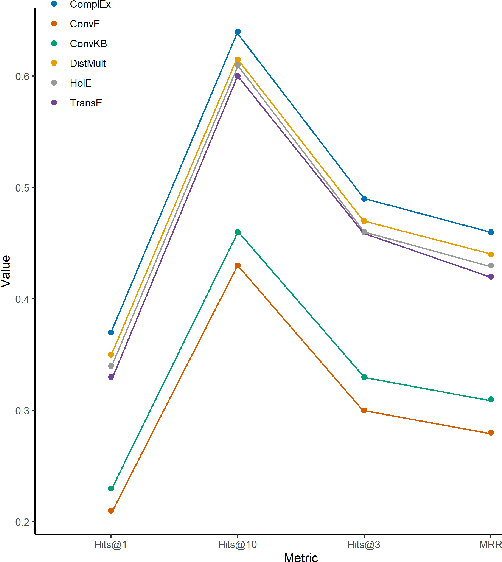
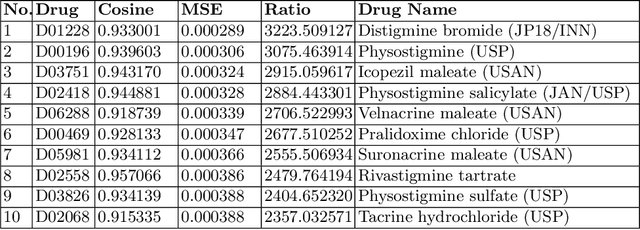
Abstract:The paper utilizes the graph embeddings generated for entities of a large biomedical database to perform link prediction to capture various new relationships among different entities. A novel node similarity measure is proposed that utilizes the graph embeddings and link prediction scores to find similarity scores among various drugs which can be used by the medical experts to recommend alternative drugs to avoid side effects from original one. Utilizing machine learning on knowledge graph for drug similarity and recommendation will be less costly and less time consuming with higher scalability as compared to traditional biomedical methods due to the dependency on costly medical equipment and experts of the latter ones.
diff-SAT -- A Software for Sampling and Probabilistic Reasoning for SAT and Answer Set Programming
Jan 03, 2021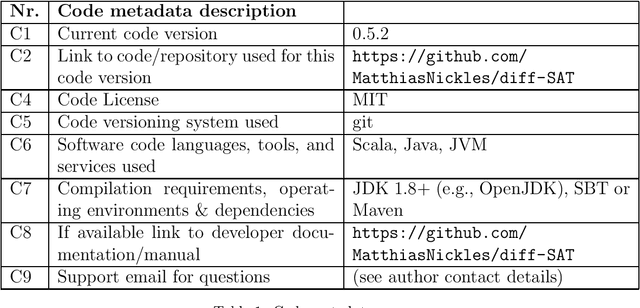
Abstract:This paper describes diff-SAT, an Answer Set and SAT solver which combines regular solving with the capability to use probabilistic clauses, facts and rules, and to sample an optimal world-view (multiset of satisfying Boolean variable assignments or answer sets) subject to user-provided probabilistic constraints. The sampling process minimizes a user-defined differentiable objective function using a gradient descent based optimization method called Differentiable Satisfiability Solving ($\partial\mathrm{SAT}$) respectively Differentiable Answer Set Programming ($\partial\mathrm{ASP}$). Use cases are i.a. probabilistic logic programming (in form of Probabilistic Answer Set Programming), Probabilistic Boolean Satisfiability solving (PSAT), and distribution-aware sampling of model multisets (answer sets or Boolean interpretations).
Differentiable Satisfiability and Differentiable Answer Set Programming for Sampling-Based Multi-Model Optimization
Dec 31, 2018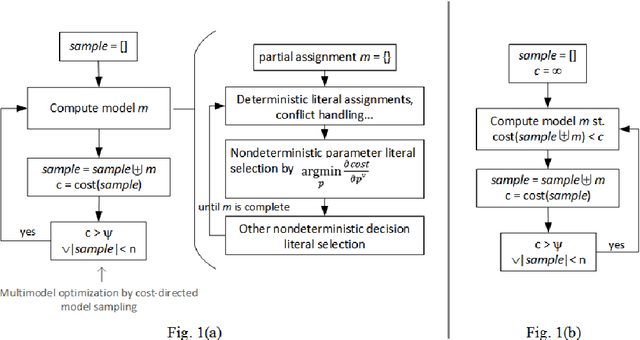
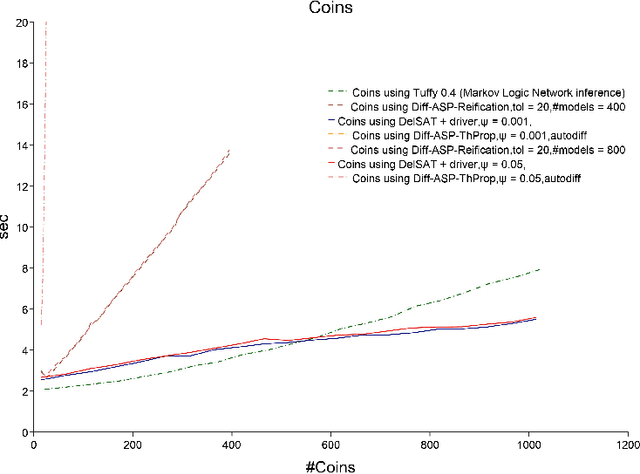
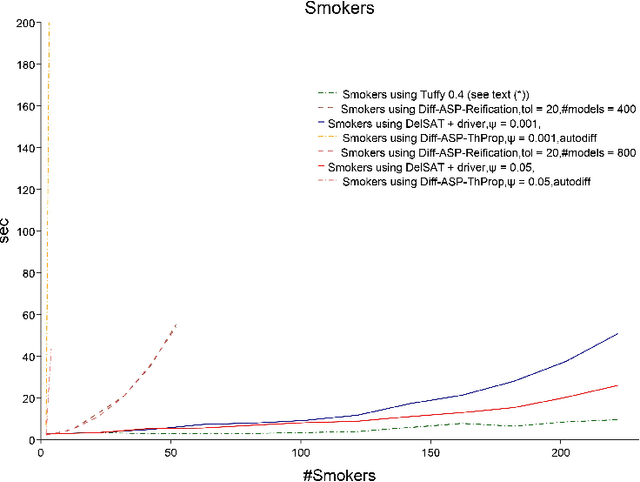
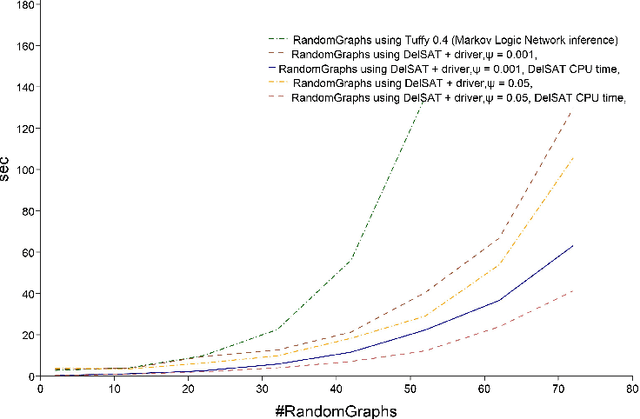
Abstract:We propose Differentiable Satisfiability and Differentiable Answer Set Programming (Differentiable SAT/ASP) for multi-model optimization. Models (answer sets or satisfying truth assignments) are sampled using a novel SAT/ASP solving approach which uses a gradient descent-based branching mechanism. Sampling proceeds until the value of a user-defined multi-model cost function reaches a given threshold. As major use cases for our approach we propose distribution-aware model sampling and expressive yet scalable probabilistic logic programming. As our main algorithmic approach to Differentiable SAT/ASP, we introduce an enhancement of the state-of-the-art CDNL/CDCL algorithm for SAT/ASP solving. Additionally, we present alternative algorithms which use an unmodified ASP solver (Clingo/clasp) and map the optimization task to conventional answer set optimization or use so-called propagators. We also report on the open source software DelSAT, a recent prototype implementation of our main algorithm, and on initial experimental results which indicate that DelSATs performance is, when applied to the use case of probabilistic logic inference, on par with Markov Logic Network (MLN) inference performance, despite having advantageous properties compared to MLNs, such as the ability to express inductive definitions and to work with probabilities as weights directly in all cases. Our experiments also indicate that our main algorithm is strongly superior in terms of performance compared to the presented alternative approaches which reduce a common instance of the general problem to regular SAT/ASP.
Embedding Cardinality Constraints in Neural Link Predictors
Dec 16, 2018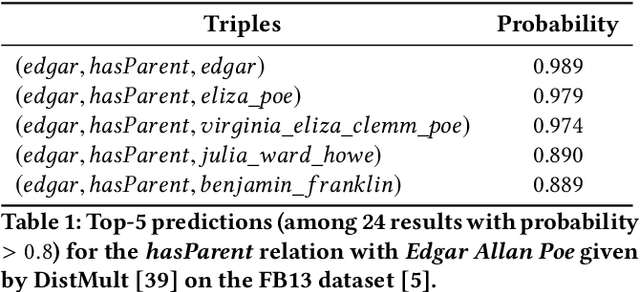
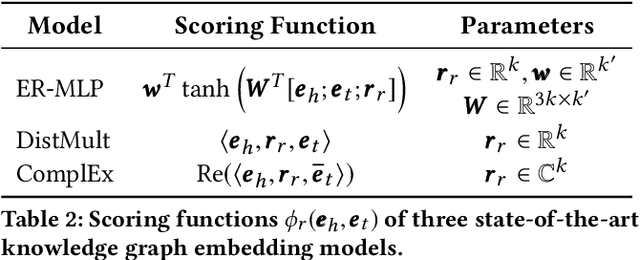
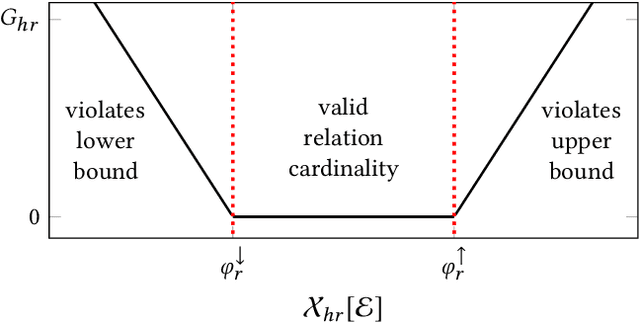

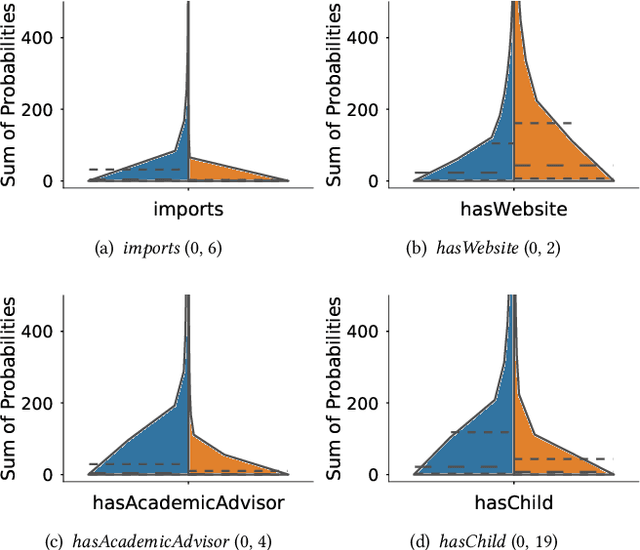
Abstract:Neural link predictors learn distributed representations of entities and relations in a knowledge graph. They are remarkably powerful in the link prediction and knowledge base completion tasks, mainly due to the learned representations that capture important statistical dependencies in the data. Recent works in the area have focused on either designing new scoring functions or incorporating extra information into the learning process to improve the representations. Yet the representations are mostly learned from the observed links between entities, ignoring commonsense or schema knowledge associated with the relations in the graph. A fundamental aspect of the topology of relational data is the cardinality information, which bounds the number of predictions given for a relation between a minimum and maximum frequency. In this paper, we propose a new regularisation approach to incorporate relation cardinality constraints to any existing neural link predictor without affecting their efficiency or scalability. Our regularisation term aims to impose boundaries on the number of predictions with high probability, thus, structuring the embeddings space to respect commonsense cardinality assumptions resulting in better representations. Experimental results on Freebase, WordNet and YAGO show that, given suitable prior knowledge, the proposed method positively impacts the predictive accuracy of downstream link prediction tasks.
PrASP Report
Dec 30, 2016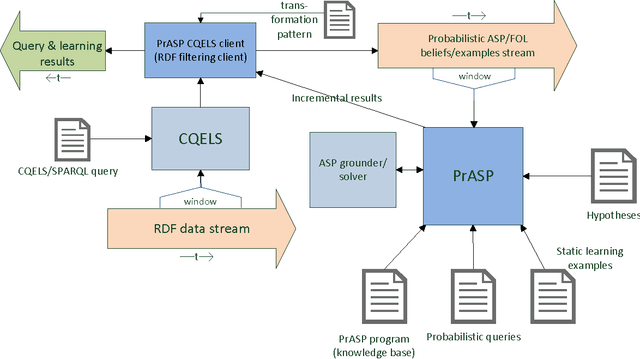
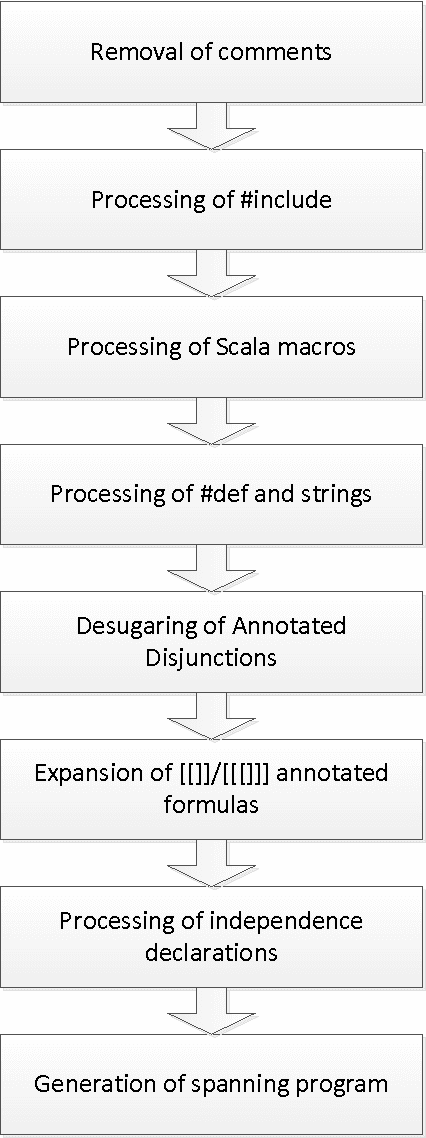
Abstract:This technical report describes the usage, syntax, semantics and core algorithms of the probabilistic inductive logic programming framework PrASP. PrASP is a research software which integrates non-monotonic reasoning based on Answer Set Programming (ASP), probabilistic inference and parameter learning. In contrast to traditional approaches to Probabilistic (Inductive) Logic Programming, our framework imposes only little restrictions on probabilistic logic programs. In particular, PrASP allows for ASP as well as First-Order Logic syntax, and for the annotation of formulas with point probabilities as well as interval probabilities. A range of widely configurable inference algorithms can be combined in a pipeline-like fashion, in order to cover a variety of use cases.
Probabilistic Inductive Logic Programming Based on Answer Set Programming
May 04, 2014Abstract:We propose a new formal language for the expressive representation of probabilistic knowledge based on Answer Set Programming (ASP). It allows for the annotation of first-order formulas as well as ASP rules and facts with probabilities and for learning of such weights from data (parameter estimation). Weighted formulas are given a semantics in terms of soft and hard constraints which determine a probability distribution over answer sets. In contrast to related approaches, we approach inference by optionally utilizing so-called streamlining XOR constraints, in order to reduce the number of computed answer sets. Our approach is prototypically implemented. Examples illustrate the introduced concepts and point at issues and topics for future research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge