Mary M. Maleckar
Balancing Fidelity, Utility, and Privacy in Synthetic Cardiac MRI Generation: A Comparative Study
Mar 04, 2026Abstract:Deep learning in cardiac MRI (CMR) is fundamentally constrained by both data scarcity and privacy regulations. This study systematically benchmarks three generative architectures: Denoising Diffusion Probabilistic Models (DDPM), Latent Diffusion Models (LDM), and Flow Matching (FM) for synthetic CMR generation. Utilizing a two-stage pipeline where anatomical masks condition image synthesis, we evaluate generated data across three critical axes: fidelity, utility, and privacy. Our results show that diffusion-based models, particularly DDPM, provide the most effective balance between downstream segmentation utility, image fidelity, and privacy preservation under limited-data conditions, while FM demonstrates promising privacy characteristics with slightly lower task-level performance. These findings quantify the trade-offs between cross-domain generalization and patient confidentiality, establishing a framework for safe and effective synthetic data augmentation in medical imaging.
Generative Modeling with Conditional Autoencoders: Building an Integrated Cell
Apr 28, 2017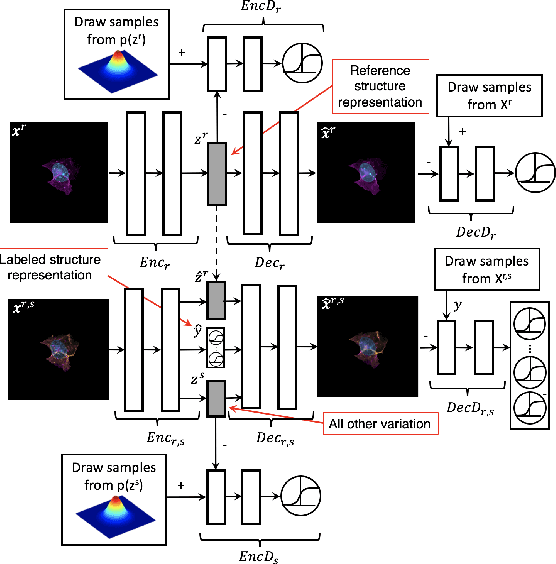
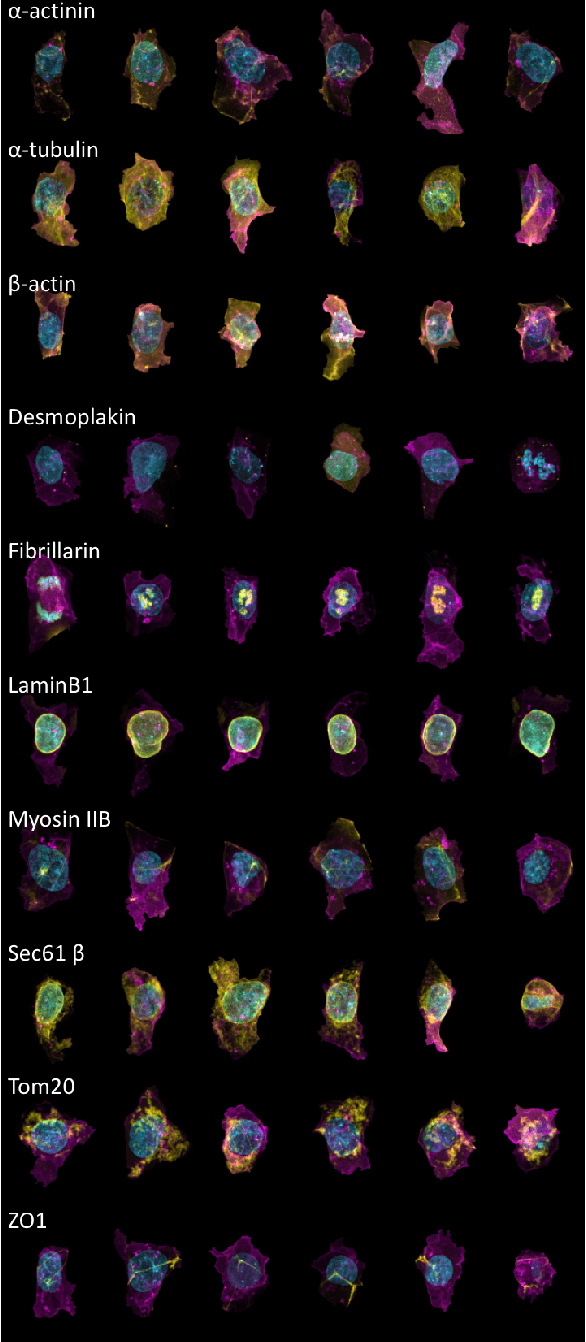
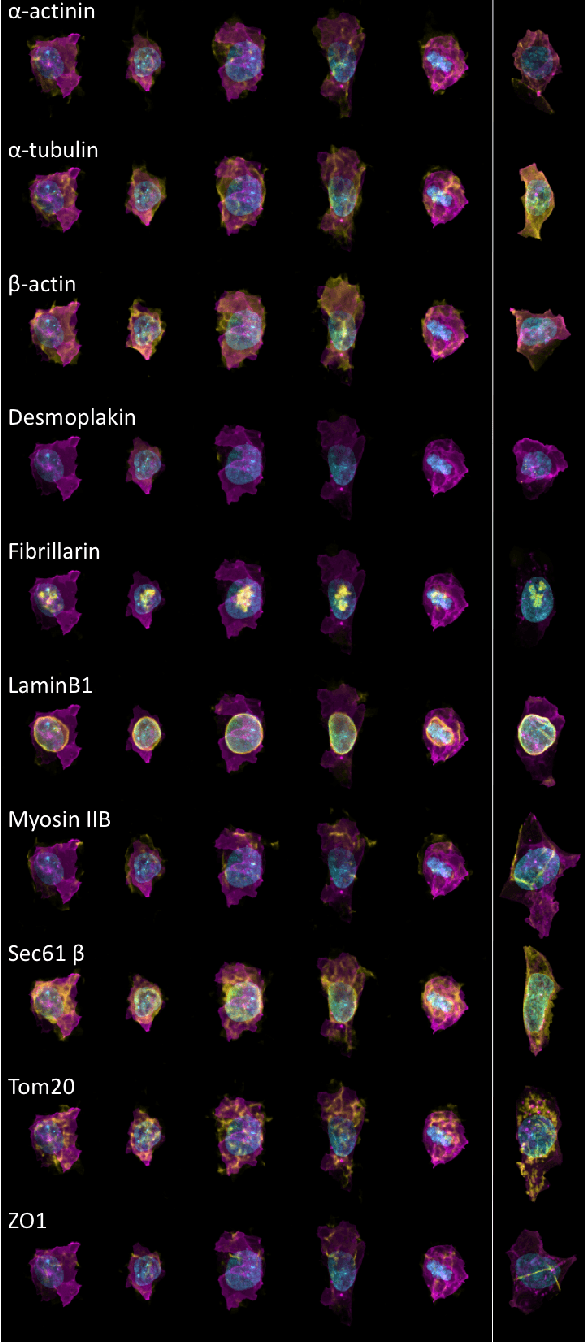
Abstract:We present a conditional generative model to learn variation in cell and nuclear morphology and the location of subcellular structures from microscopy images. Our model generalizes to a wide range of subcellular localization and allows for a probabilistic interpretation of cell and nuclear morphology and structure localization from fluorescence images. We demonstrate the effectiveness of our approach by producing photo-realistic cell images using our generative model. The conditional nature of the model provides the ability to predict the localization of unobserved structures given cell and nuclear morphology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge