Lorenzo Cerrone
SynCellFactory: Generative Data Augmentation for Cell Tracking
Apr 26, 2024Abstract:Cell tracking remains a pivotal yet challenging task in biomedical research. The full potential of deep learning for this purpose is often untapped due to the limited availability of comprehensive and varied training data sets. In this paper, we present SynCellFactory, a generative cell video augmentation. At the heart of SynCellFactory lies the ControlNet architecture, which has been fine-tuned to synthesize cell imagery with photorealistic accuracy in style and motion patterns. This technique enables the creation of synthetic yet realistic cell videos that mirror the complexity of authentic microscopy time-lapses. Our experiments demonstrate that SynCellFactory boosts the performance of well-established deep learning models for cell tracking, particularly when original training data is sparse.
CellTypeGraph: A New Geometric Computer Vision Benchmark
May 17, 2022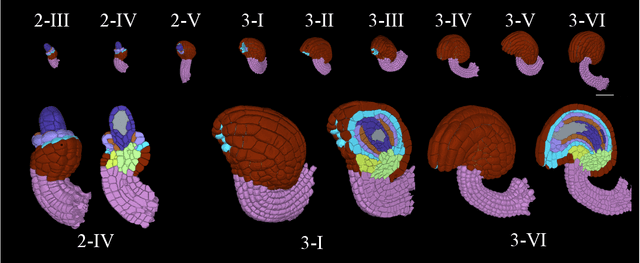
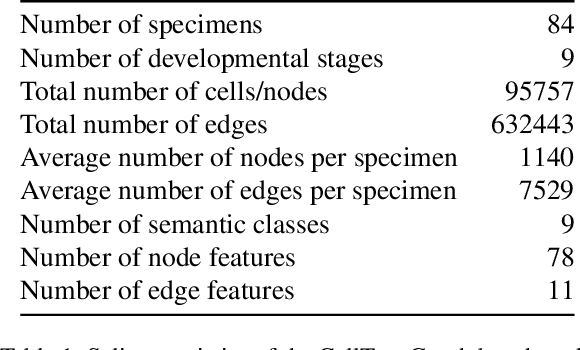

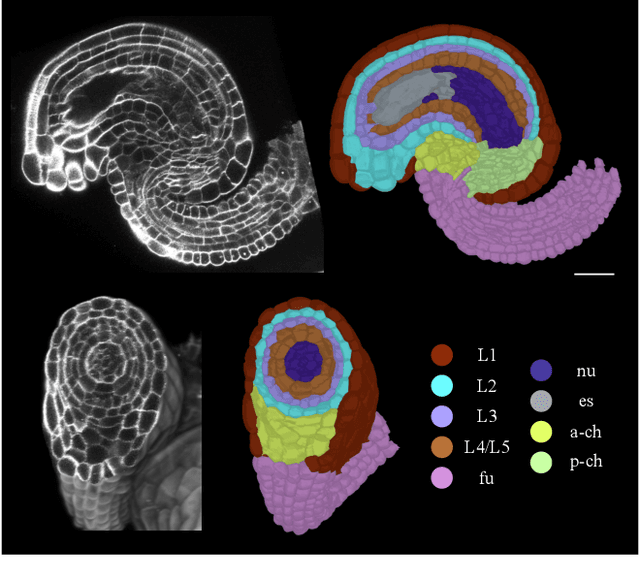
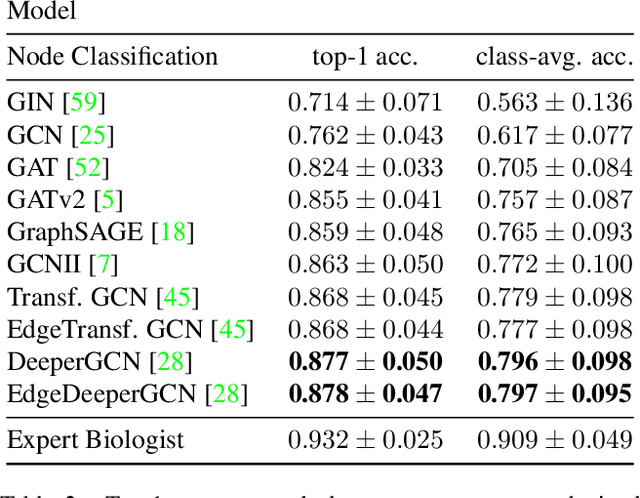
Abstract:Classifying all cells in an organ is a relevant and difficult problem from plant developmental biology. We here abstract the problem into a new benchmark for node classification in a geo-referenced graph. Solving it requires learning the spatial layout of the organ including symmetries. To allow the convenient testing of new geometrical learning methods, the benchmark of Arabidopsis thaliana ovules is made available as a PyTorch data loader, along with a large number of precomputed features. Finally, we benchmark eight recent graph neural network architectures, finding that DeeperGCN currently works best on this problem.
End-to-End Learned Random Walker for Seeded Image Segmentation
May 22, 2019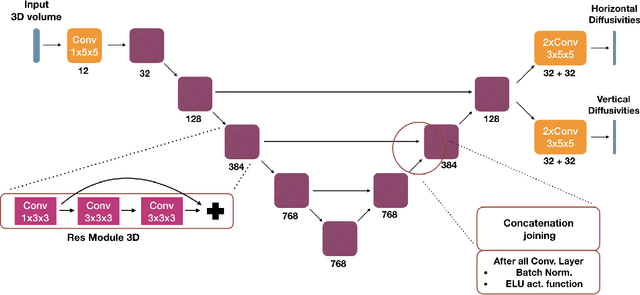
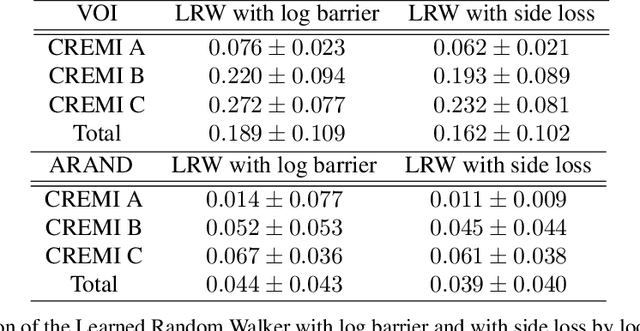

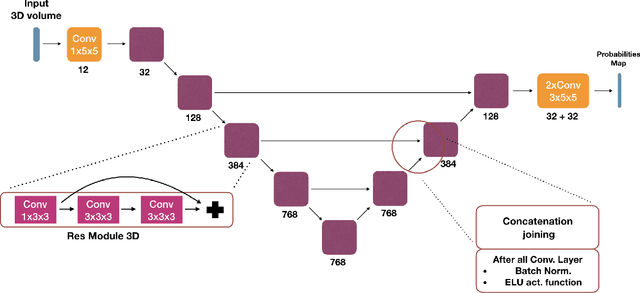
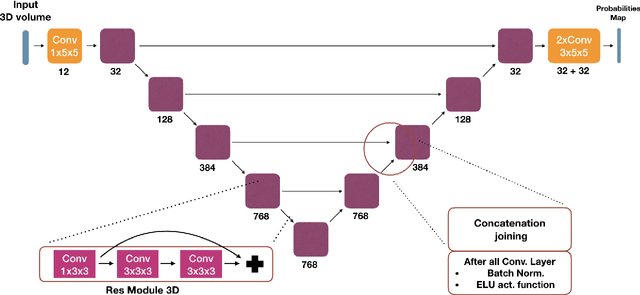
Abstract:We present an end-to-end learned algorithm for seeded segmentation. Our method is based on the Random Walker algorithm, where we predict the edge weights of the underlying graph using a convolutional neural network. This can be interpreted as learning context-dependent diffusivities for a linear diffusion process. Besides calculating the exact gradient for optimizing these diffusivities, we also propose simplifications that sparsely sample the gradient and still yield competitive results. The proposed method achieves the currently best results on a seeded version of the CREMI neuron segmentation challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge