Jayaraman J. Thiagarajan
Lawrence Livermore National Laboratory, Livermore, CA
Can Deep Clinical Models Handle Real-World Domain Shifts?
Sep 20, 2018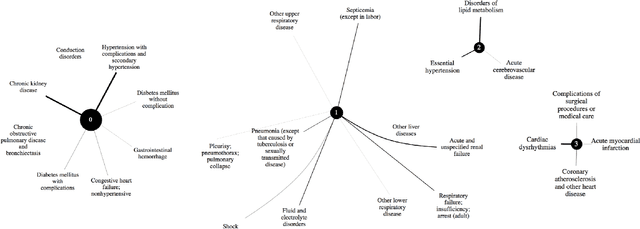
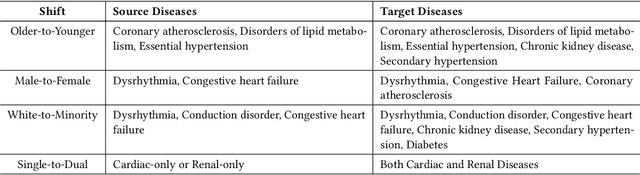
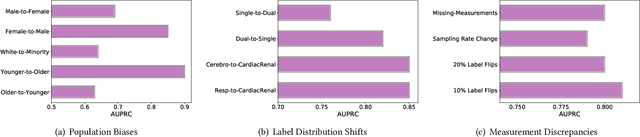
Abstract:The hypothesis that computational models can be reliable enough to be adopted in prognosis and patient care is revolutionizing healthcare. Deep learning, in particular, has been a game changer in building predictive models, thereby leading to community-wide data curation efforts. However, due to the inherent variabilities in population characteristics and biological systems, these models are often biased to the training datasets. This can be limiting when models are deployed in new environments, particularly when there are systematic domain shifts not known a priori. In this paper, we formalize these challenges by emulating a large class of domain shifts that can occur in clinical settings, and argue that evaluating the behavior of predictive models in light of those shifts is an effective way of quantifying the reliability of clinical models. More specifically, we develop an approach for building challenging scenarios, based on analysis of \textit{disease landscapes}, and utilize unsupervised domain adaptation to compensate for the domain shifts. Using the openly available MIMIC-III EHR dataset for phenotyping, we generate a large class of scenarios and evaluate the ability of deep clinical models in those cases. For the first time, our work sheds light into data regimes where deep clinical models can fail to generalize, due to significant changes in the disease landscapes between the source and target landscapes. This study emphasizes the need for sophisticated evaluation mechanisms driven by real-world domain shifts to build effective AI solutions for healthcare.
Controlled Random Search Improves Sample Mining and Hyper-Parameter Optimization
Sep 05, 2018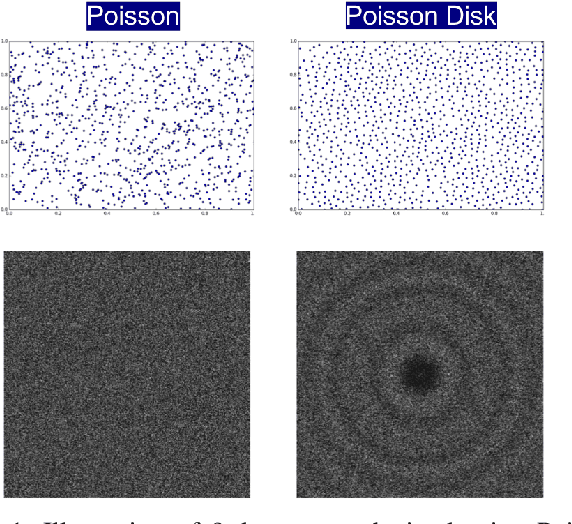
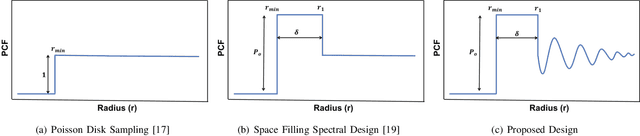
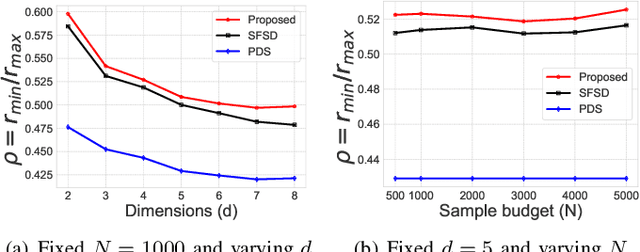
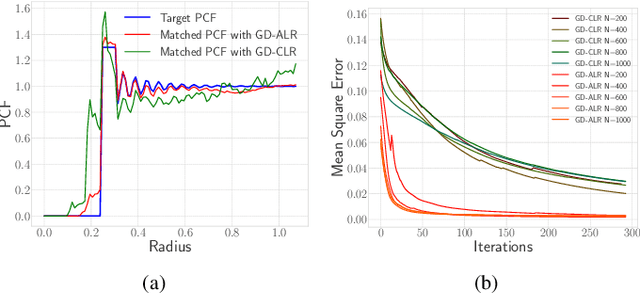
Abstract:A common challenge in machine learning and related fields is the need to efficiently explore high dimensional parameter spaces using small numbers of samples. Typical examples are hyper-parameter optimization in deep learning and sample mining in predictive modeling tasks. All such problems trade-off exploration, which samples the space without knowledge of the target function, and exploitation where information from previous evaluations is used in an adaptive feedback loop. Much of the recent focus has been on the exploitation while exploration is done with simple designs such as Latin hypercube or even uniform random sampling. In this paper, we introduce optimal space-filling sample designs for effective exploration of high dimensional spaces. Specifically, we propose a new parameterized family of sample designs called space-filling spectral designs, and introduce a framework to choose optimal designs for a given sample size and dimension. Furthermore, we present an efficient algorithm to synthesize a given spectral design. Finally, we evaluate the performance of spectral designs in both data space and model space applications. The data space exploration is targeted at recovering complex regression functions in high dimensional spaces. The model space exploration focuses on selecting hyper-parameters for a given neural network architecture. Our empirical studies demonstrate that the proposed approach consistently outperforms state-of-the-art techniques, particularly with smaller design sizes.
Triplet Network with Attention for Speaker Diarization
Aug 04, 2018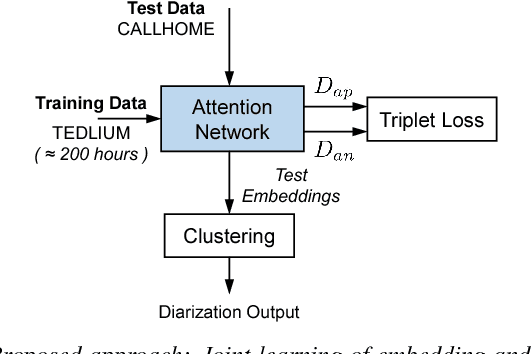
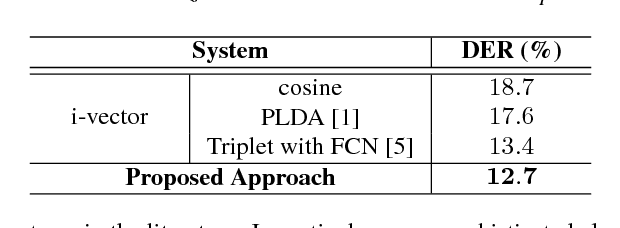

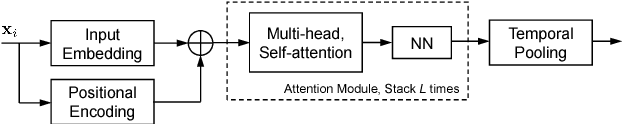
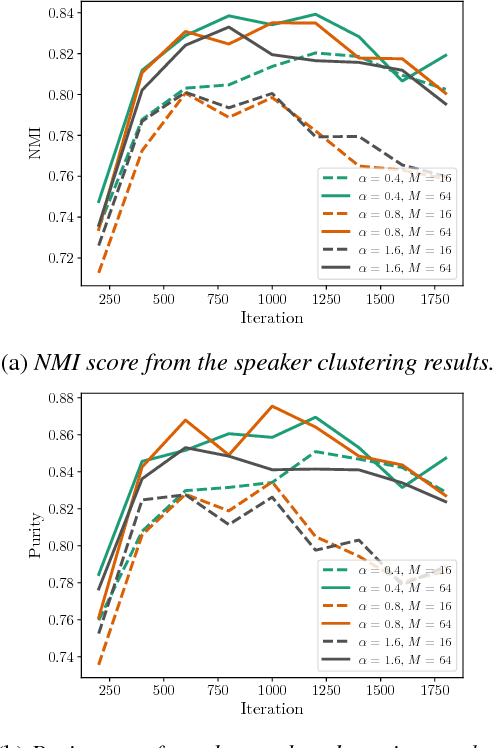
Abstract:In automatic speech processing systems, speaker diarization is a crucial front-end component to separate segments from different speakers. Inspired by the recent success of deep neural networks (DNNs) in semantic inferencing, triplet loss-based architectures have been successfully used for this problem. However, existing work utilizes conventional i-vectors as the input representation and builds simple fully connected networks for metric learning, thus not fully leveraging the modeling power of DNN architectures. This paper investigates the importance of learning effective representations from the sequences directly in metric learning pipelines for speaker diarization. More specifically, we propose to employ attention models to learn embeddings and the metric jointly in an end-to-end fashion. Experiments are conducted on the CALLHOME conversational speech corpus. The diarization results demonstrate that, besides providing a unified model, the proposed approach achieves improved performance when compared against existing approaches.
Lose The Views: Limited Angle CT Reconstruction via Implicit Sinogram Completion
Jul 11, 2018



Abstract:Computed Tomography (CT) reconstruction is a fundamental component to a wide variety of applications ranging from security, to healthcare. The classical techniques require measuring projections, called sinograms, from a full 180$^\circ$ view of the object. This is impractical in a limited angle scenario, when the viewing angle is less than 180$^\circ$, which can occur due to different factors including restrictions on scanning time, limited flexibility of scanner rotation, etc. The sinograms obtained as a result, cause existing techniques to produce highly artifact-laden reconstructions. In this paper, we propose to address this problem through implicit sinogram completion, on a challenging real world dataset containing scans of common checked-in luggage. We propose a system, consisting of 1D and 2D convolutional neural networks, that operates on a limited angle sinogram to directly produce the best estimate of a reconstruction. Next, we use the x-ray transform on this reconstruction to obtain a "completed" sinogram, as if it came from a full 180$^\circ$ measurement. We feed this to standard analytical and iterative reconstruction techniques to obtain the final reconstruction. We show with extensive experimentation that this combined strategy outperforms many competitive baselines. We also propose a measure of confidence for the reconstruction that enables a practitioner to gauge the reliability of a prediction made by our network. We show that this measure is a strong indicator of quality as measured by the PSNR, while not requiring ground truth at test time. Finally, using a segmentation experiment, we show that our reconstruction preserves the 3D structure of objects effectively.
A Generative Modeling Approach to Limited Channel ECG Classification
Jun 14, 2018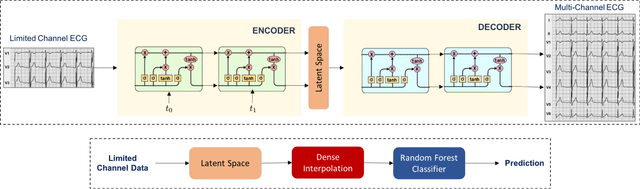
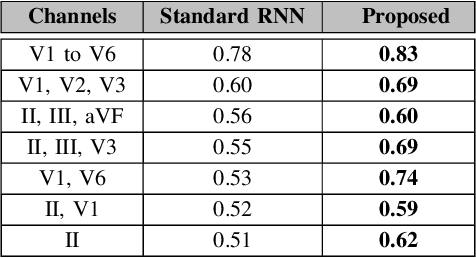

Abstract:Processing temporal sequences is central to a variety of applications in health care, and in particular multi-channel Electrocardiogram (ECG) is a highly prevalent diagnostic modality that relies on robust sequence modeling. While Recurrent Neural Networks (RNNs) have led to significant advances in automated diagnosis with time-series data, they perform poorly when models are trained using a limited set of channels. A crucial limitation of existing solutions is that they rely solely on discriminative models, which tend to generalize poorly in such scenarios. In order to combat this limitation, we develop a generative modeling approach to limited channel ECG classification. This approach first uses a Seq2Seq model to implicitly generate the missing channel information, and then uses the latent representation to perform the actual supervisory task. This decoupling enables the use of unsupervised data and also provides highly robust metric spaces for subsequent discriminative learning. Our experiments with the Physionet dataset clearly evidence the effectiveness of our approach over standard RNNs in disease prediction.
An Unsupervised Approach to Solving Inverse Problems using Generative Adversarial Networks
Jun 04, 2018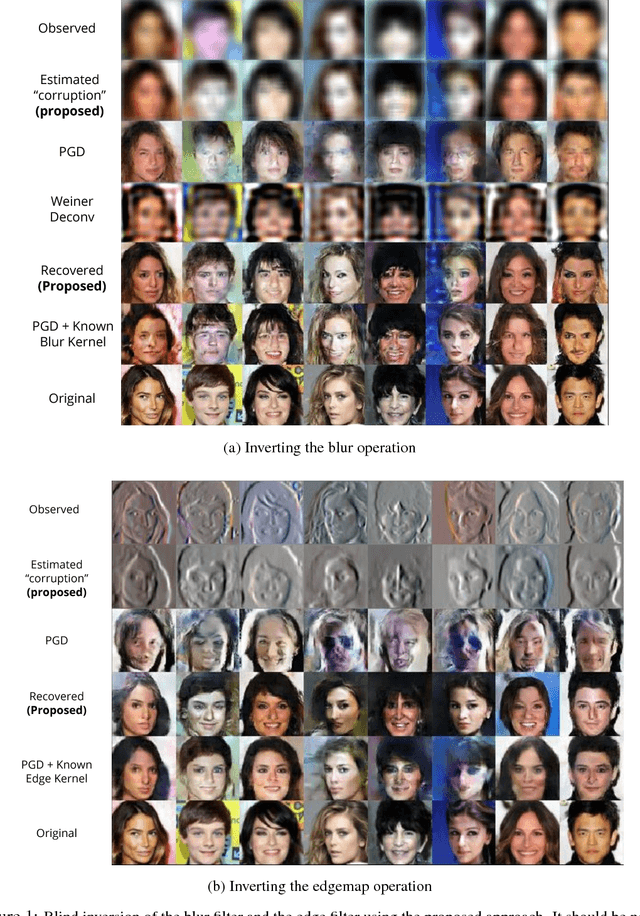
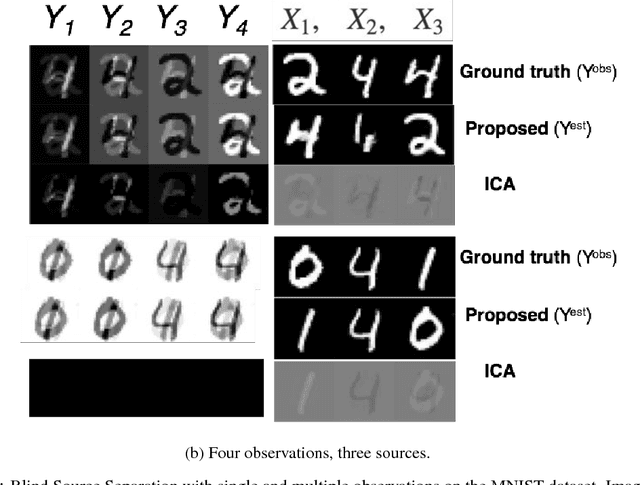

Abstract:Solving inverse problems continues to be a challenge in a wide array of applications ranging from deblurring, image inpainting, source separation etc. Most existing techniques solve such inverse problems by either explicitly or implicitly finding the inverse of the model. The former class of techniques require explicit knowledge of the measurement process which can be unrealistic, and rely on strong analytical regularizers to constrain the solution space, which often do not generalize well. The latter approaches have had remarkable success in part due to deep learning, but require a large collection of source-observation pairs, which can be prohibitively expensive. In this paper, we propose an unsupervised technique to solve inverse problems with generative adversarial networks (GANs). Using a pre-trained GAN in the space of source signals, we show that one can reliably recover solutions to under determined problems in a `blind' fashion, i.e., without knowledge of the measurement process. We solve this by making successive estimates on the model and the solution in an iterative fashion. We show promising results in three challenging applications -- blind source separation, image deblurring, and recovering an image from its edge map, and perform better than several baselines.
MARGIN: Uncovering Deep Neural Networks using Graph Signal Analysis
Feb 10, 2018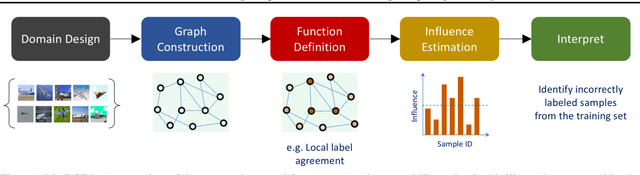
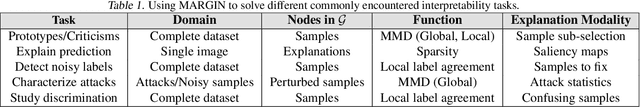

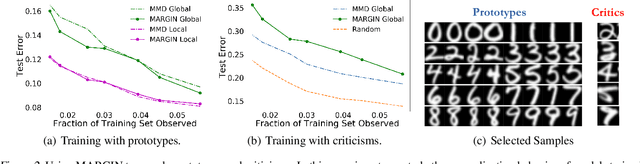
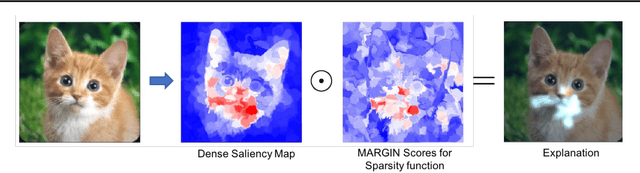
Abstract:Interpretability has emerged as a crucial aspect of machine learning, aimed at providing insights into the working of complex neural networks. However, existing solutions vary vastly based on the nature of the interpretability task, with each use case requiring substantial time and effort. This paper introduces MARGIN, a simple yet general approach to address a large set of interpretability tasks ranging from identifying prototypes to explaining image predictions. MARGIN exploits ideas rooted in graph signal analysis to determine influential nodes in a graph, which are defined as those nodes that maximally describe a function defined on the graph. By carefully defining task-specific graphs and functions, we demonstrate that MARGIN outperforms existing approaches in a number of disparate interpretability challenges.
Exploring High-Dimensional Structure via Axis-Aligned Decomposition of Linear Projections
Dec 20, 2017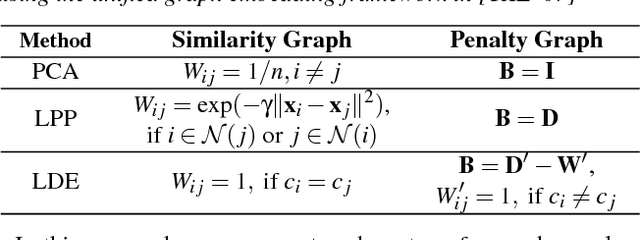
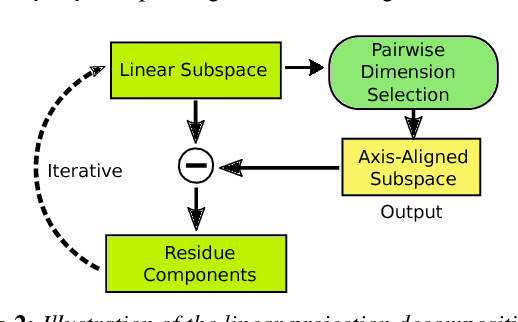
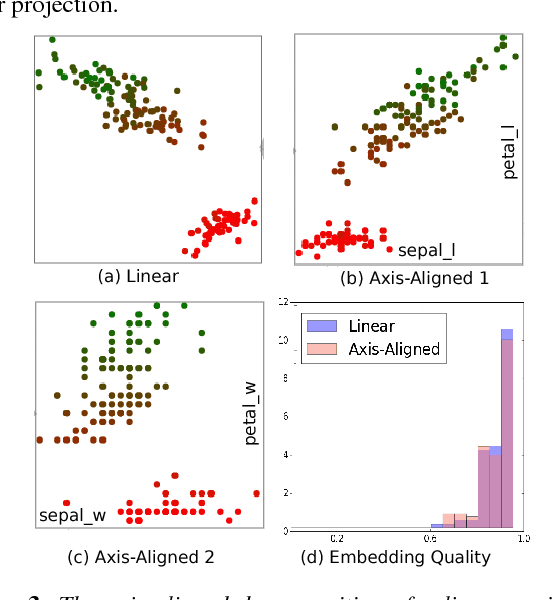
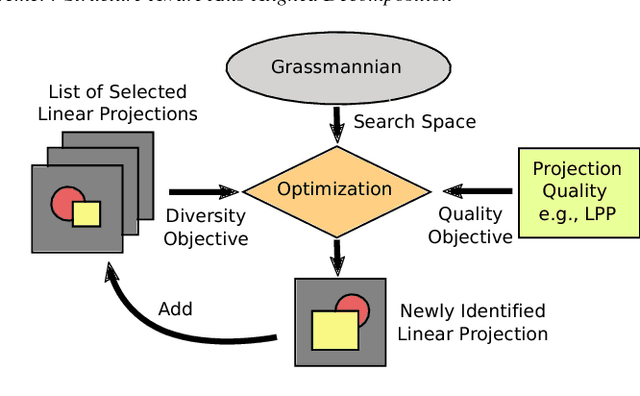
Abstract:Two-dimensional embeddings remain the dominant approach to visualize high dimensional data. The choice of embeddings ranges from highly non-linear ones, which can capture complex relationships but are difficult to interpret quantitatively, to axis-aligned projections, which are easy to interpret but are limited to bivariate relationships. Linear project can be considered as a compromise between complexity and interpretability, as they allow explicit axes labels, yet provide significantly more degrees of freedom compared to axis-aligned projections. Nevertheless, interpreting the axes directions, which are linear combinations often with many non-trivial components, remains difficult. To address this problem we introduce a structure aware decomposition of (multiple) linear projections into sparse sets of axis aligned projections, which jointly capture all information of the original linear ones. In particular, we use tools from Dempster-Shafer theory to formally define how relevant a given axis aligned project is to explain the neighborhood relations displayed in some linear projection. Furthermore, we introduce a new approach to discover a diverse set of high quality linear projections and show that in practice the information of $k$ linear projections is often jointly encoded in $\sim k$ axis aligned plots. We have integrated these ideas into an interactive visualization system that allows users to jointly browse both linear projections and their axis aligned representatives. Using a number of case studies we show how the resulting plots lead to more intuitive visualizations and new insight.
A Spectral Approach for the Design of Experiments: Design, Analysis and Algorithms
Dec 16, 2017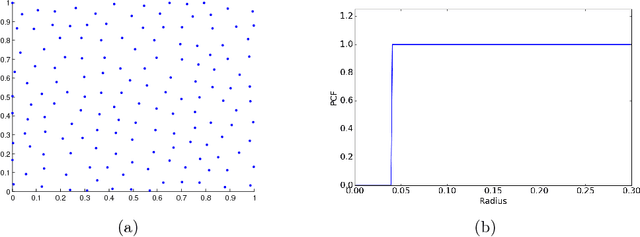
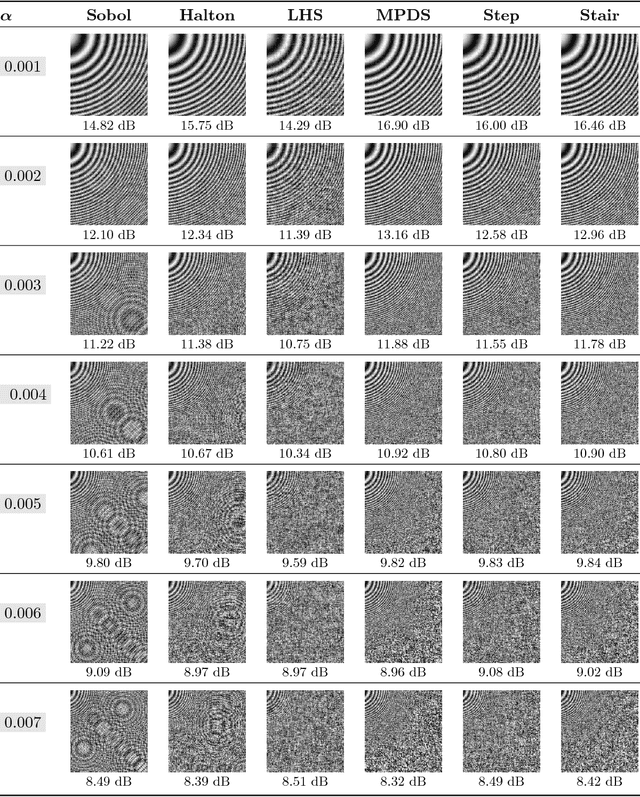

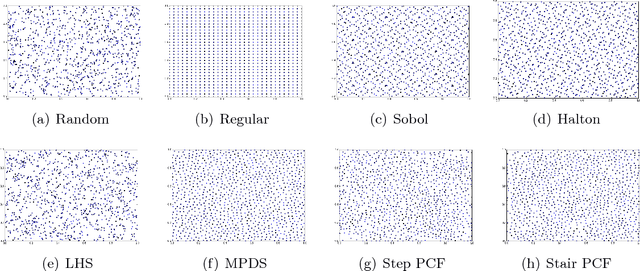
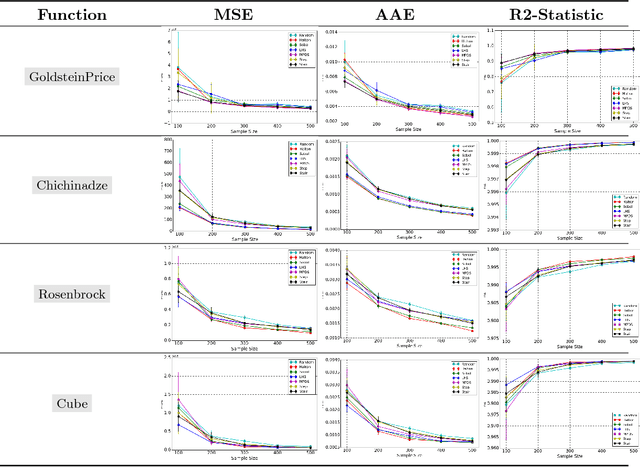
Abstract:This paper proposes a new approach to construct high quality space-filling sample designs. First, we propose a novel technique to quantify the space-filling property and optimally trade-off uniformity and randomness in sample designs in arbitrary dimensions. Second, we connect the proposed metric (defined in the spatial domain) to the objective measure of the design performance (defined in the spectral domain). This connection serves as an analytic framework for evaluating the qualitative properties of space-filling designs in general. Using the theoretical insights provided by this spatial-spectral analysis, we derive the notion of optimal space-filling designs, which we refer to as space-filling spectral designs. Third, we propose an efficient estimator to evaluate the space-filling properties of sample designs in arbitrary dimensions and use it to develop an optimization framework to generate high quality space-filling designs. Finally, we carry out a detailed performance comparison on two different applications in 2 to 6 dimensions: a) image reconstruction and b) surrogate modeling on several benchmark optimization functions and an inertial confinement fusion (ICF) simulation code. We demonstrate that the propose spectral designs significantly outperform existing approaches especially in high dimensions.
Attend and Diagnose: Clinical Time Series Analysis using Attention Models
Nov 19, 2017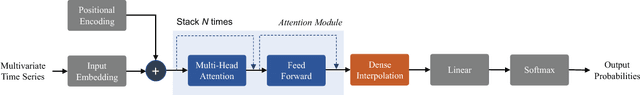
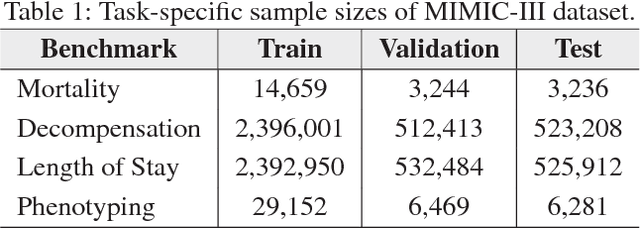
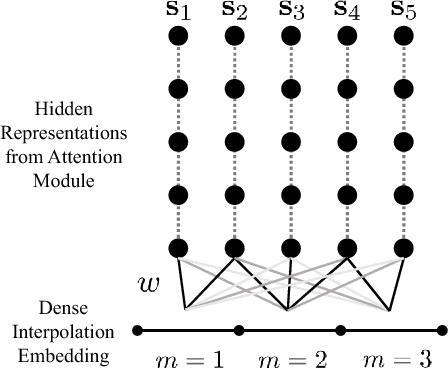
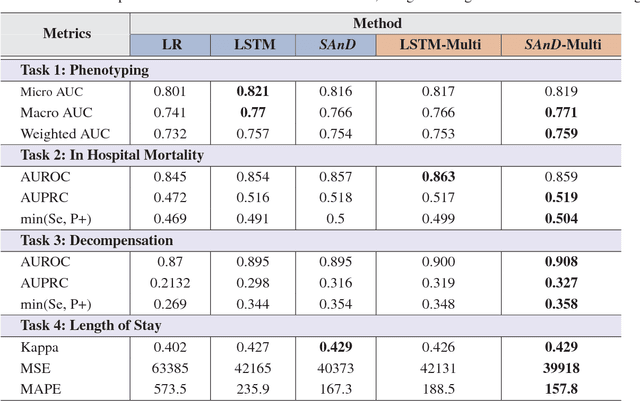
Abstract:With widespread adoption of electronic health records, there is an increased emphasis for predictive models that can effectively deal with clinical time-series data. Powered by Recurrent Neural Network (RNN) architectures with Long Short-Term Memory (LSTM) units, deep neural networks have achieved state-of-the-art results in several clinical prediction tasks. Despite the success of RNNs, its sequential nature prohibits parallelized computing, thus making it inefficient particularly when processing long sequences. Recently, architectures which are based solely on attention mechanisms have shown remarkable success in transduction tasks in NLP, while being computationally superior. In this paper, for the first time, we utilize attention models for clinical time-series modeling, thereby dispensing recurrence entirely. We develop the \textit{SAnD} (Simply Attend and Diagnose) architecture, which employs a masked, self-attention mechanism, and uses positional encoding and dense interpolation strategies for incorporating temporal order. Furthermore, we develop a multi-task variant of \textit{SAnD} to jointly infer models with multiple diagnosis tasks. Using the recent MIMIC-III benchmark datasets, we demonstrate that the proposed approach achieves state-of-the-art performance in all tasks, outperforming LSTM models and classical baselines with hand-engineered features.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge