Krzysztof Koras
Unified Molecule Generation and Property Prediction
Apr 23, 2025Abstract:Modeling the joint distribution of the data samples and their properties allows to construct a single model for both data generation and property prediction, with synergistic capabilities reaching beyond purely generative or predictive models. However, training joint models presents daunting architectural and optimization challenges. Here, we propose Hyformer, a transformer-based joint model that successfully blends the generative and predictive functionalities, using an alternating attention mask together with a unified pre-training scheme. We show that Hyformer rivals other joint models, as well as state-of-the-art molecule generation and property prediction models. Additionally, we show the benefits of joint modeling in downstream tasks of molecular representation learning, hit identification and antimicrobial peptide design.
A generative recommender system with GMM prior for cancer drug generation and sensitivity prediction
Jun 07, 2022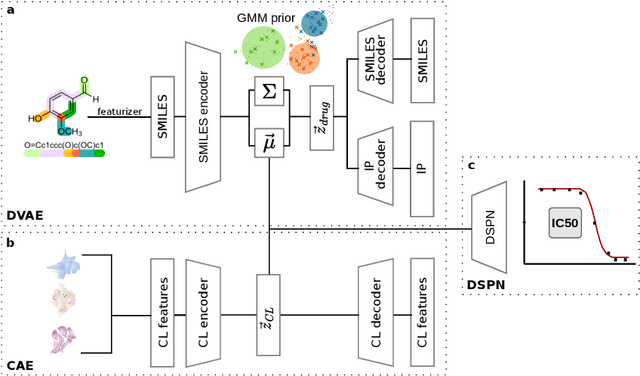
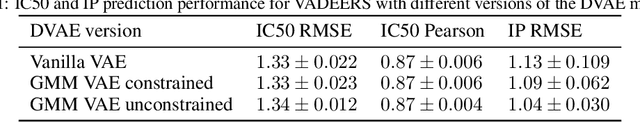

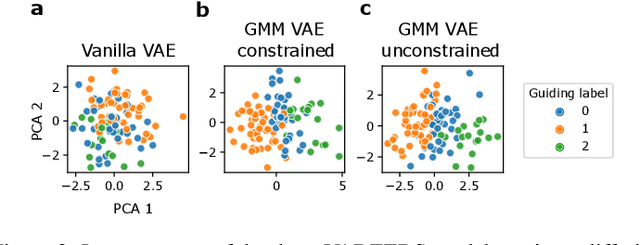
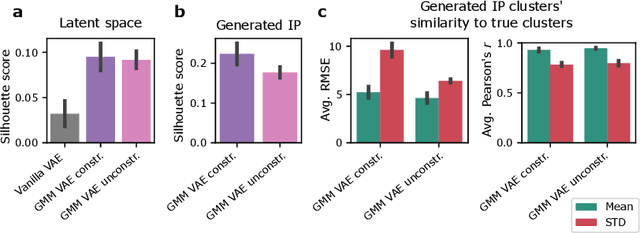
Abstract:Recent emergence of high-throughput drug screening assays sparkled an intensive development of machine learning methods, including models for prediction of sensitivity of cancer cell lines to anti-cancer drugs, as well as methods for generation of potential drug candidates. However, a concept of generation of compounds with specific properties and simultaneous modeling of their efficacy against cancer cell lines has not been comprehensively explored. To address this need, we present VADEERS, a Variational Autoencoder-based Drug Efficacy Estimation Recommender System. The generation of compounds is performed by a novel variational autoencoder with a semi-supervised Gaussian Mixture Model (GMM) prior. The prior defines a clustering in the latent space, where the clusters are associated with specific drug properties. In addition, VADEERS is equipped with a cell line autoencoder and a sensitivity prediction network. The model combines data for SMILES string representations of anti-cancer drugs, their inhibition profiles against a panel of protein kinases, cell lines biological features and measurements of the sensitivity of the cell lines to the drugs. The evaluated variants of VADEERS achieve a high r=0.87 Pearson correlation between true and predicted drug sensitivity estimates. We train the GMM prior in such a way that the clusters in the latent space correspond to a pre-computed clustering of the drugs by their inhibitory profiles. We show that the learned latent representations and new generated data points accurately reflect the given clustering. In summary, VADEERS offers a comprehensive model of drugs and cell lines properties and relationships between them, as well as a guided generation of novel compounds.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge