Joseph Khoury
Enhancing Network Security Management in Water Systems using FM-based Attack Attribution
Mar 03, 2025



Abstract:Water systems are vital components of modern infrastructure, yet they are increasingly susceptible to sophisticated cyber attacks with potentially dire consequences on public health and safety. While state-of-the-art machine learning techniques effectively detect anomalies, contemporary model-agnostic attack attribution methods using LIME, SHAP, and LEMNA are deemed impractical for large-scale, interdependent water systems. This is due to the intricate interconnectivity and dynamic interactions that define these complex environments. Such methods primarily emphasize individual feature importance while falling short of addressing the crucial sensor-actuator interactions in water systems, which limits their effectiveness in identifying root cause attacks. To this end, we propose a novel model-agnostic Factorization Machines (FM)-based approach that capitalizes on water system sensor-actuator interactions to provide granular explanations and attributions for cyber attacks. For instance, an anomaly in an actuator pump activity can be attributed to a top root cause attack candidates, a list of water pressure sensors, which is derived from the underlying linear and quadratic effects captured by our approach. We validate our method using two real-world water system specific datasets, SWaT and WADI, demonstrating its superior performance over traditional attribution methods. In multi-feature cyber attack scenarios involving intricate sensor-actuator interactions, our FM-based attack attribution method effectively ranks attack root causes, achieving approximately 20% average improvement over SHAP and LEMNA.
Enhancing Reverse Engineering: Investigating and Benchmarking Large Language Models for Vulnerability Analysis in Decompiled Binaries
Nov 07, 2024Abstract:Security experts reverse engineer (decompile) binary code to identify critical security vulnerabilities. The limited access to source code in vital systems - such as firmware, drivers, and proprietary software used in Critical Infrastructures (CI) - makes this analysis even more crucial on the binary level. Even with available source code, a semantic gap persists after compilation between the source and the binary code executed by the processor. This gap may hinder the detection of vulnerabilities in source code. That being said, current research on Large Language Models (LLMs) overlooks the significance of decompiled binaries in this area by focusing solely on source code. In this work, we are the first to empirically uncover the substantial semantic limitations of state-of-the-art LLMs when it comes to analyzing vulnerabilities in decompiled binaries, largely due to the absence of relevant datasets. To bridge the gap, we introduce DeBinVul, a novel decompiled binary code vulnerability dataset. Our dataset is multi-architecture and multi-optimization, focusing on C/C++ due to their wide usage in CI and association with numerous vulnerabilities. Specifically, we curate 150,872 samples of vulnerable and non-vulnerable decompiled binary code for the task of (i) identifying; (ii) classifying; (iii) describing vulnerabilities; and (iv) recovering function names in the domain of decompiled binaries. Subsequently, we fine-tune state-of-the-art LLMs using DeBinVul and report on a performance increase of 19%, 24%, and 21% in the capabilities of CodeLlama, Llama3, and CodeGen2 respectively, in detecting binary code vulnerabilities. Additionally, using DeBinVul, we report a high performance of 80-90% on the vulnerability classification task. Furthermore, we report improved performance in function name recovery and vulnerability description tasks.
Enhancing Source Code Security with LLMs: Demystifying The Challenges and Generating Reliable Repairs
Sep 01, 2024



Abstract:With the recent unprecedented advancements in Artificial Intelligence (AI) computing, progress in Large Language Models (LLMs) is accelerating rapidly, presenting challenges in establishing clear guidelines, particularly in the field of security. That being said, we thoroughly identify and describe three main technical challenges in the security and software engineering literature that spans the entire LLM workflow, namely; \textbf{\textit{(i)}} Data Collection and Labeling; \textbf{\textit{(ii)}} System Design and Learning; and \textbf{\textit{(iii)}} Performance Evaluation. Building upon these challenges, this paper introduces \texttt{SecRepair}, an instruction-based LLM system designed to reliably \textit{identify}, \textit{describe}, and automatically \textit{repair} vulnerable source code. Our system is accompanied by a list of actionable guides on \textbf{\textit{(i)}} Data Preparation and Augmentation Techniques; \textbf{\textit{(ii)}} Selecting and Adapting state-of-the-art LLM Models; \textbf{\textit{(iii)}} Evaluation Procedures. \texttt{SecRepair} uses a reinforcement learning-based fine-tuning with a semantic reward that caters to the functionality and security aspects of the generated code. Our empirical analysis shows that \texttt{SecRepair} achieves a \textit{12}\% improvement in security code repair compared to other LLMs when trained using reinforcement learning. Furthermore, we demonstrate the capabilities of \texttt{SecRepair} in generating reliable, functional, and compilable security code repairs against real-world test cases using automated evaluation metrics.
LLM-Powered Code Vulnerability Repair with Reinforcement Learning and Semantic Reward
Jan 07, 2024



Abstract:In software development, the predominant emphasis on functionality often supersedes security concerns, a trend gaining momentum with AI-driven automation tools like GitHub Copilot. These tools significantly improve developers' efficiency in functional code development. Nevertheless, it remains a notable concern that such tools are also responsible for creating insecure code, predominantly because of pre-training on publicly available repositories with vulnerable code. Moreover, developers are called the "weakest link in the chain" since they have very minimal knowledge of code security. Although existing solutions provide a reasonable solution to vulnerable code, they must adequately describe and educate the developers on code security to ensure that the security issues are not repeated. Therefore we introduce a multipurpose code vulnerability analysis system \texttt{SecRepair}, powered by a large language model, CodeGen2 assisting the developer in identifying and generating fixed code along with a complete description of the vulnerability with a code comment. Our innovative methodology uses a reinforcement learning paradigm to generate code comments augmented by a semantic reward mechanism. Inspired by how humans fix code issues, we propose an instruction-based dataset suitable for vulnerability analysis with LLMs. We further identify zero-day and N-day vulnerabilities in 6 Open Source IoT Operating Systems on GitHub. Our findings underscore that incorporating reinforcement learning coupled with semantic reward augments our model's performance, thereby fortifying its capacity to address code vulnerabilities with improved efficacy.
Hierarchical Phenotyping and Graph Modeling of Spatial Architecture in Lymphoid Neoplasms
Jun 30, 2021
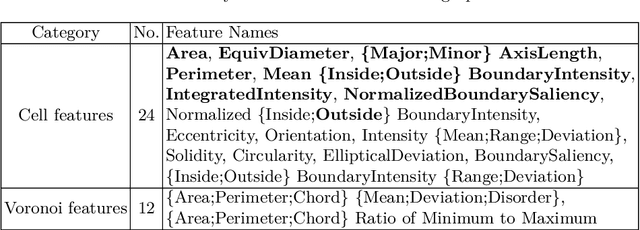
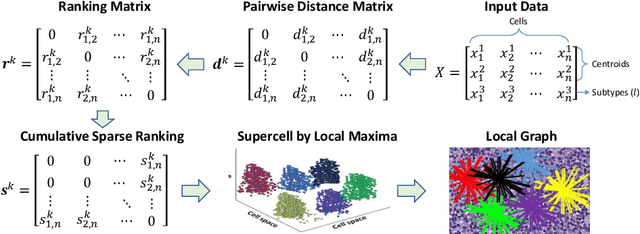
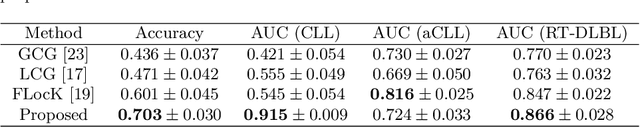
Abstract:The cells and their spatial patterns in the tumor microenvironment (TME) play a key role in tumor evolution, and yet remains an understudied topic in computational pathology. This study, to the best of our knowledge, is among the first to hybrid local and global graph methods to profile orchestration and interaction of cellular components. To address the challenge in hematolymphoid cancers where the cell classes in TME are unclear, we first implemented cell level unsupervised learning and identified two new cell subtypes. Local cell graphs or supercells were built for each image by considering the individual cell's geospatial location and classes. Then, we applied supercell level clustering and identified two new cell communities. In the end, we built global graphs to abstract spatial interaction patterns and extract features for disease diagnosis. We evaluate the proposed algorithm on H\&E slides of 60 hematolymphoid neoplasm patients and further compared it with three cell level graph-based algorithms, including the global cell graph, cluster cell graph, and FLocK. The proposed algorithm achieves a mean diagnosis accuracy of 0.703 with the repeated 5-fold cross-validation scheme. In conclusion, our algorithm shows superior performance over the existing methods and can be potentially applied to other cancer types.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge