Marleen de Bruijne
for the ALFA study
An End-to-end Approach to Semantic Segmentation with 3D CNN and Posterior-CRF in Medical Images
Nov 08, 2018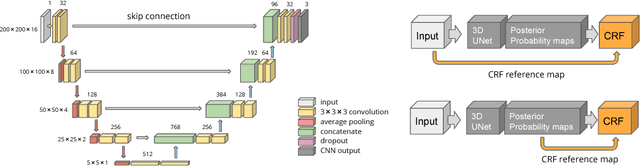
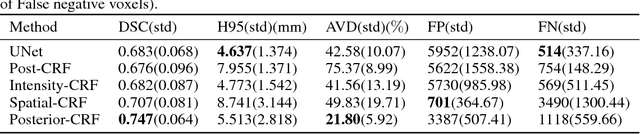
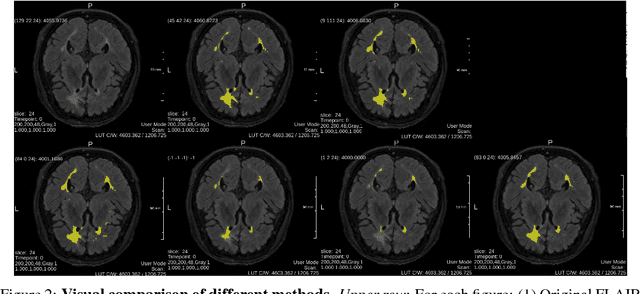
Abstract:Fully-connected Conditional Random Field (CRF) is often used as post-processing to refine voxel classification results by encouraging spatial coherence. In this paper, we propose a new end-to-end training method called Posterior-CRF. In contrast with previous approaches which use the original image intensity in the CRF, our approach applies 3D, fully connected CRF to the posterior probabilities from a CNN and optimizes both CNN and CRF together. The experiments on white matter hyperintensities segmentation demonstrate that our method outperforms CNN, post-processing CRF and different end-to-end training CRF approaches.
3D Regression Neural Network for the Quantification of Enlarged Perivascular Spaces in Brain MRI
Oct 28, 2018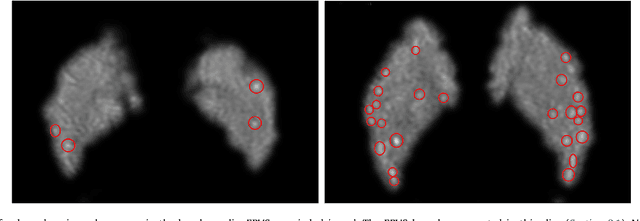
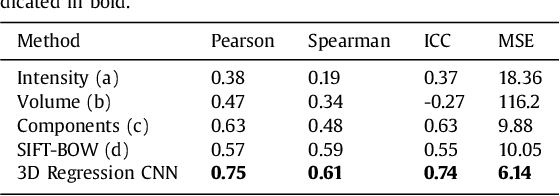
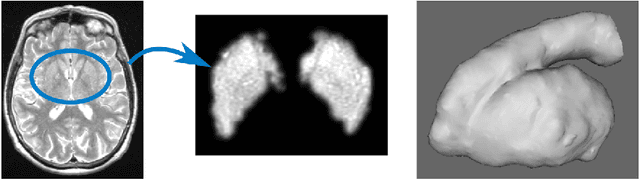
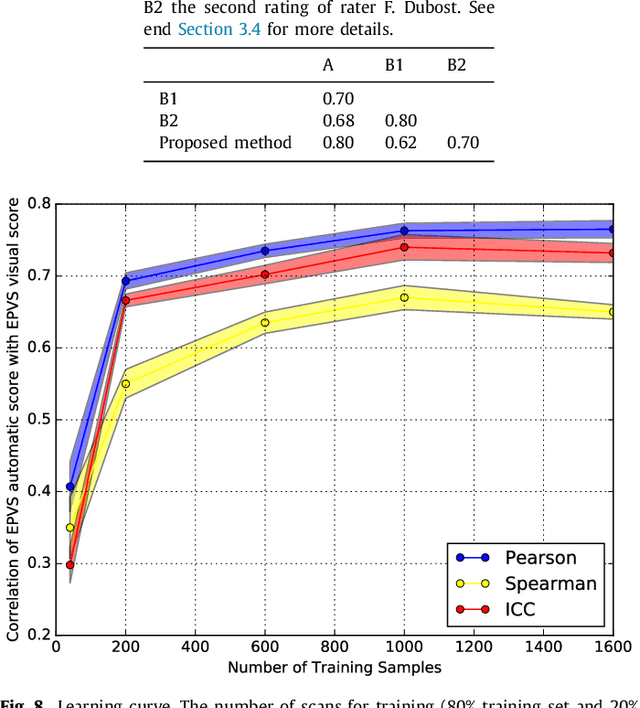
Abstract:Enlarged perivascular spaces (EPVS) in the brain are an emerging imaging marker for cerebral small vessel disease, and have been shown to be related to increased risk of various neurological diseases, including stroke and dementia. Automatic quantification of EPVS would greatly help to advance research into its etiology and its potential as a risk indicator of disease. We propose a convolutional network regression method to quantify the extent of EPVS in the basal ganglia from 3D brain MRI. We first segment the basal ganglia and subsequently apply a 3D convolutional regression network designed for small object detection within this region of interest. The network takes an image as input, and outputs a quantification score of EPVS. The network has significantly more convolution operations than pooling ones and no final activation, allowing it to span the space of real numbers. We validated our approach using a dataset of 2000 brain MRI scans scored visually. Experiments with varying sizes of training and test sets showed that a good performance can be achieved with a training set of only 200 scans. With a training set of 1000 scans, the intraclass correlation coefficient (ICC) between our scoring method and the expert's visual score was 0.74. Our method outperforms by a large margin - more than 0.10 - four more conventional automated approaches based on intensities, scale-invariant feature transform, and random forest. We show that the network learns the structures of interest and investigate the influence of hyper-parameters on the performance. We also evaluate the reproducibility of our network using a set of 60 subjects scanned twice (scan-rescan reproducibility). On this set our network achieves an ICC of 0.93, while the intrarater agreement reaches 0.80. Furthermore, the automatic EPVS scoring correlates similarly to age as visual scoring.
Learning to quantify emphysema extent: What labels do we need?
Oct 17, 2018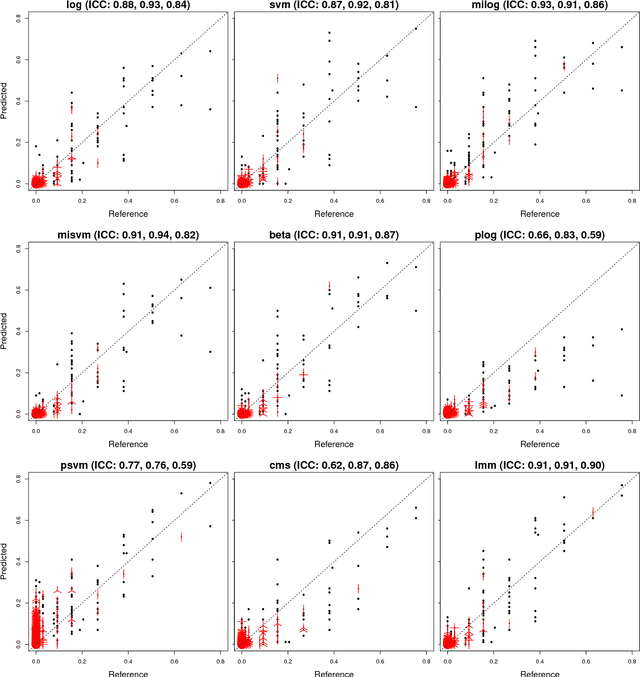
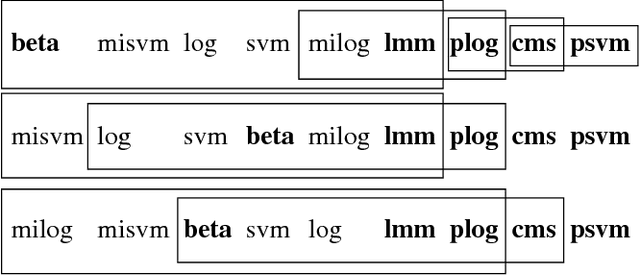

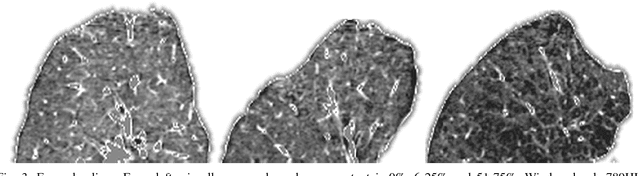
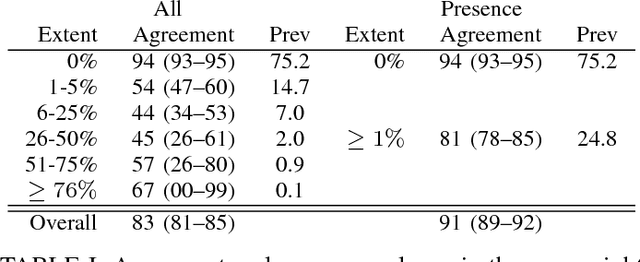
Abstract:Accurate assessment of pulmonary emphysema is crucial to assess disease severity and subtype, to monitor disease progression and to predict lung cancer risk. However, visual assessment is time-consuming and subject to substantial inter-rater variability and standard densitometry approaches to quantify emphysema remain inferior to visual scoring. We explore if machine learning methods that learn from a large dataset of visually assessed CT scans can provide accurate estimates of emphysema extent. We further investigate if machine learning algorithms that learn from a scoring of emphysema extent can outperform algorithms that learn only from a scoring of emphysema presence. We compare four Multiple Instance Learning classifiers that are trained on emphysema presence labels, and five Learning with Label Proportions classifiers that are trained on emphysema extent labels. We evaluate performance on 600 low-dose CT scans from the Danish Lung Cancer Screening Trial and find that learning from emphysema presence labels, which are much easier to obtain, gives equally good performance to learning from emphysema extent labels. The best classifiers achieve intra-class correlation coefficients around 0.90 and average overall agreement with raters of 78% and 79% on six emphysema extent classes versus inter-rater agreement of 83%.
Automatic Emphysema Detection using Weakly Labeled HRCT Lung Images
Oct 01, 2018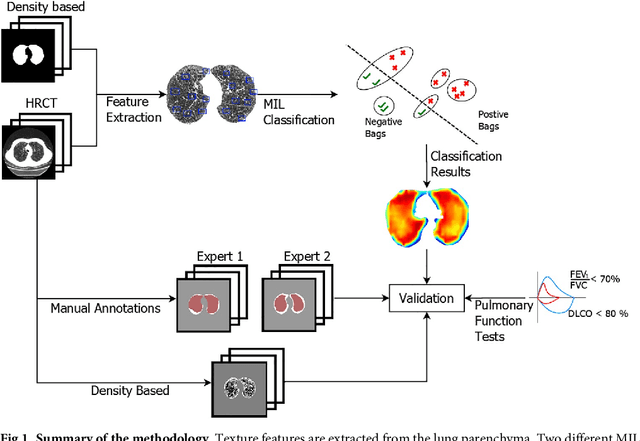
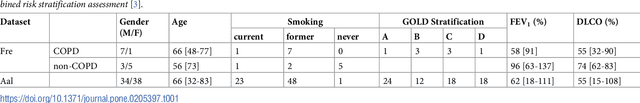
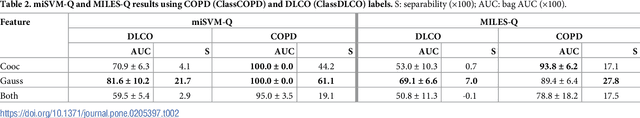
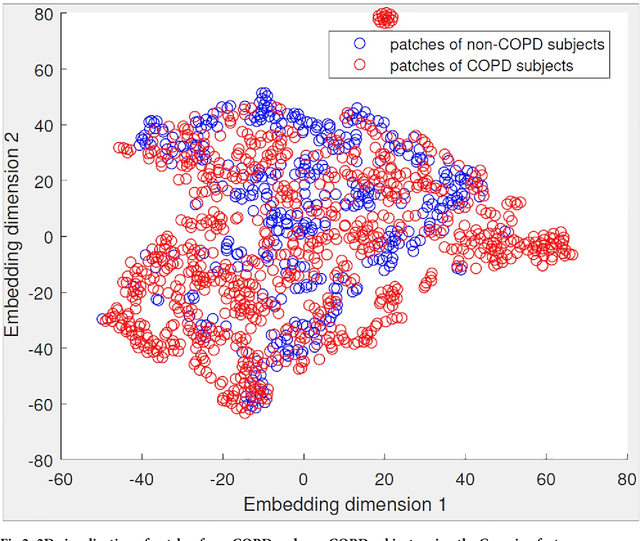
Abstract:A method for automatically quantifying emphysema regions using High-Resolution Computed Tomography (HRCT) scans of patients with chronic obstructive pulmonary disease (COPD) that does not require manually annotated scans for training is presented. HRCT scans of controls and of COPD patients with diverse disease severity are acquired at two different centers. Textural features from co-occurrence matrices and Gaussian filter banks are used to characterize the lung parenchyma in the scans. Two robust versions of multiple instance learning (MIL) classifiers, miSVM and MILES, are investigated. The classifiers are trained with the weak labels extracted from the forced expiratory volume in one minute (FEV$_1$) and diffusing capacity of the lungs for carbon monoxide (DLCO). At test time, the classifiers output a patient label indicating overall COPD diagnosis and local labels indicating the presence of emphysema. The classifier performance is compared with manual annotations by two radiologists, a classical density based method, and pulmonary function tests (PFTs). The miSVM classifier performed better than MILES on both patient and emphysema classification. The classifier has a stronger correlation with PFT than the density based method, the percentage of emphysema in the intersection of annotations from both radiologists, and the percentage of emphysema annotated by one of the radiologists. The correlation between the classifier and the PFT is only outperformed by the second radiologist. The method is therefore promising for facilitating assessment of emphysema and reducing inter-observer variability.
Not-so-supervised: a survey of semi-supervised, multi-instance, and transfer learning in medical image analysis
Sep 14, 2018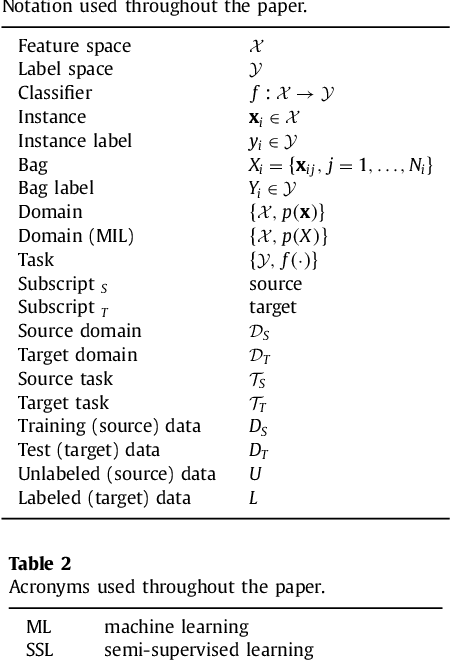
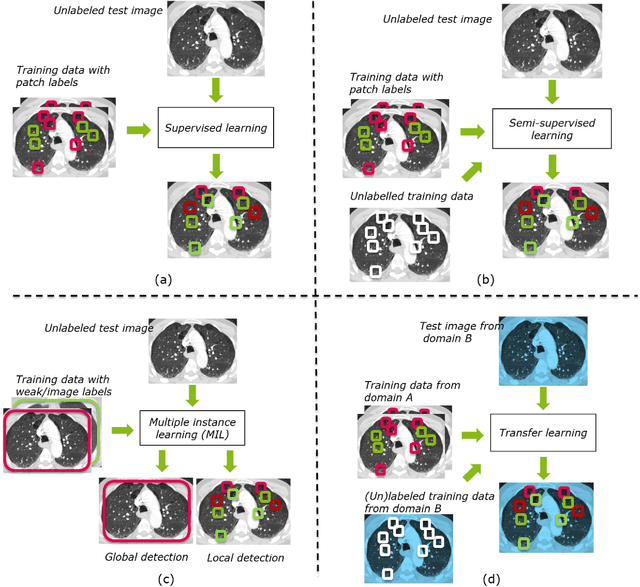

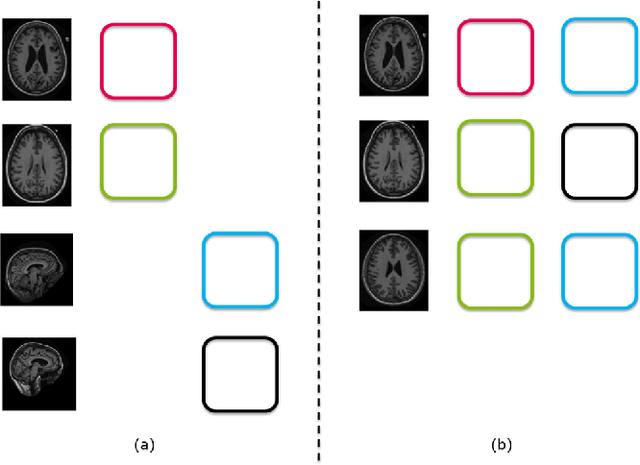
Abstract:Machine learning (ML) algorithms have made a tremendous impact in the field of medical imaging. While medical imaging datasets have been growing in size, a challenge for supervised ML algorithms that is frequently mentioned is the lack of annotated data. As a result, various methods which can learn with less/other types of supervision, have been proposed. We review semi-supervised, multiple instance, and transfer learning in medical imaging, both in diagnosis/detection or segmentation tasks. We also discuss connections between these learning scenarios, and opportunities for future research.
Deep Learning from Label Proportions for Emphysema Quantification
Jul 23, 2018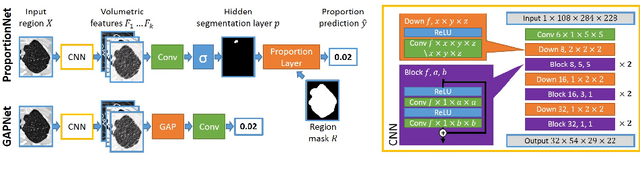
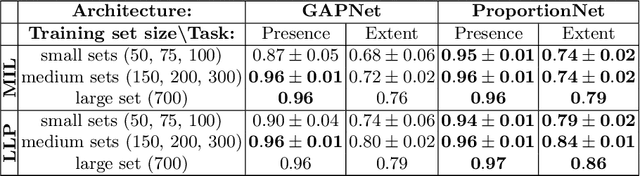
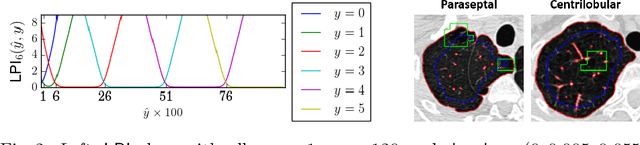
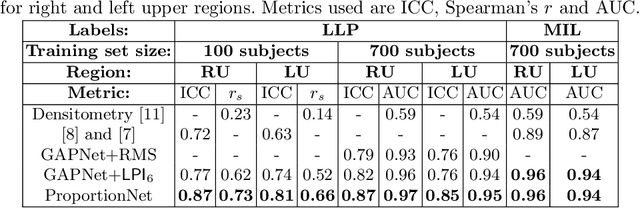
Abstract:We propose an end-to-end deep learning method that learns to estimate emphysema extent from proportions of the diseased tissue. These proportions were visually estimated by experts using a standard grading system, in which grades correspond to intervals (label example: 1-5% of diseased tissue). The proposed architecture encodes the knowledge that the labels represent a volumetric proportion. A custom loss is designed to learn with intervals. Thus, during training, our network learns to segment the diseased tissue such that its proportions fit the ground truth intervals. Our architecture and loss combined improve the performance substantially (8% ICC) compared to a more conventional regression network. We outperform traditional lung densitometry and two recently published methods for emphysema quantification by a large margin (at least 7% AUC and 15% ICC), and achieve near-human-level performance. Moreover, our method generates emphysema segmentations that predict the spatial distribution of emphysema at human level.
Hydranet: Data Augmentation for Regression Neural Networks
Jul 12, 2018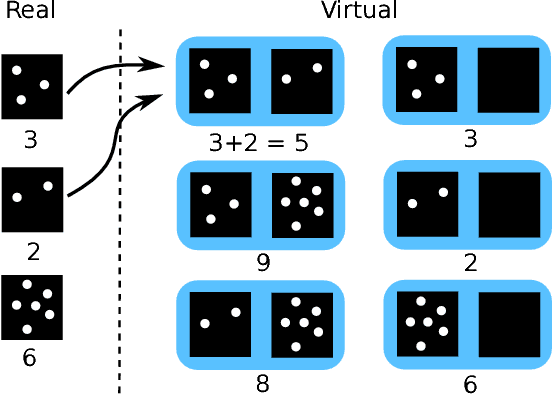
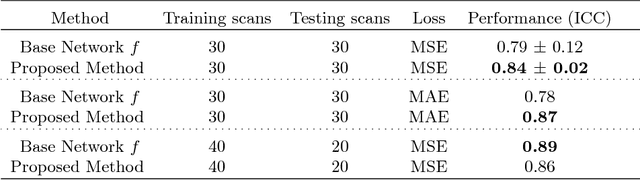
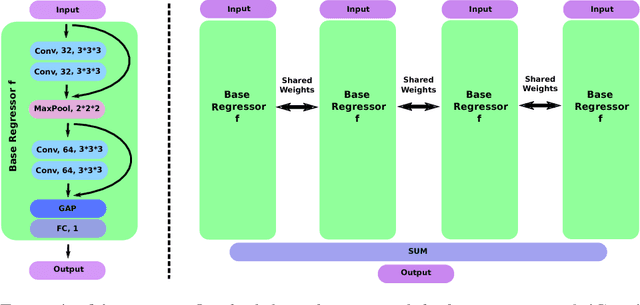
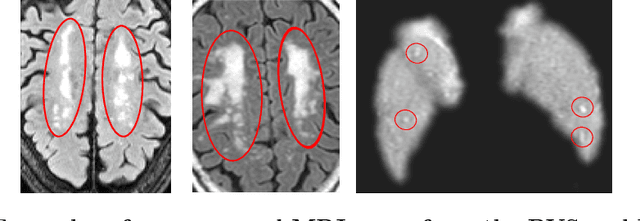
Abstract:Despite recent efforts, deep learning techniques remain often heavily dependent on a large quantity of labeled data. This problem is even more challenging in medical image analysis where the annotator expertise is often scarce. In this paper we propose a novel data-augmentation method to regularize neural network regressors, learning from a single global label per image. The principle of the method is to create new samples by recombining existing ones. We demonstrate the performance of our algorithm on two tasks: the regression of number of enlarged perivascular spaces in the basal ganglia; and the regression of white matter hyperintensities volume. We show that the proposed method improves the performance even when more basic data augmentation is used. Furthermore we reached an intraclass correlation coefficient between ground truth and network predictions of 0.73 on the first task and 0.86 on the second task, only using between 25 and 30 scans with a single global label per scan for training. To achieve a similar correlation on the first task, state-of-the-art methods needed more than 1000 training scans.
Extracting Tree-structures in CT data by Tracking Multiple Statistically Ranked Hypotheses
Jun 23, 2018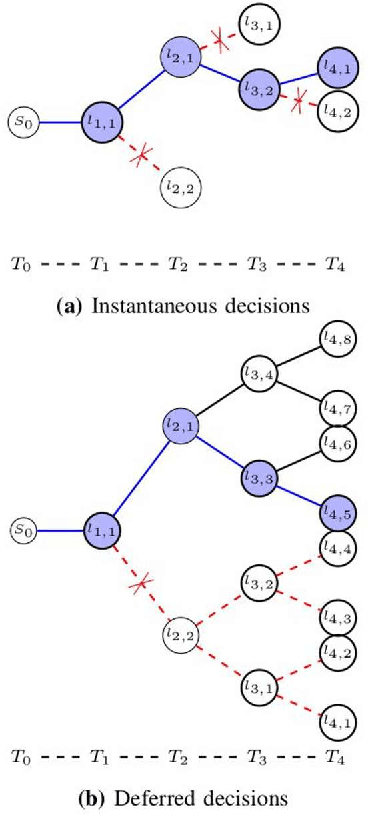
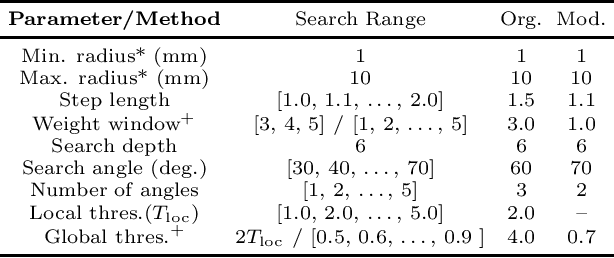

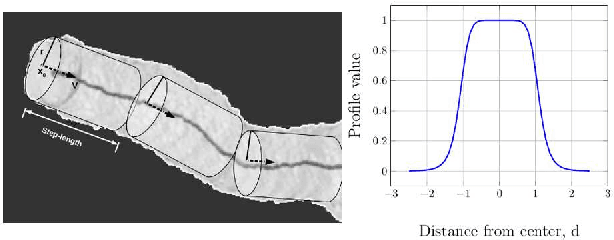
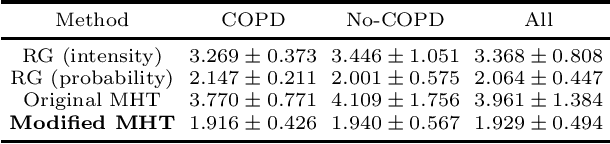
Abstract:In this work, we adapt a method based on multiple hypothesis tracking (MHT) that has been shown to give state-of-the-art vessel segmentation results in interactive settings, for the purpose of extracting trees. Regularly spaced tubular templates are fit to image data forming local hypotheses. These local hypotheses are used to construct the MHT tree, which is then traversed to make segmentation decisions. However, some critical parameters in this method are scale-dependent and have an adverse effect when tracking structures of varying dimensions. We propose to use statistical ranking of local hypotheses in constructing the MHT tree, which yields a probabilistic interpretation of scores across scales and helps alleviate the scale-dependence of MHT parameters. This enables our method to track trees starting from a single seed point. Our method is evaluated on chest CT data to extract airway trees and coronary arteries. In both cases, we show that our method performs significantly better than the original MHT method.
Feature learning based on visual similarity triplets in medical image analysis: A case study of emphysema in chest CT scans
Jun 19, 2018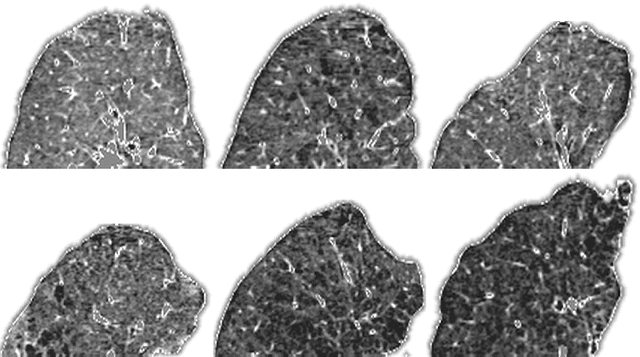
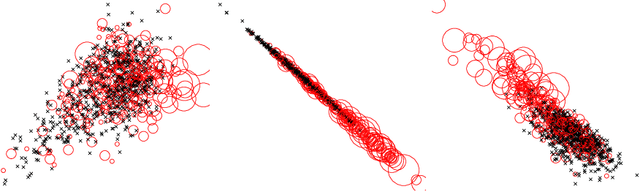



Abstract:Supervised feature learning using convolutional neural networks (CNNs) can provide concise and disease relevant representations of medical images. However, training CNNs requires annotated image data. Annotating medical images can be a time-consuming task and even expert annotations are subject to substantial inter- and intra-rater variability. Assessing visual similarity of images instead of indicating specific pathologies or estimating disease severity could allow non-experts to participate, help uncover new patterns, and possibly reduce rater variability. We consider the task of assessing emphysema extent in chest CT scans. We derive visual similarity triplets from visually assessed emphysema extent and learn a low dimensional embedding using CNNs. We evaluate the networks on 973 images, and show that the CNNs can learn disease relevant feature representations from derived similarity triplets. To our knowledge this is the first medical image application where similarity triplets has been used to learn a feature representation that can be used for embedding unseen test images
Extraction of Airways using Graph Neural Networks
Apr 12, 2018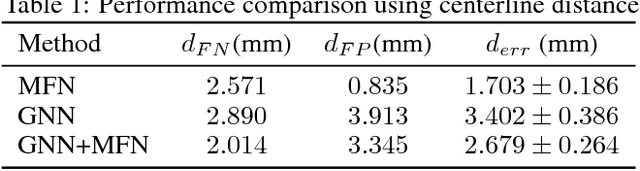
Abstract:We present extraction of tree structures, such as airways, from image data as a graph refinement task. To this end, we propose a graph auto-encoder model that uses an encoder based on graph neural networks (GNNs) to learn embeddings from input node features and a decoder to predict connections between nodes. Performance of the GNN model is compared with mean-field networks in their ability to extract airways from 3D chest CT scans.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge