Kyung-Ah Sohn
Image Distortion Detection using Convolutional Neural Network
May 28, 2018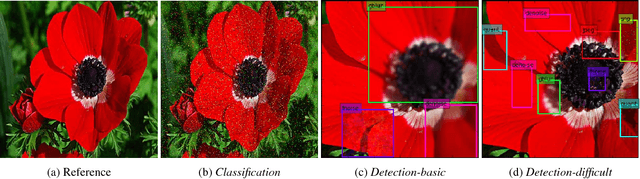
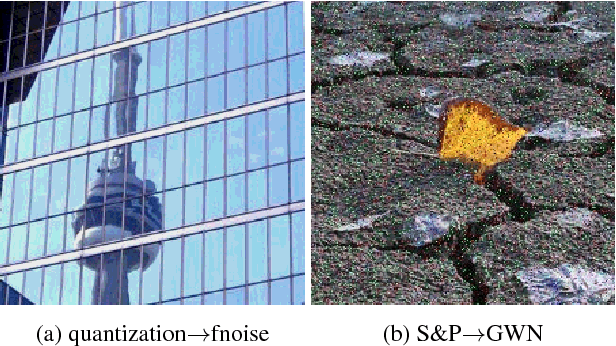
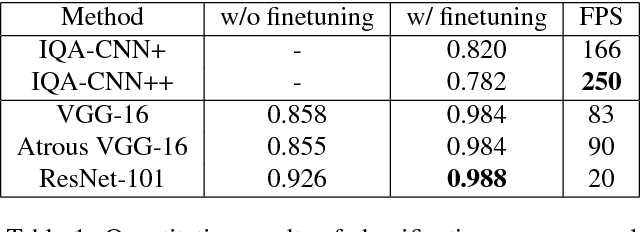

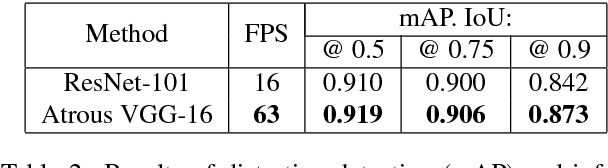
Abstract:Image distortion classification and detection is an important task in many applications. For example when compressing images, if we know the exact location of the distortion, then it is possible to re-compress images by adjusting the local compression level dynamically. In this paper, we address the problem of detecting the distortion region and classifying the distortion type of a given image. We show that our model significantly outperforms the state-of-the-art distortion classifier, and report accurate detection results for the first time. We expect that such results prove the usefulness of our approach in many potential applications such as image compression or distortion restoration.
Investigating the feature collection for semantic segmentation via single skip connection
Oct 23, 2017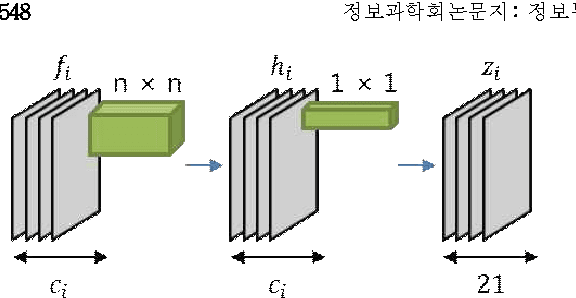
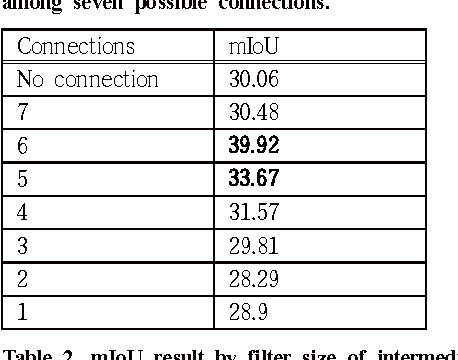
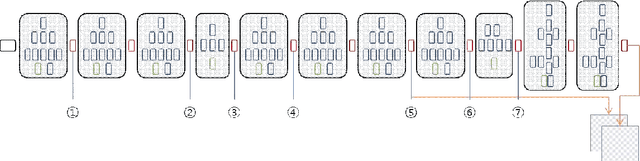
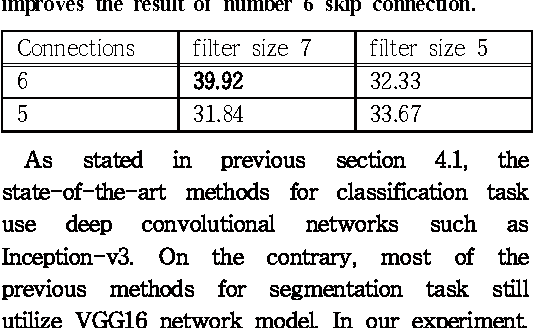
Abstract:Since the study of deep convolutional neural network became prevalent, one of the important discoveries is that a feature map from a convolutional network can be extracted before going into the fully connected layer and can be used as a saliency map for object detection. Furthermore, the model can use features from each different layer for accurate object detection: the features from different layers can have different properties. As the model goes deeper, it has many latent skip connections and feature maps to elaborate object detection. Although there are many intermediate layers that we can use for semantic segmentation through skip connection, still the characteristics of each skip connection and the best skip connection for this task are uncertain. Therefore, in this study, we exhaustively research skip connections of state-of-the-art deep convolutional networks and investigate the characteristics of the features from each intermediate layer. In addition, this study would suggest how to use a recent deep neural network model for semantic segmentation and it would therefore become a cornerstone for later studies with the state-of-the-art network models.
Enhancing the Performance of Convolutional Neural Networks on Quality Degraded Datasets
Oct 18, 2017
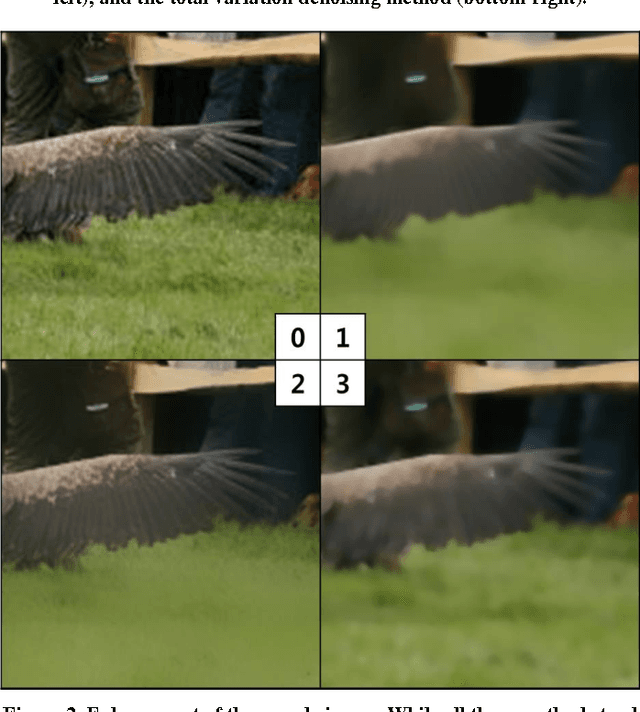
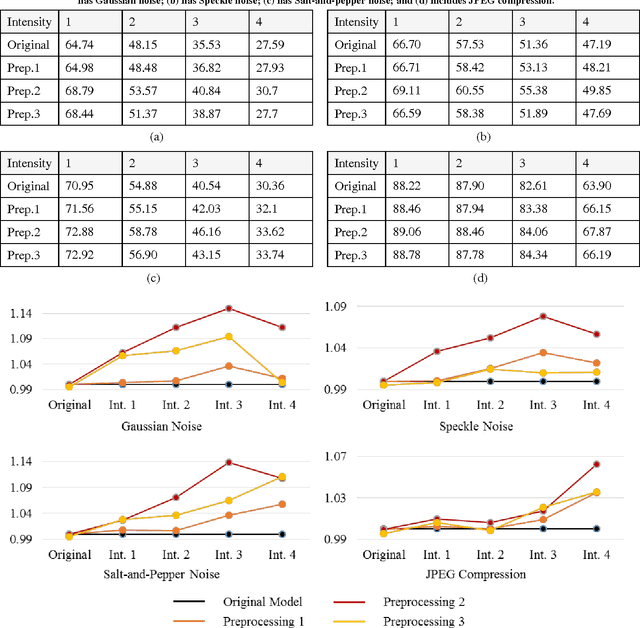

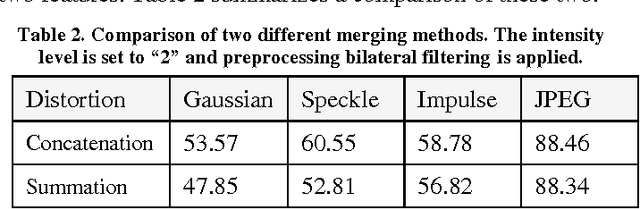
Abstract:Despite the appeal of deep neural networks that largely replace the traditional handmade filters, they still suffer from isolated cases that cannot be properly handled only by the training of convolutional filters. Abnormal factors, including real-world noise, blur, or other quality degradations, ruin the output of a neural network. These unexpected problems can produce critical complications, and it is surprising that there has only been minimal research into the effects of noise in the deep neural network model. Therefore, we present an exhaustive investigation into the effect of noise in image classification and suggest a generalized architecture of a dual-channel model to treat quality degraded input images. We compare the proposed dual-channel model with a simple single model and show it improves the overall performance of neural networks on various types of quality degraded input datasets.
A hierarchical Dirichlet process mixture model for haplotype reconstruction from multi-population data
Aug 20, 2009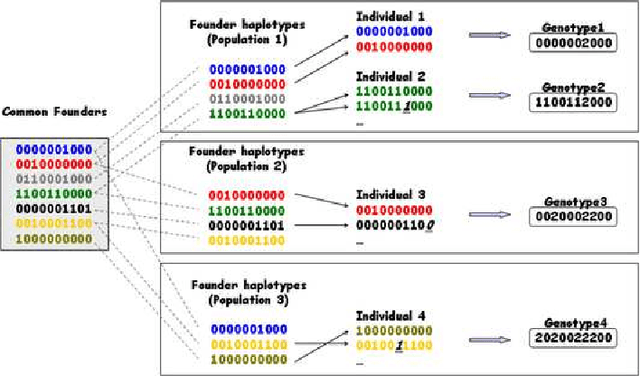
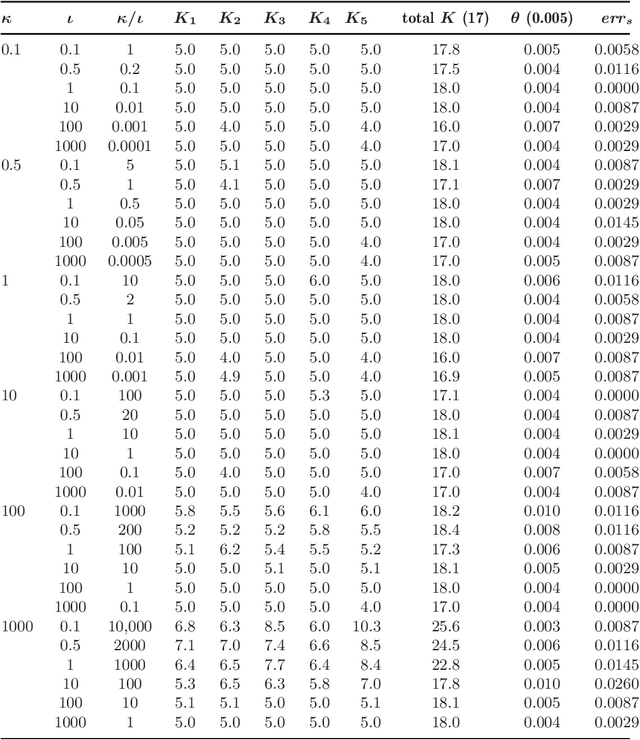
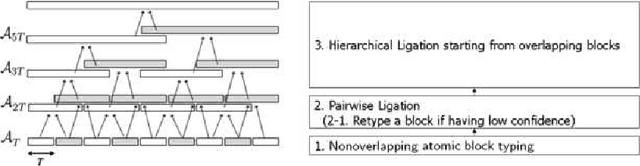
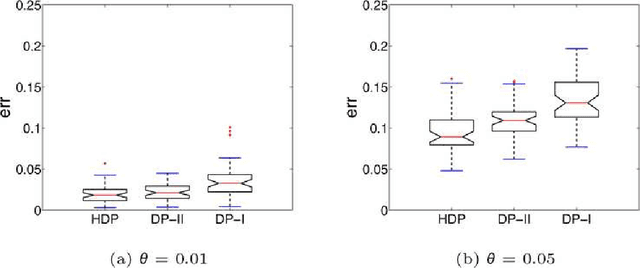
Abstract:The perennial problem of "how many clusters?" remains an issue of substantial interest in data mining and machine learning communities, and becomes particularly salient in large data sets such as populational genomic data where the number of clusters needs to be relatively large and open-ended. This problem gets further complicated in a co-clustering scenario in which one needs to solve multiple clustering problems simultaneously because of the presence of common centroids (e.g., ancestors) shared by clusters (e.g., possible descents from a certain ancestor) from different multiple-cluster samples (e.g., different human subpopulations). In this paper we present a hierarchical nonparametric Bayesian model to address this problem in the context of multi-population haplotype inference. Uncovering the haplotypes of single nucleotide polymorphisms is essential for many biological and medical applications. While it is uncommon for the genotype data to be pooled from multiple ethnically distinct populations, few existing programs have explicitly leveraged the individual ethnic information for haplotype inference. In this paper we present a new haplotype inference program, Haploi, which makes use of such information and is readily applicable to genotype sequences with thousands of SNPs from heterogeneous populations, with competent and sometimes superior speed and accuracy comparing to the state-of-the-art programs. Underlying Haploi is a new haplotype distribution model based on a nonparametric Bayesian formalism known as the hierarchical Dirichlet process, which represents a tractable surrogate to the coalescent process. The proposed model is exchangeable, unbounded, and capable of coupling demographic information of different populations.
* Published in at http://dx.doi.org/10.1214/08-AOAS225 the Annals of Applied Statistics (http://www.imstat.org/aoas/) by the Institute of Mathematical Statistics (http://www.imstat.org)
A Multivariate Regression Approach to Association Analysis of Quantitative Trait Network
Nov 13, 2008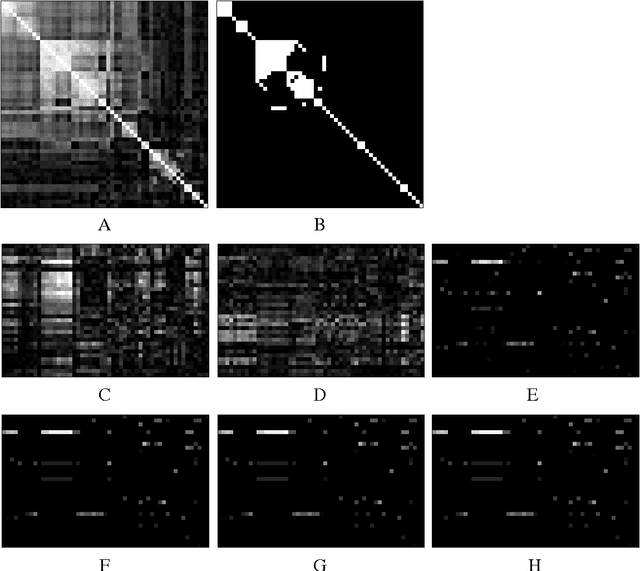
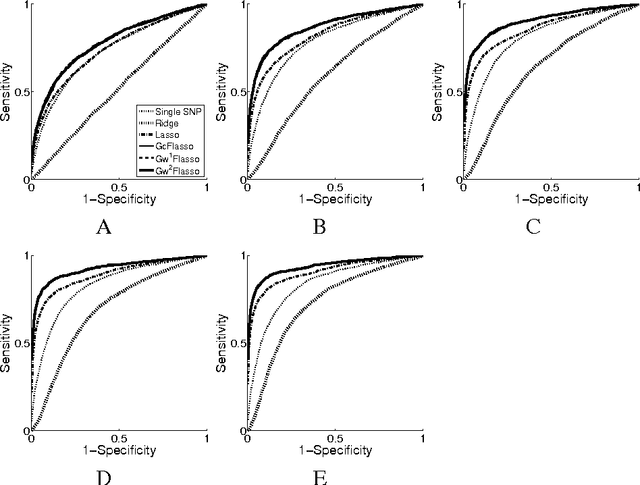
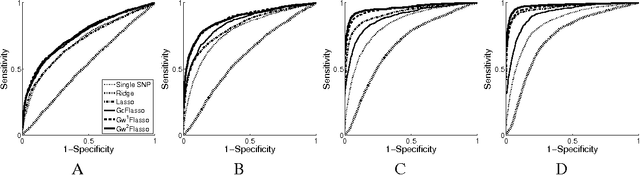
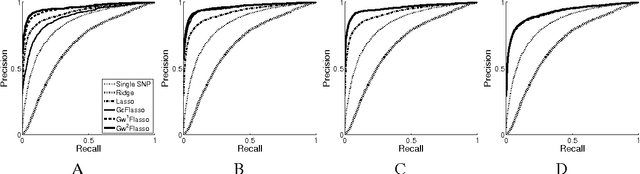
Abstract:Many complex disease syndromes such as asthma consist of a large number of highly related, rather than independent, clinical phenotypes, raising a new technical challenge in identifying genetic variations associated simultaneously with correlated traits. In this study, we propose a new statistical framework called graph-guided fused lasso (GFlasso) to address this issue in a principled way. Our approach explicitly represents the dependency structure among the quantitative traits as a network, and leverages this trait network to encode structured regularizations in a multivariate regression model over the genotypes and traits, so that the genetic markers that jointly influence subgroups of highly correlated traits can be detected with high sensitivity and specificity. While most of the traditional methods examined each phenotype independently and combined the results afterwards, our approach analyzes all of the traits jointly in a single statistical method, and borrow information across correlated phenotypes to discover the genetic markers that perturbe a subset of correlated triats jointly rather than a single trait. Using simulated datasets based on the HapMap consortium data and an asthma dataset, we compare the performance of our method with the single-marker analysis, and other sparse regression methods such as the ridge regression and the lasso that do not use any structural information in the traits. Our results show that there is a significant advantage in detecting the true causal SNPs when we incorporate the correlation pattern in traits using our proposed methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge