Katja Hose
Amalgam: Hybrid LLM-PGM Synthesis Algorithm for Accuracy and Realism
Mar 28, 2026Abstract:To generate synthetic datasets, e.g., in domains such as healthcare, the literature proposes approaches of two main types: Probabilistic Graphical Models (PGMs) and Deep Learning models, such as LLMs. While PGMs produce synthetic data that can be used for advanced analytics, they do not support complex schemas and datasets. LLMs on the other hand, support complex schemas but produce skewed dataset distributions, which are less useful for advanced analytics. In this paper, we therefore present Amalgam, a hybrid LLM-PGM data synthesis algorithm supporting both advanced analytics, realism, and tangible privacy properties. We show that Amalgam synthesizes data with an average 91 % $χ^2 P$ value and scores 3.8/5 for realism using our proposed metric, where state-of-the-art is 3.3 and real data is 4.7.
MultiHal: Multilingual Dataset for Knowledge-Graph Grounded Evaluation of LLM Hallucinations
May 20, 2025



Abstract:Large Language Models (LLMs) have inherent limitations of faithfulness and factuality, commonly referred to as hallucinations. Several benchmarks have been developed that provide a test bed for factuality evaluation within the context of English-centric datasets, while relying on supplementary informative context like web links or text passages but ignoring the available structured factual resources. To this end, Knowledge Graphs (KGs) have been identified as a useful aid for hallucination mitigation, as they provide a structured way to represent the facts about entities and their relations with minimal linguistic overhead. We bridge the lack of KG paths and multilinguality for factual language modeling within the existing hallucination evaluation benchmarks and propose a KG-based multilingual, multihop benchmark called \textbf{MultiHal} framed for generative text evaluation. As part of our data collection pipeline, we mined 140k KG-paths from open-domain KGs, from which we pruned noisy KG-paths, curating a high-quality subset of 25.9k. Our baseline evaluation shows an absolute scale increase by approximately 0.12 to 0.36 points for the semantic similarity score in KG-RAG over vanilla QA across multiple languages and multiple models, demonstrating the potential of KG integration. We anticipate MultiHal will foster future research towards several graph-based hallucination mitigation and fact-checking tasks.
The Limits of Graph Samplers for Training Inductive Recommender Systems: Extended results
May 20, 2025



Abstract:Inductive Recommender Systems are capable of recommending for new users and with new items thus avoiding the need to retrain after new data reaches the system. However, these methods are still trained on all the data available, requiring multiple days to train a single model, without counting hyperparameter tuning. In this work we focus on graph-based recommender systems, i.e., systems that model the data as a heterogeneous network. In other applications, graph sampling allows to study a subgraph and generalize the findings to the original graph. Thus, we investigate the applicability of sampling techniques for this task. We test on three real world datasets, with three state-of-the-art inductive methods, and using six different sampling methods. We find that its possible to maintain performance using only 50% of the training data with up to 86% percent decrease in training time; however, using less training data leads to far worse performance. Further, we find that when it comes to data for recommendations, graph sampling should also account for the temporal dimension. Therefore, we find that if higher data reduction is needed, new graph based sampling techniques should be studied and new inductive methods should be designed.
Semantic Web: Past, Present, and Future
Dec 22, 2024Abstract:Ever since the vision was formulated, the Semantic Web has inspired many generations of innovations. Semantic technologies have been used to share vast amounts of information on the Web, enhance them with semantics to give them meaning, and enable inference and reasoning on them. Throughout the years, semantic technologies, and in particular knowledge graphs, have been used in search engines, data integration, enterprise settings, and machine learning. In this paper, we recap the classical concepts and foundations of the Semantic Web as well as modern and recent concepts and applications, building upon these foundations. The classical topics we cover include knowledge representation, creating and validating knowledge on the Web, reasoning and linking, and distributed querying. We enhance this classical view of the so-called ``Semantic Web Layer Cake'' with an update of recent concepts that include provenance, security and trust, as well as a discussion of practical impacts from industry-led contributions. We conclude with an outlook on the future directions of the Semantic Web.
* Extended Version 2024-12-13 of TGDK 2(1): 3:1-3:37 (2024) If you like to contribute, please contact the first author and visit: https://github.com/ascherp/semantic-web-primer Please cite this paper as, see https://dblp.org/rec/journals/tgdk/ScherpG0HV24.html?view=bibtex
Knowledge Graphs, Large Language Models, and Hallucinations: An NLP Perspective
Nov 21, 2024


Abstract:Large Language Models (LLMs) have revolutionized Natural Language Processing (NLP) based applications including automated text generation, question answering, chatbots, and others. However, they face a significant challenge: hallucinations, where models produce plausible-sounding but factually incorrect responses. This undermines trust and limits the applicability of LLMs in different domains. Knowledge Graphs (KGs), on the other hand, provide a structured collection of interconnected facts represented as entities (nodes) and their relationships (edges). In recent research, KGs have been leveraged to provide context that can fill gaps in an LLM understanding of certain topics offering a promising approach to mitigate hallucinations in LLMs, enhancing their reliability and accuracy while benefiting from their wide applicability. Nonetheless, it is still a very active area of research with various unresolved open problems. In this paper, we discuss these open challenges covering state-of-the-art datasets and benchmarks as well as methods for knowledge integration and evaluating hallucinations. In our discussion, we consider the current use of KGs in LLM systems and identify future directions within each of these challenges.
The Susceptibility of Example-Based Explainability Methods to Class Outliers
Aug 01, 2024



Abstract:This study explores the impact of class outliers on the effectiveness of example-based explainability methods for black-box machine learning models. We reformulate existing explainability evaluation metrics, such as correctness and relevance, specifically for example-based methods, and introduce a new metric, distinguishability. Using these metrics, we highlight the shortcomings of current example-based explainability methods, including those who attempt to suppress class outliers. We conduct experiments on two datasets, a text classification dataset and an image classification dataset, and evaluate the performance of four state-of-the-art explainability methods. Our findings underscore the need for robust techniques to tackle the challenges posed by class outliers.
AIDE: Antithetical, Intent-based, and Diverse Example-Based Explanations
Jul 22, 2024



Abstract:For many use-cases, it is often important to explain the prediction of a black-box model by identifying the most influential training data samples. Existing approaches lack customization for user intent and often provide a homogeneous set of explanation samples, failing to reveal the model's reasoning from different angles. In this paper, we propose AIDE, an approach for providing antithetical (i.e., contrastive), intent-based, diverse explanations for opaque and complex models. AIDE distinguishes three types of explainability intents: interpreting a correct, investigating a wrong, and clarifying an ambiguous prediction. For each intent, AIDE selects an appropriate set of influential training samples that support or oppose the prediction either directly or by contrast. To provide a succinct summary, AIDE uses diversity-aware sampling to avoid redundancy and increase coverage of the training data. We demonstrate the effectiveness of AIDE on image and text classification tasks, in three ways: quantitatively, assessing correctness and continuity; qualitatively, comparing anecdotal evidence from AIDE and other example-based approaches; and via a user study, evaluating multiple aspects of AIDE. The results show that AIDE addresses the limitations of existing methods and exhibits desirable traits for an explainability method.
Hospitalization Length of Stay Prediction using Patient Event Sequences
Mar 20, 2023



Abstract:Predicting patients hospital length of stay (LOS) is essential for improving resource allocation and supporting decision-making in healthcare organizations. This paper proposes a novel approach for predicting LOS by modeling patient information as sequences of events. Specifically, we present a transformer-based model, termed Medic-BERT (M-BERT), for LOS prediction using the unique features describing patients medical event sequences. We performed empirical experiments on a cohort of more than 45k emergency care patients from a large Danish hospital. Experimental results show that M-BERT can achieve high accuracy on a variety of LOS problems and outperforms traditional nonsequence-based machine learning approaches.
Linked Data Science Powered by Knowledge Graphs
Mar 09, 2023



Abstract:In recent years, we have witnessed a growing interest in data science not only from academia but particularly from companies investing in data science platforms to analyze large amounts of data. In this process, a myriad of data science artifacts, such as datasets and pipeline scripts, are created. Yet, there has so far been no systematic attempt to holistically exploit the collected knowledge and experiences that are implicitly contained in the specification of these pipelines, e.g., compatible datasets, cleansing steps, ML algorithms, parameters, etc. Instead, data scientists still spend a considerable amount of their time trying to recover relevant information and experiences from colleagues, trial and error, lengthy exploration, etc. In this paper, we, therefore, propose a scalable system (KGLiDS) that employs machine learning to extract the semantics of data science pipelines and captures them in a knowledge graph, which can then be exploited to assist data scientists in various ways. This abstraction is the key to enabling Linked Data Science since it allows us to share the essence of pipelines between platforms, companies, and institutions without revealing critical internal information and instead focusing on the semantics of what is being processed and how. Our comprehensive evaluation uses thousands of datasets and more than thirteen thousand pipeline scripts extracted from data discovery benchmarks and the Kaggle portal and shows that KGLiDS significantly outperforms state-of-the-art systems on related tasks, such as dataset recommendation and pipeline classification.
Simple and Powerful Architecture for Inductive Recommendation Using Knowledge Graph Convolutions
Sep 13, 2022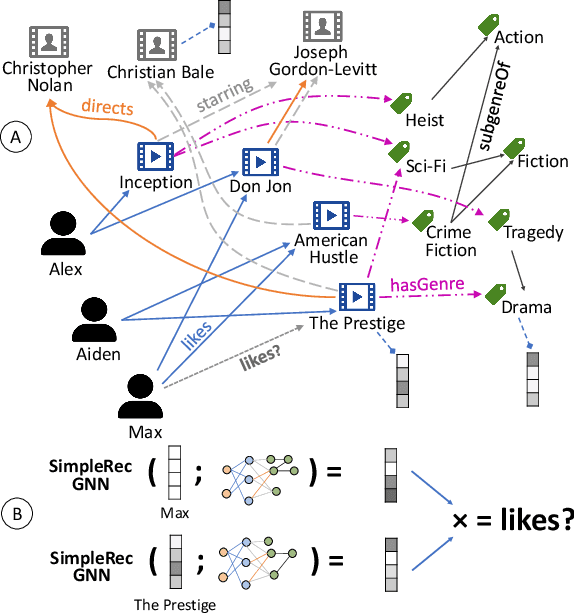
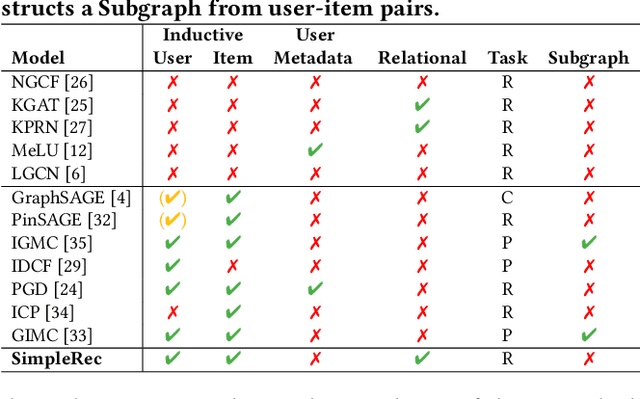
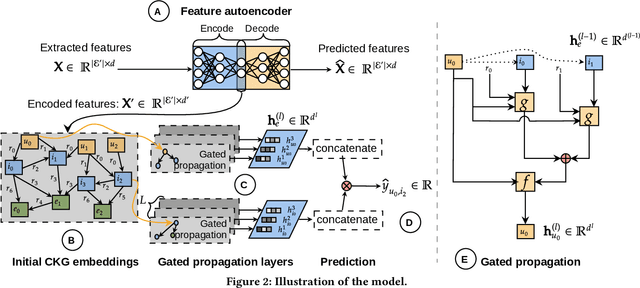
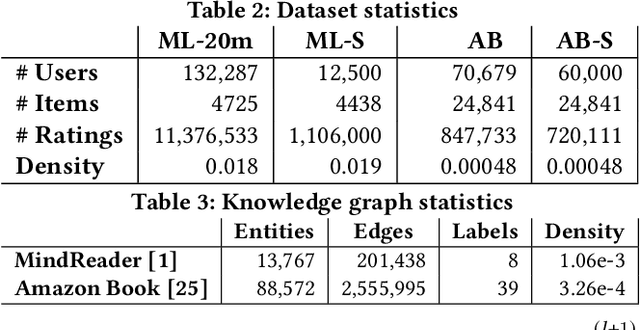
Abstract:Using graph models with relational information in recommender systems has shown promising results. Yet, most methods are transductive, i.e., they are based on dimensionality reduction architectures. Hence, they require heavy retraining every time new items or users are added. Conversely, inductive methods promise to solve these issues. Nonetheless, all inductive methods rely only on interactions, making recommendations for users with few interactions sub-optimal and even impossible for new items. Therefore, we focus on inductive methods able to also exploit knowledge graphs (KGs). In this work, we propose SimpleRec, a strong baseline that uses a graph neural network and a KG to provide better recommendations than related inductive methods for new users and items. We show that it is unnecessary to create complex model architectures for user representations, but it is enough to allow users to be represented by the few ratings they provide and the indirect connections among them without any user metadata. As a result, we re-evaluate state-of-the-art methods, identify better evaluation protocols, highlight unwarranted conclusions from previous proposals, and showcase a novel, stronger baseline for this task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge