Marick Laé
Scale dependant layer for self-supervised nuclei encoding
Jul 22, 2022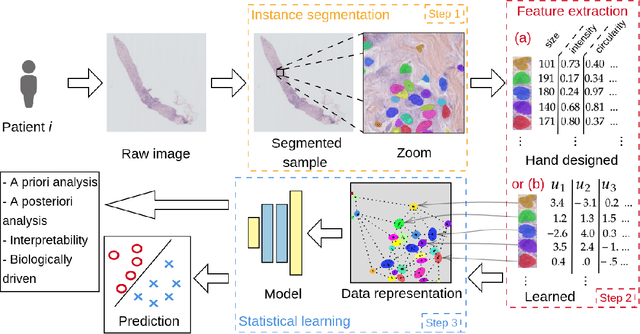
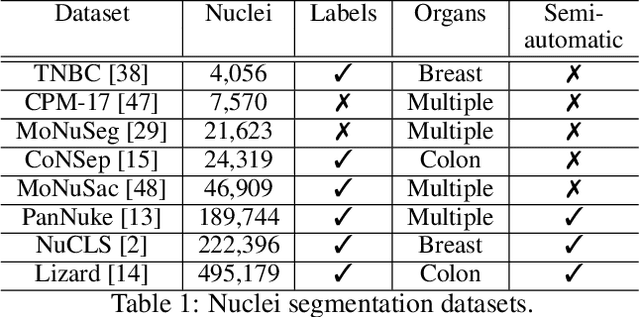
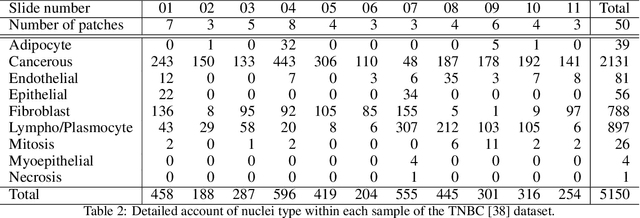
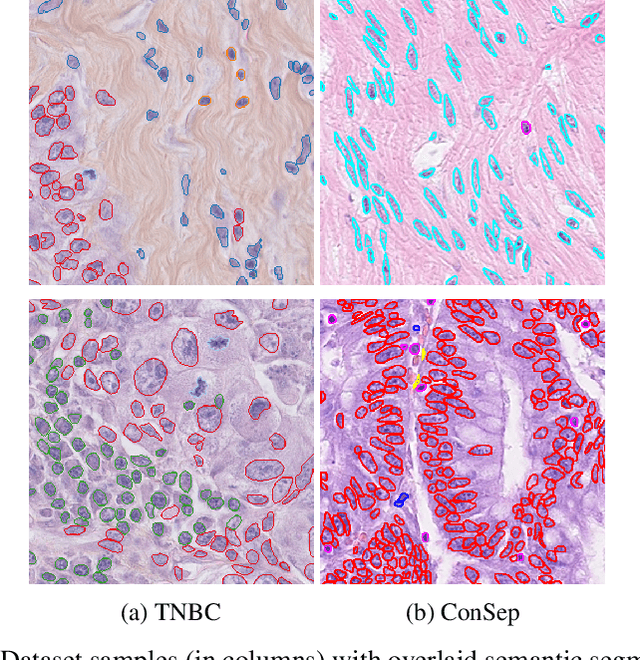
Abstract:Recent developments in self-supervised learning give us the possibility to further reduce human intervention in multi-step pipelines where the focus evolves around particular objects of interest. In the present paper, the focus lays in the nuclei in histopathology images. In particular we aim at extracting cellular information in an unsupervised manner for a downstream task. As nuclei present themselves in a variety of sizes, we propose a new Scale-dependant convolutional layer to bypass scaling issues when resizing nuclei. On three nuclei datasets, we benchmark the following methods: handcrafted, pre-trained ResNet, supervised ResNet and self-supervised features. We show that the proposed convolution layer boosts performance and that this layer combined with Barlows-Twins allows for better nuclei encoding compared to the supervised paradigm in the low sample setting and outperforms all other proposed unsupervised methods. In addition, we extend the existing TNBC dataset to incorporate nuclei class annotation in order to enrich and publicly release a small sample setting dataset for nuclei segmentation and classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge