Luca Muscarnera
TheProphetAI
Elimination-compensation pruning for fully-connected neural networks
Feb 24, 2026Abstract:The unmatched ability of Deep Neural Networks in capturing complex patterns in large and noisy datasets is often associated with their large hypothesis space, and consequently to the vast amount of parameters that characterize model architectures. Pruning techniques affirmed themselves as valid tools to extract sparse representations of neural networks parameters, carefully balancing between compression and preservation of information. However, a fundamental assumption behind pruning is that expendable weights should have small impact on the error of the network, while highly important weights should tend to have a larger influence on the inference. We argue that this idea could be generalized; what if a weight is not simply removed but also compensated with a perturbation of the adjacent bias, which does not contribute to the network sparsity? Our work introduces a novel pruning method in which the importance measure of each weight is computed considering the output behavior after an optimal perturbation of its adjacent bias, efficiently computable by automatic differentiation. These perturbations can be then applied directly after the removal of each weight, independently of each other. After deriving analytical expressions for the aforementioned quantities, numerical experiments are conducted to benchmark this technique against some of the most popular pruning strategies, demonstrating an intrinsic efficiency of the proposed approach in very diverse machine learning scenarios. Finally, our findings are discussed and the theoretical implications of our results are presented.
Knowledge-Informed Kernel State Reconstruction for Interpretable Dynamical System Discovery
Jan 29, 2026Abstract:Recovering governing equations from data is central to scientific discovery, yet existing methods often break down under noisy, partial observations, or rely on black-box latent dynamics that obscure mechanism. We introduce MAAT (Model Aware Approximation of Trajectories), a framework for symbolic discovery built on knowledge-informed Kernel State Reconstruction. MAAT formulates state reconstruction in a reproducing kernel Hilbert space and directly incorporates structural and semantic priors such as non-negativity, conservation laws, and domain-specific observation models into the reconstruction objective, while accommodating heterogeneous sampling and measurement granularity. This yields smooth, physically consistent state estimates with analytic time derivatives, providing a principled interface between fragmented sensor data and symbolic regression. Across twelve diverse scientific benchmarks and multiple noise regimes, MAAT substantially reduces state-estimation MSE for trajectories and derivatives used by downstream symbolic regression relative to strong baselines.
Emergence of Structure in Ensembles of Random Neural Networks
May 15, 2025



Abstract:Randomness is ubiquitous in many applications across data science and machine learning. Remarkably, systems composed of random components often display emergent global behaviors that appear deterministic, manifesting a transition from microscopic disorder to macroscopic organization. In this work, we introduce a theoretical model for studying the emergence of collective behaviors in ensembles of random classifiers. We argue that, if the ensemble is weighted through the Gibbs measure defined by adopting the classification loss as an energy, then there exists a finite temperature parameter for the distribution such that the classification is optimal, with respect to the loss (or the energy). Interestingly, for the case in which samples are generated by a Gaussian distribution and labels are constructed by employing a teacher perceptron, we analytically prove and numerically confirm that such optimal temperature does not depend neither on the teacher classifier (which is, by construction of the learning problem, unknown), nor on the number of random classifiers, highlighting the universal nature of the observed behavior. Experiments on the MNIST dataset underline the relevance of this phenomenon in high-quality, noiseless, datasets. Finally, a physical analogy allows us to shed light on the self-organizing nature of the studied phenomenon.
DeepProphet2 -- A Deep Learning Gene Recommendation Engine
Aug 03, 2022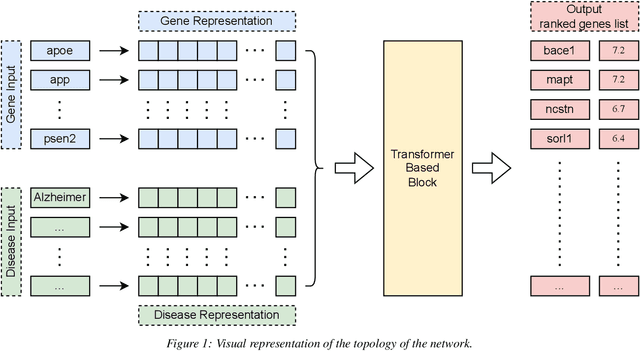
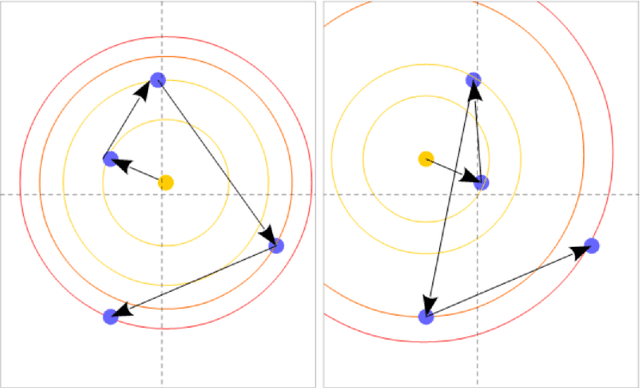
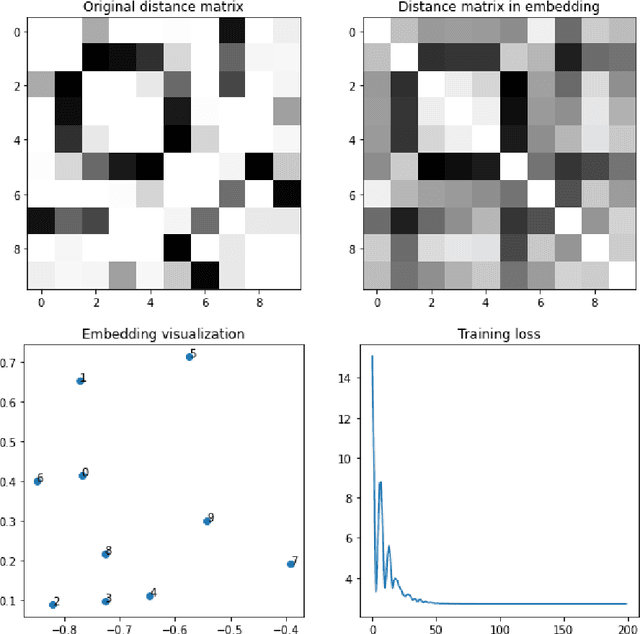
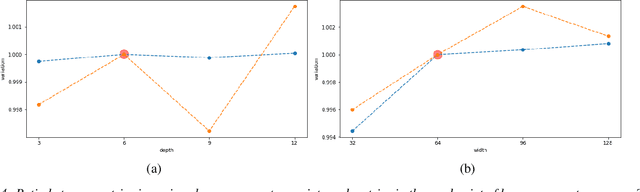
Abstract:New powerful tools for tackling life science problems have been created by recent advances in machine learning. The purpose of the paper is to discuss the potential advantages of gene recommendation performed by artificial intelligence (AI). Indeed, gene recommendation engines try to solve this problem: if the user is interested in a set of genes, which other genes are likely to be related to the starting set and should be investigated? This task was solved with a custom deep learning recommendation engine, DeepProphet2 (DP2), which is freely available to researchers worldwide via www.generecommender.com. Hereafter, insights behind the algorithm and its practical applications are illustrated. The gene recommendation problem can be addressed by mapping the genes to a metric space where a distance can be defined to represent the real semantic distance between them. To achieve this objective a transformer-based model has been trained on a well-curated freely available paper corpus, PubMed. The paper describes multiple optimization procedures that were employed to obtain the best bias-variance trade-off, focusing on embedding size and network depth. In this context, the model's ability to discover sets of genes implicated in diseases and pathways was assessed through cross-validation. A simple assumption guided the procedure: the network had no direct knowledge of pathways and diseases but learned genes' similarities and the interactions among them. Moreover, to further investigate the space where the neural network represents genes, the dimensionality of the embedding was reduced, and the results were projected onto a human-comprehensible space. In conclusion, a set of use cases illustrates the algorithm's potential applications in a real word setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge