Majid Sarrafzadeh
Target-Focused Feature Selection Using a Bayesian Approach
Sep 15, 2019
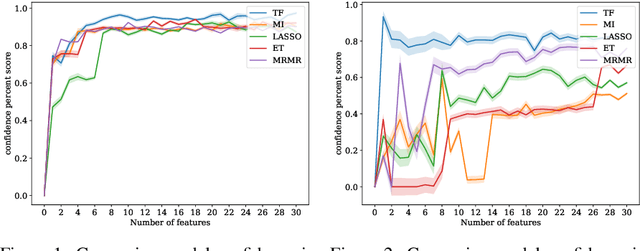
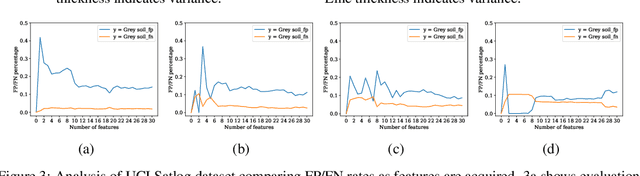
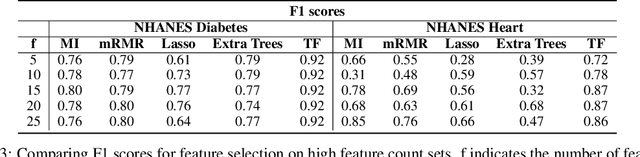
Abstract:In many real-world scenarios where data is high dimensional, test time acquisition of features is a non-trivial task due to costs associated with feature acquisition and evaluating feature value. The need for highly confident models with an extremely frugal acquisition of features can be addressed by allowing a feature selection method to become target aware. We introduce an approach to feature selection that is based on Bayesian learning, allowing us to report target-specific levels of uncertainty, false positive, and false negative rates. In addition, measuring uncertainty lifts the restriction on feature selection being target agnostic, allowing for feature acquisition based on a single target of focus out of many. We show that acquiring features for a specific target is at least as good as common linear feature selection approaches for small non-sparse datasets, and surpasses these when faced with real-world healthcare data that is larger in scale and in sparseness.
TAPER: Time-Aware Patient EHR Representation
Aug 16, 2019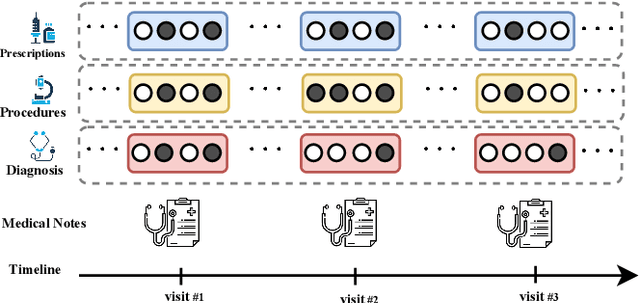
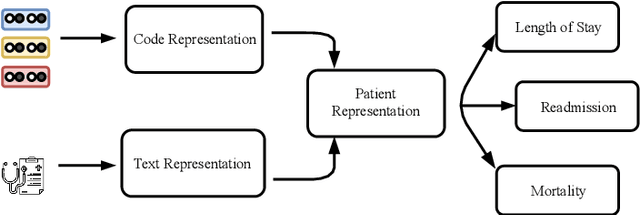
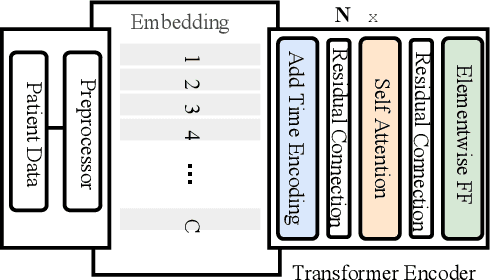
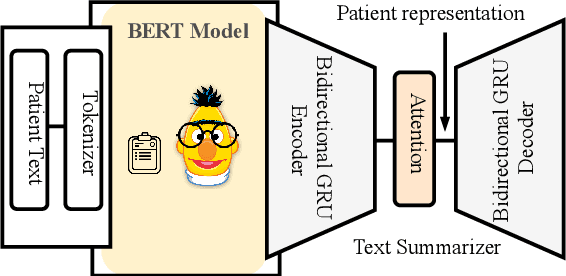
Abstract:Effective representation learning of electronic health records is a challenging task and is becoming more important as the availability of such data is becoming pervasive. The data contained in these records are irregular and contain multiple modalities such as notes, and medical codes. They are preempted by medical conditions the patient may have, and are typically jotted down by medical staff. Accompanying codes are notes containing valuable information about patients beyond the structured information contained in electronic health records. We use transformer networks and the recently proposed BERT language model to embed these data streams into a unified vector representation. The presented approach effectively encodes a patient's visit data into a single distributed representation, which can be used for downstream tasks. Our model demonstrates superior performance and generalization on mortality, readmission and length of stay tasks using the publicly available MIMIC-III ICU dataset.
Generative Imputation and Stochastic Prediction
May 22, 2019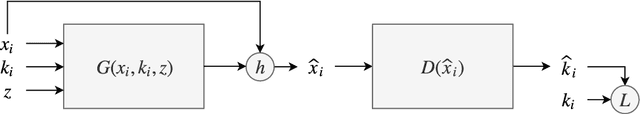
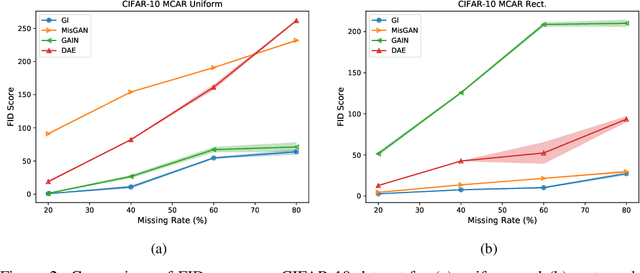
Abstract:In many machine learning applications, we are faced with incomplete datasets. In the literature, missing data imputation techniques have been mostly concerned with filling missing values. However, the existence of missing values is synonymous with uncertainties not only over the distribution of missing values but also over target class assignments that require careful consideration. The objectives of this paper are twofold. First, we proposed a method for generating imputations from the conditional distribution of missing values given observed values. Second, we use the generated samples to estimate the distribution of target assignments given incomplete data. In order to generate imputations, we train a simple and effective generator network to generate imputations that a discriminator network is tasked to distinguish. Following this, a predictor network is trained using imputed samples from the generator network to capture the classification uncertainties and make predictions accordingly. The proposed method is evaluated on CIFAR-10 image dataset as well as two real-world tabular classification datasets, under various missingness rates and structures. Our experimental results show the effectiveness of the proposed method in generating imputations, as well as providing estimates for the class uncertainties in a classification task when faced with missing values.
Nutrition and Health Data for Cost-Sensitive Learning
Feb 19, 2019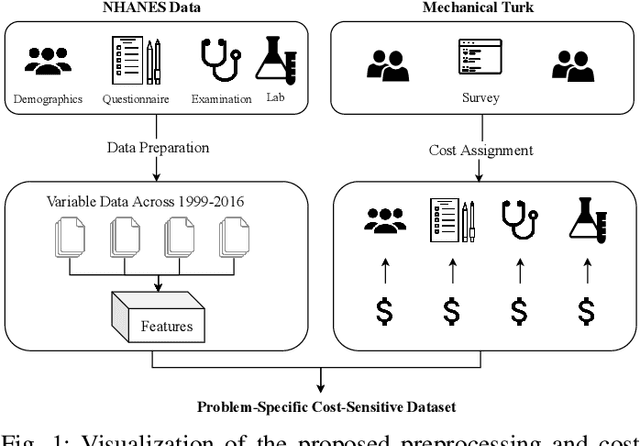
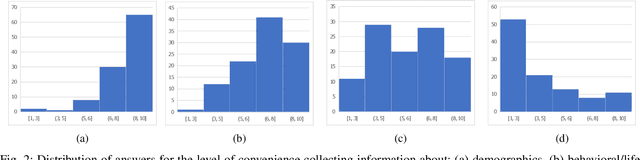
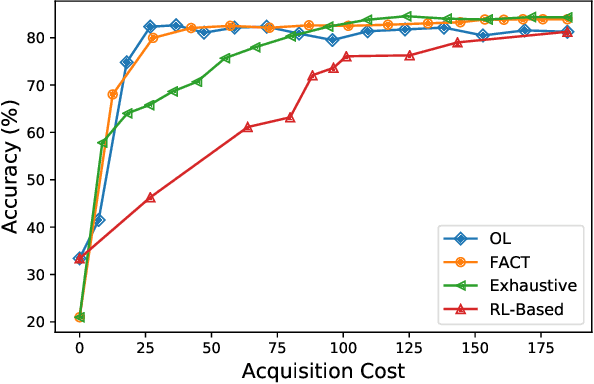
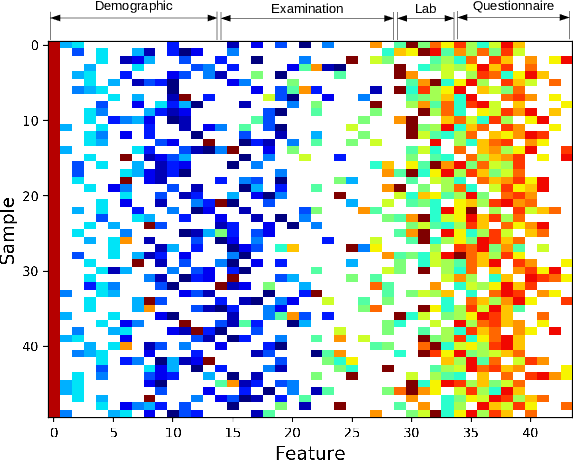
Abstract:Traditionally, machine learning algorithms have been focused on modeling dynamics of a certain dataset at hand for which all features are available for free. However, there are many concerns such as monetary data collection costs, patient discomfort in medical procedures, and privacy impacts of data collection that require careful consideration in any health analytics system. An efficient solution would only acquire a subset of features based on the value it provides whilst considering acquisition costs. Moreover, datasets that provide feature costs are very limited, especially in healthcare. In this paper, we provide a health dataset as well as a method for assigning feature costs based on the total level of inconvenience asking for each feature entails. Furthermore, based on the suggested dataset, we provide a comparison of recent and state-of-the-art approaches to cost-sensitive feature acquisition and learning. Specifically, we analyze the performance of major sensitivity-based and reinforcement learning based methods in the literature on three different problems in the health domain, including diabetes, heart disease, and hypertension classification.
Unsupervised Prediction of Negative Health Events Ahead of Time
Jan 31, 2019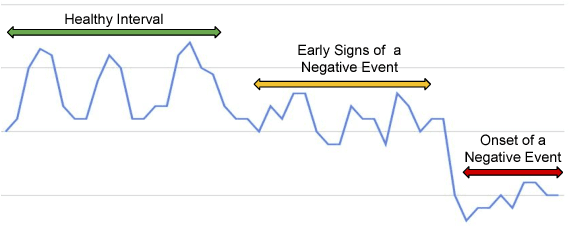
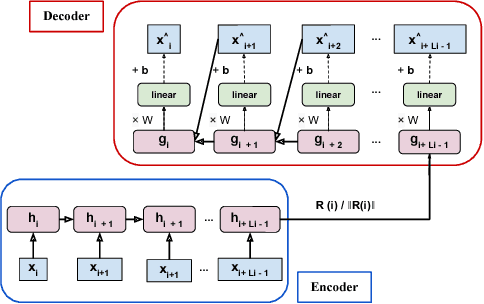
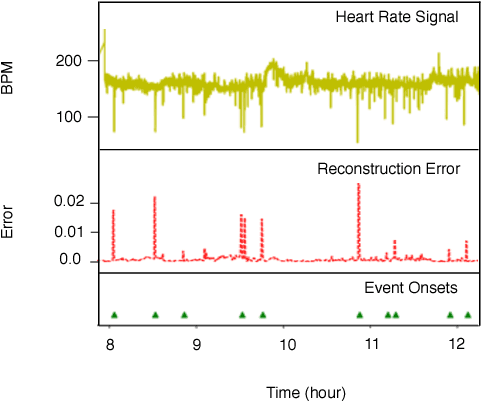
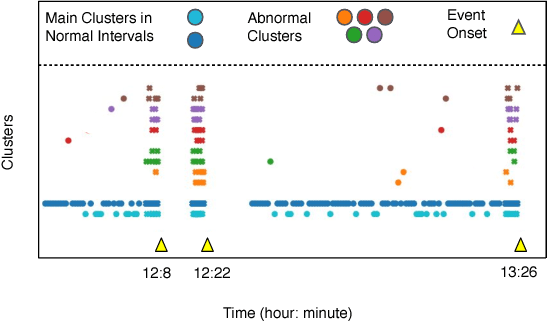
Abstract:The emergence of continuous health monitoring and the availability of an enormous amount of time series data has provided a great opportunity for the advancement of personal health tracking. In recent years, unsupervised learning methods have drawn special attention of researchers to tackle the sparse annotation of health data and real-time detection of anomalies has been a central problem of interest. However, one problem that has not been well addressed before is the early prediction of forthcoming negative health events. Early signs of an event can introduce subtle and gradual changes in the health signal prior to its onset, detection of which can be invaluable in effective prevention. In this study, we first demonstrate our observations on the shortcoming of widely adopted anomaly detection methods in uncovering the changes prior to a negative health event. We then propose a framework which relies on online clustering of signal segment representations which are automatically learned by a specially designed LSTM auto-encoder. We show the effectiveness of our approach by predicting Bradycardia events in infants using MIT-PICS dataset 1.3 minutes ahead of time with 68\% AUC score on average, using no label supervision. Results of our study can indicate the viability of our approach in the early detection of health events in other applications as well.
Opportunistic Learning: Budgeted Cost-Sensitive Learning from Data Streams
Jan 02, 2019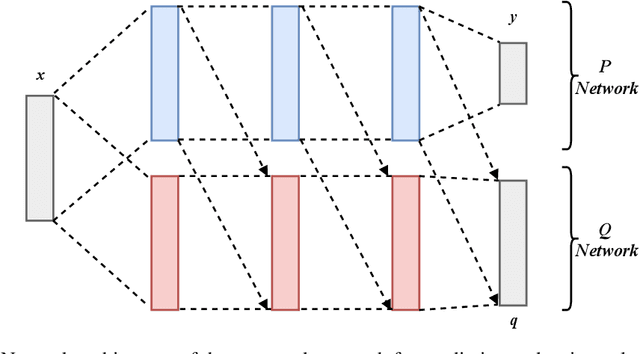
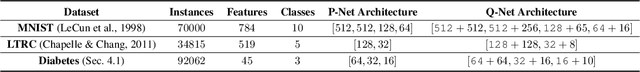
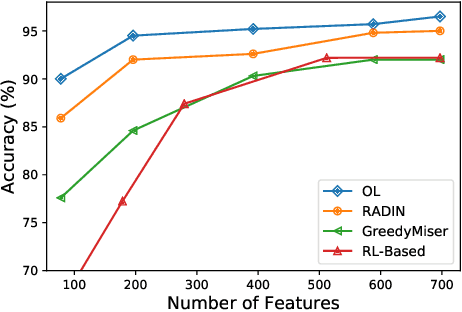
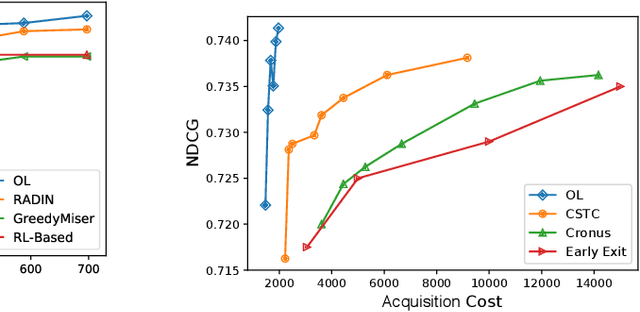
Abstract:In many real-world learning scenarios, features are only acquirable at a cost constrained under a budget. In this paper, we propose a novel approach for cost-sensitive feature acquisition at the prediction-time. The suggested method acquires features incrementally based on a context-aware feature-value function. We formulate the problem in the reinforcement learning paradigm, and introduce a reward function based on the utility of each feature. Specifically, MC dropout sampling is used to measure expected variations of the model uncertainty which is used as a feature-value function. Furthermore, we suggest sharing representations between the class predictor and value function estimator networks. The suggested approach is completely online and is readily applicable to stream learning setups. The solution is evaluated on three different datasets including the well-known MNIST dataset as a benchmark as well as two cost-sensitive datasets: Yahoo Learning to Rank and a dataset in the medical domain for diabetes classification. According to the results, the proposed method is able to efficiently acquire features and make accurate predictions.
* https://openreview.net/forum?id=S1eOHo09KX
Dynamic Feature Acquisition Using Denoising Autoencoders
Nov 03, 2018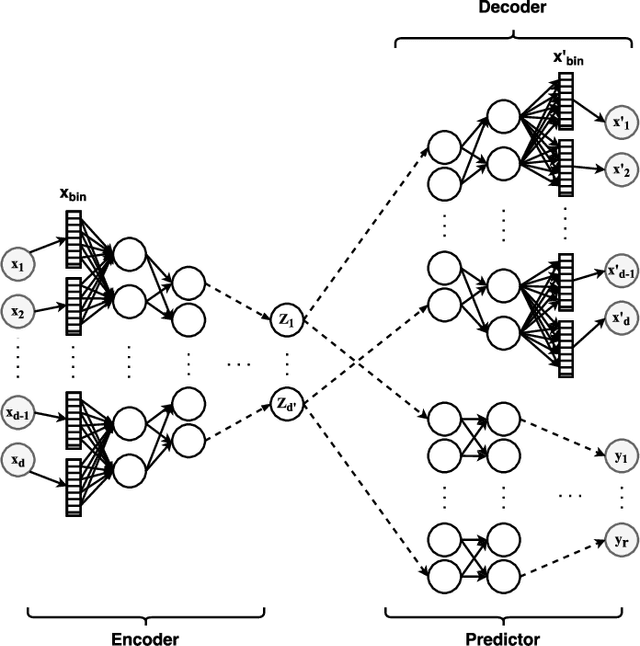
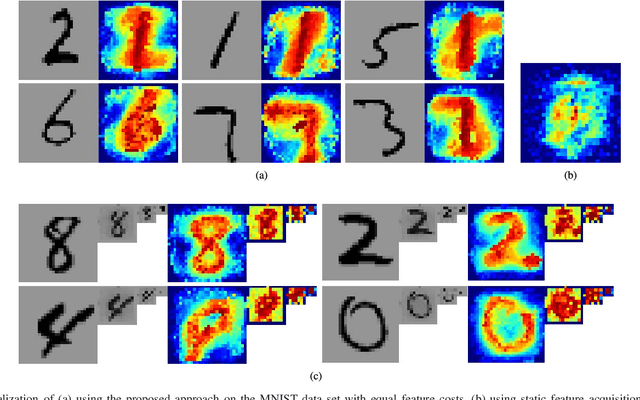
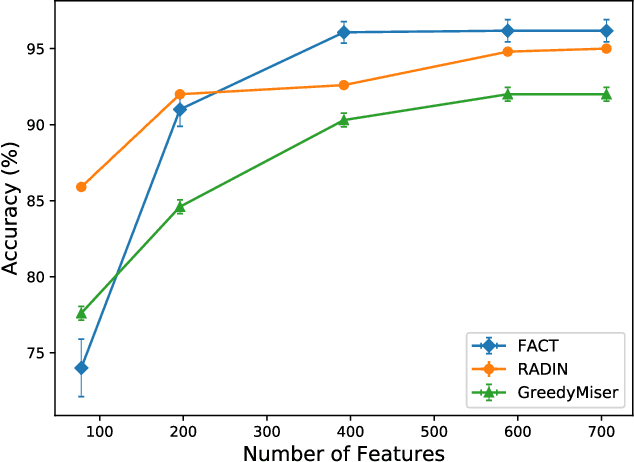
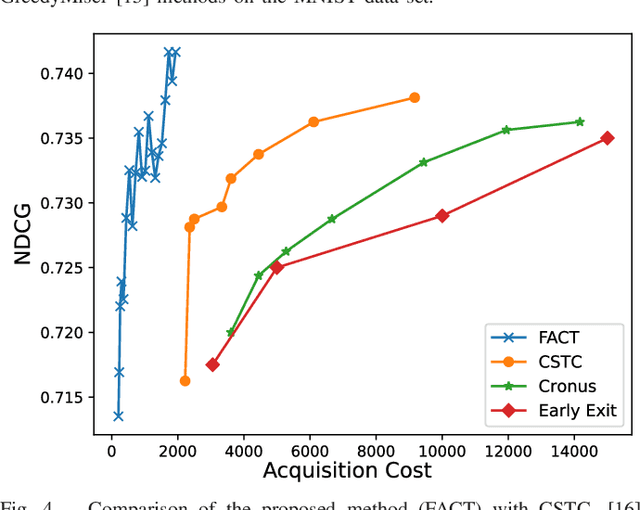
Abstract:In real-world scenarios, different features have different acquisition costs at test-time which necessitates cost-aware methods to optimize the cost and performance trade-off. This paper introduces a novel and scalable approach for cost-aware feature acquisition at test-time. The method incrementally asks for features based on the available context that are known feature values. The proposed method is based on sensitivity analysis in neural networks and density estimation using denoising autoencoders with binary representation layers. In the proposed architecture, a denoising autoencoder is used to handle unknown features (i.e., features that are yet to be acquired), and the sensitivity of predictions with respect to each unknown feature is used as a context-dependent measure of informativeness. We evaluated the proposed method on eight different real-world datasets as well as one synthesized dataset and compared its performance with several other approaches in the literature. According to the results, the suggested method is capable of efficiently acquiring features at test-time in a cost- and context-aware fashion.
ECG Heartbeat Classification: A Deep Transferable Representation
Jul 12, 2018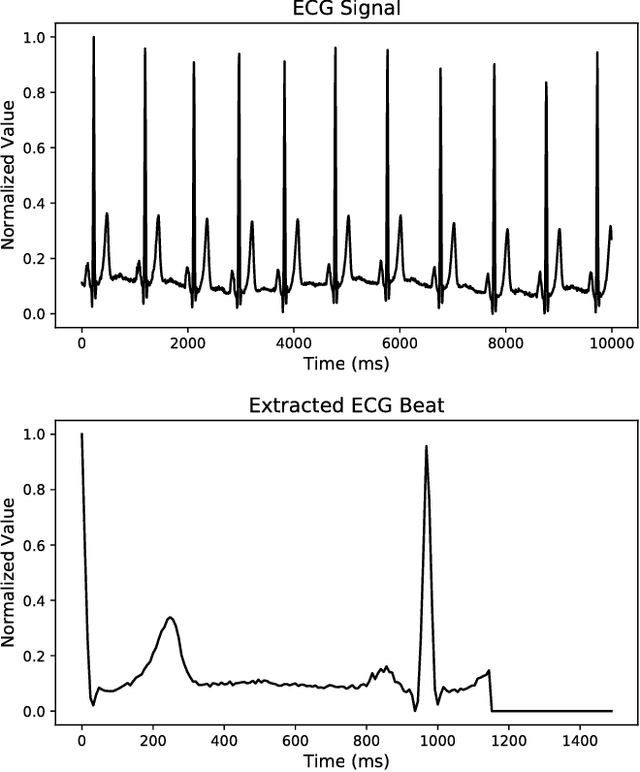
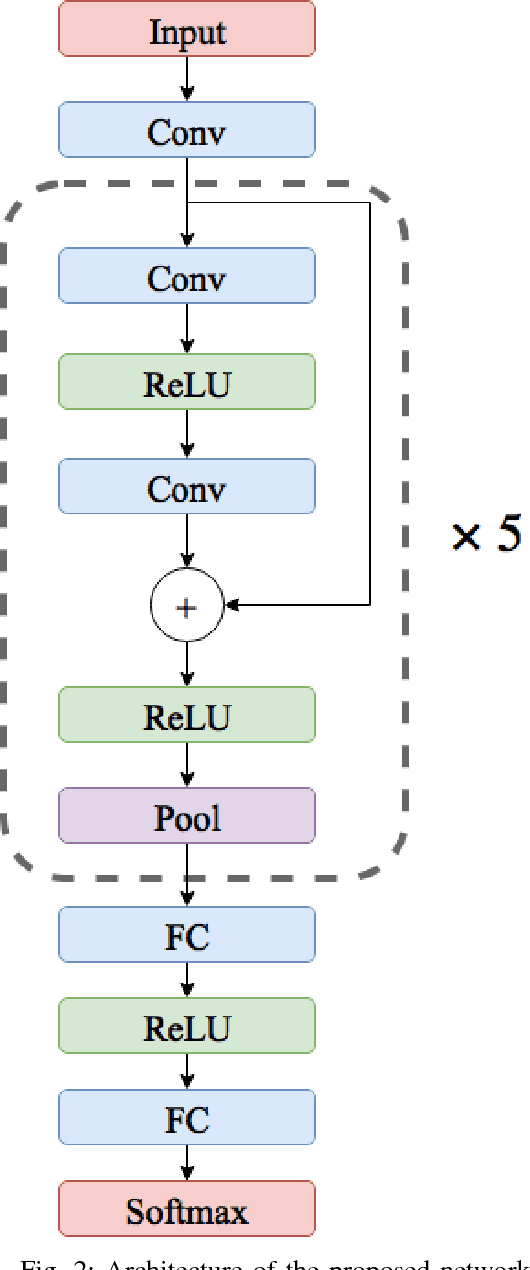
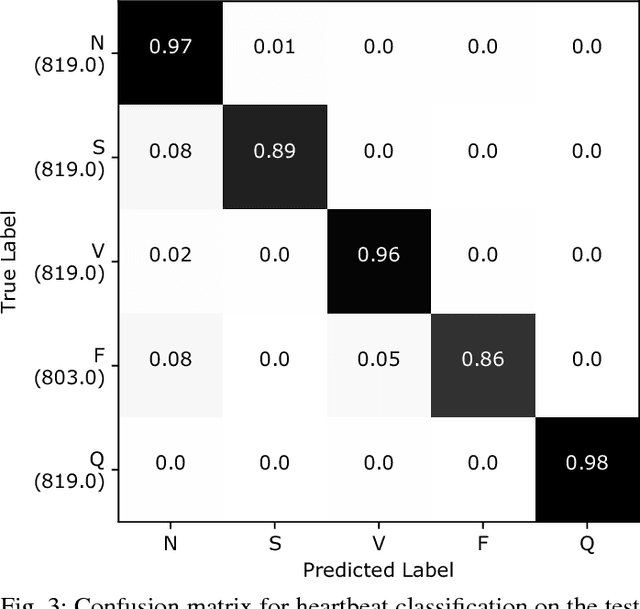
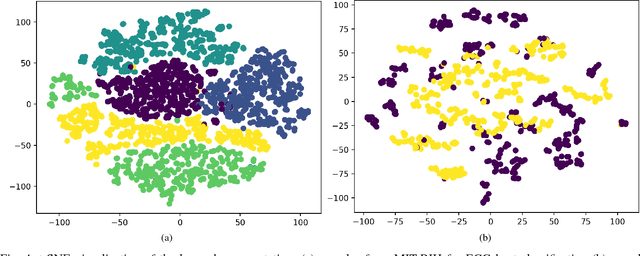
Abstract:Electrocardiogram (ECG) can be reliably used as a measure to monitor the functionality of the cardiovascular system. Recently, there has been a great attention towards accurate categorization of heartbeats. While there are many commonalities between different ECG conditions, the focus of most studies has been classifying a set of conditions on a dataset annotated for that task rather than learning and employing a transferable knowledge between different tasks. In this paper, we propose a method based on deep convolutional neural networks for the classification of heartbeats which is able to accurately classify five different arrhythmias in accordance with the AAMI EC57 standard. Furthermore, we suggest a method for transferring the knowledge acquired on this task to the myocardial infarction (MI) classification task. We evaluated the proposed method on PhysionNet's MIT-BIH and PTB Diagnostics datasets. According to the results, the suggested method is able to make predictions with the average accuracies of 93.4% and 95.9% on arrhythmia classification and MI classification, respectively.
HeteroMed: Heterogeneous Information Network for Medical Diagnosis
Apr 22, 2018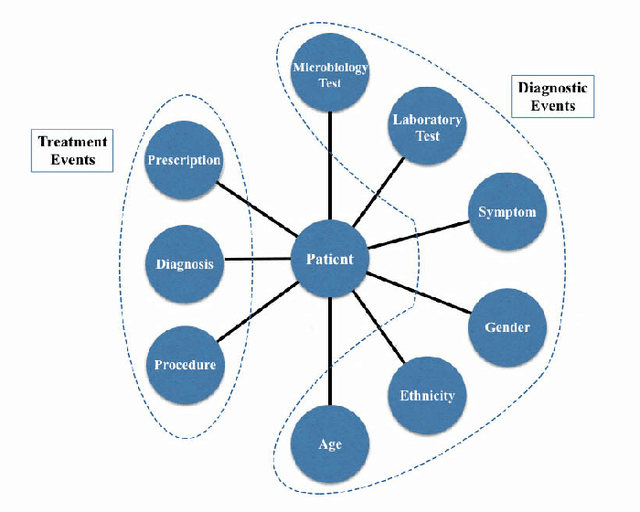
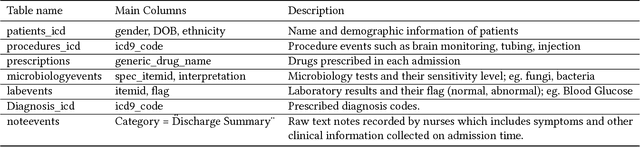

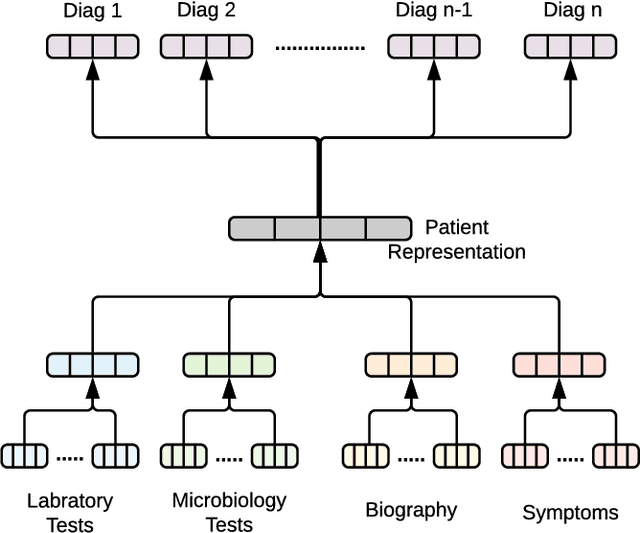
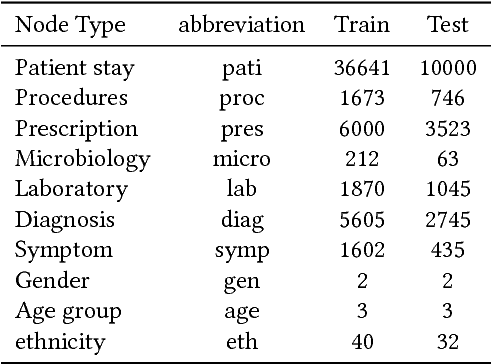
Abstract:With the recent availability of Electronic Health Records (EHR) and great opportunities they offer for advancing medical informatics, there has been growing interest in mining EHR for improving quality of care. Disease diagnosis due to its sensitive nature, huge costs of error, and complexity has become an increasingly important focus of research in past years. Existing studies model EHR by capturing co-occurrence of clinical events to learn their latent embeddings. However, relations among clinical events carry various semantics and contribute differently to disease diagnosis which gives precedence to a more advanced modeling of heterogeneous data types and relations in EHR data than existing solutions. To address these issues, we represent how high-dimensional EHR data and its rich relationships can be suitably translated into HeteroMed, a heterogeneous information network for robust medical diagnosis. Our modeling approach allows for straightforward handling of missing values and heterogeneity of data. HeteroMed exploits metapaths to capture higher level and semantically important relations contributing to disease diagnosis. Furthermore, it employs a joint embedding framework to tailor clinical event representations to the disease diagnosis goal. To the best of our knowledge, this is the first study to use Heterogeneous Information Network for modeling clinical data and disease diagnosis. Experimental results of our study show superior performance of HeteroMed compared to prior methods in prediction of exact diagnosis codes and general disease cohorts. Moreover, HeteroMed outperforms baseline models in capturing similarities of clinical events which are examined qualitatively through case studies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge