James Thomas
Exploring the use of a Large Language Model for data extraction in systematic reviews: a rapid feasibility study
May 23, 2024



Abstract:This paper describes a rapid feasibility study of using GPT-4, a large language model (LLM), to (semi)automate data extraction in systematic reviews. Despite the recent surge of interest in LLMs there is still a lack of understanding of how to design LLM-based automation tools and how to robustly evaluate their performance. During the 2023 Evidence Synthesis Hackathon we conducted two feasibility studies. Firstly, to automatically extract study characteristics from human clinical, animal, and social science domain studies. We used two studies from each category for prompt-development; and ten for evaluation. Secondly, we used the LLM to predict Participants, Interventions, Controls and Outcomes (PICOs) labelled within 100 abstracts in the EBM-NLP dataset. Overall, results indicated an accuracy of around 80%, with some variability between domains (82% for human clinical, 80% for animal, and 72% for studies of human social sciences). Causal inference methods and study design were the data extraction items with the most errors. In the PICO study, participants and intervention/control showed high accuracy (>80%), outcomes were more challenging. Evaluation was done manually; scoring methods such as BLEU and ROUGE showed limited value. We observed variability in the LLMs predictions and changes in response quality. This paper presents a template for future evaluations of LLMs in the context of data extraction for systematic review automation. Our results show that there might be value in using LLMs, for example as second or third reviewers. However, caution is advised when integrating models such as GPT-4 into tools. Further research on stability and reliability in practical settings is warranted for each type of data that is processed by the LLM.
* Conference proceedings, peer-reviewed and presented at the 3rd Workshop on Augmented Intelligence for Technology-Assisted Reviews Systems, Glasgow, 2024
Constructing Artificial Data for Fine-tuning for Low-Resource Biomedical Text Tagging with Applications in PICO Annotation
Oct 21, 2019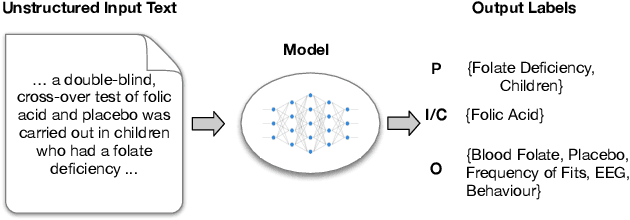

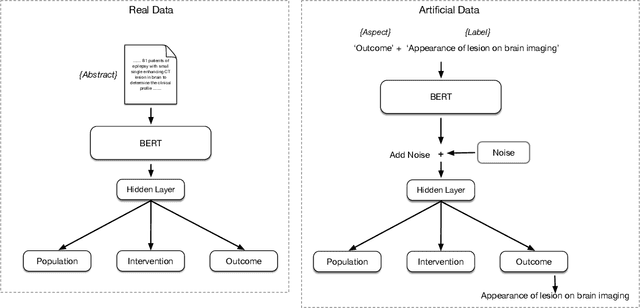
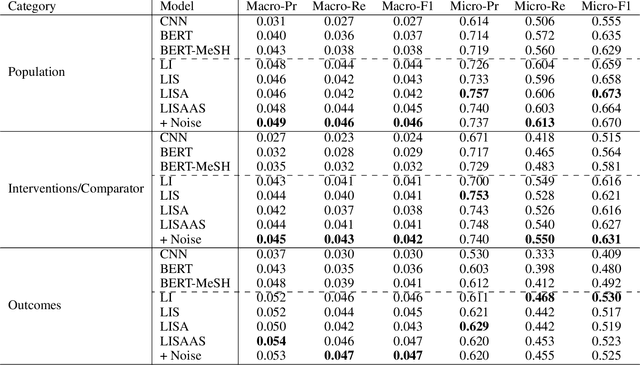
Abstract:Biomedical text tagging systems are plagued by the dearth of labeled training data. There have been recent attempts at using pre-trained encoders to deal with this issue. Pre-trained encoder provides representation of the input text which is then fed to task-specific layers for classification. The entire network is fine-tuned on the labeled data from the target task. Unfortunately, a low-resource biomedical task often has too few labeled instances for satisfactory fine-tuning. Also, if the label space is large, it contains few or no labeled instances for majority of the labels. Most biomedical tagging systems treat labels as indexes, ignoring the fact that these labels are often concepts expressed in natural language e.g. `Appearance of lesion on brain imaging'. To address these issues, we propose constructing extra labeled instances using label-text (i.e. label's name) as input for the corresponding label-index (i.e. label's index). In fact, we propose a number of strategies for manufacturing multiple artificial labeled instances from a single label. The network is then fine-tuned on a combination of real and these newly constructed artificial labeled instances. We evaluate the proposed approach on an important low-resource biomedical task called \textit{PICO annotation}, which requires tagging raw text describing clinical trials with labels corresponding to different aspects of the trial i.e. PICO (Population, Intervention/Control, Outcome) characteristics of the trial. Our empirical results show that the proposed method achieves a new state-of-the-art performance for PICO annotation with very significant improvements over competitive baselines.
Structured Multi-Label Biomedical Text Tagging via Attentive Neural Tree Decoding
Oct 02, 2018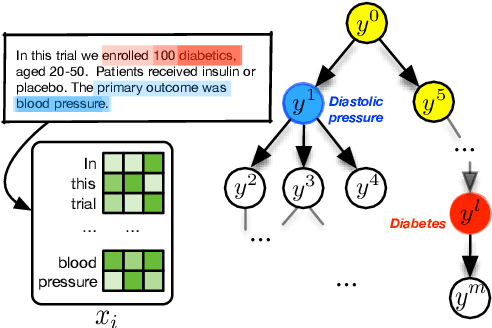

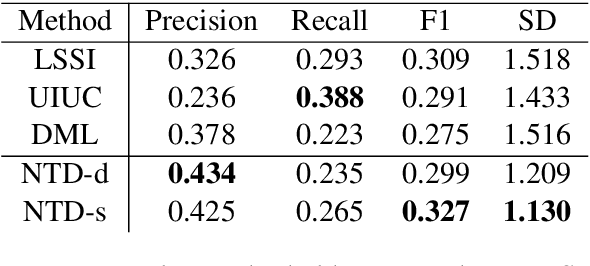
Abstract:We propose a model for tagging unstructured texts with an arbitrary number of terms drawn from a tree-structured vocabulary (i.e., an ontology). We treat this as a special case of sequence-to-sequence learning in which the decoder begins at the root node of an ontological tree and recursively elects to expand child nodes as a function of the input text, the current node, and the latent decoder state. In our experiments the proposed method outperforms state-of-the-art approaches on the important task of automatically assigning MeSH terms to biomedical abstracts.
Improving Active Learning in Systematic Reviews
Jan 29, 2018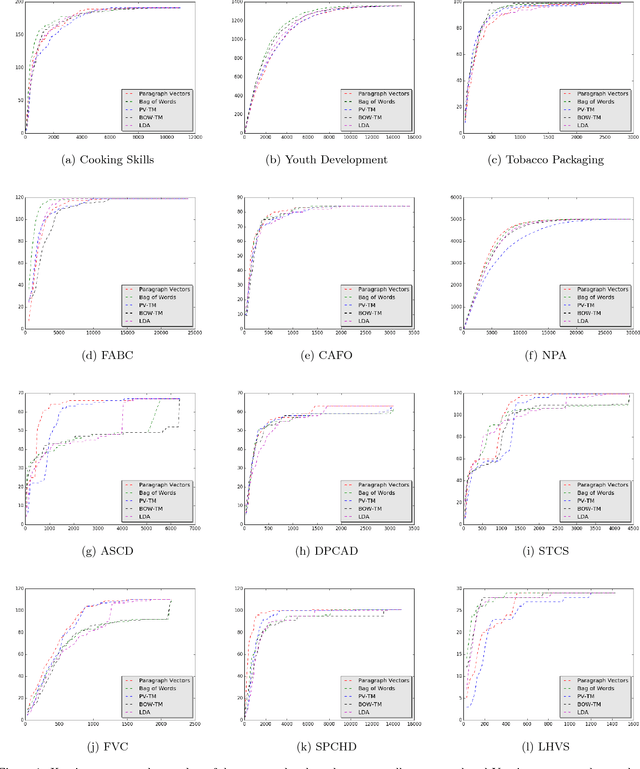
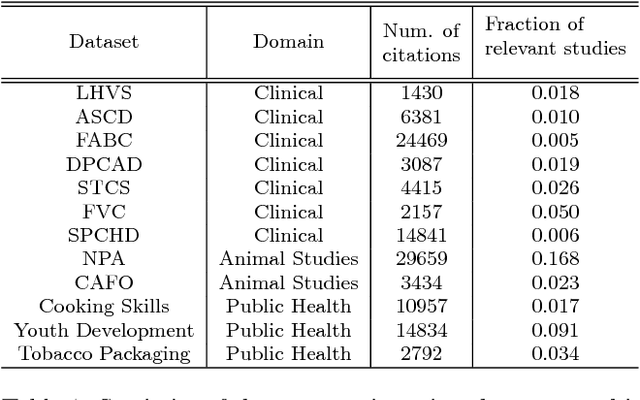
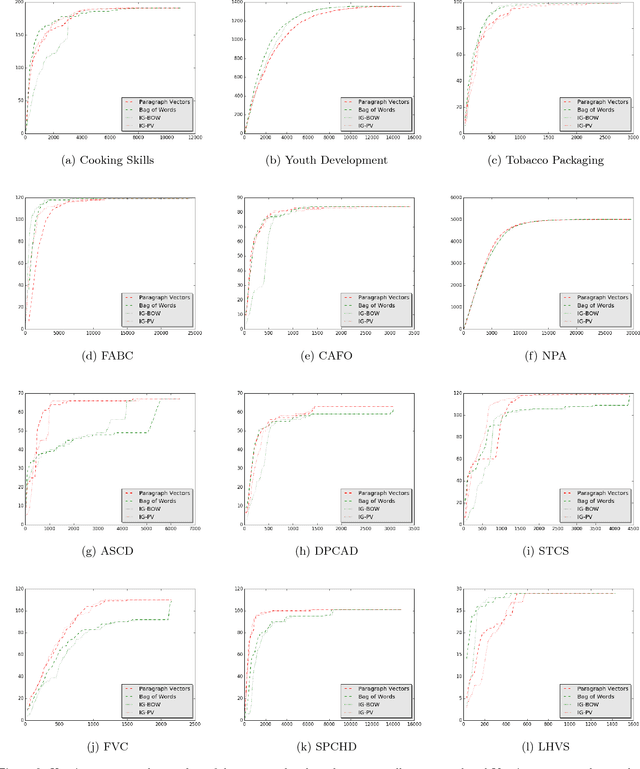
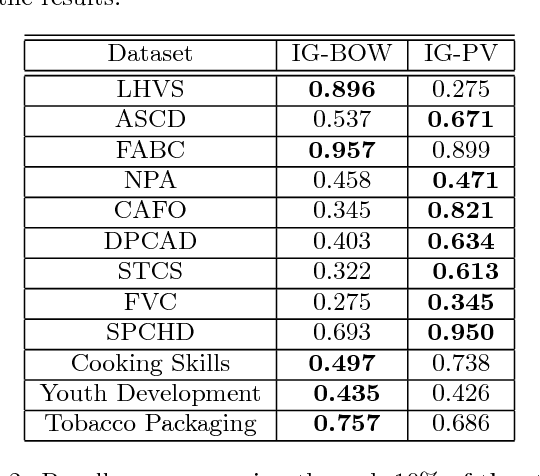
Abstract:Systematic reviews are essential to summarizing the results of different clinical and social science studies. The first step in a systematic review task is to identify all the studies relevant to the review. The task of identifying relevant studies for a given systematic review is usually performed manually, and as a result, involves substantial amounts of expensive human resource. Lately, there have been some attempts to reduce this manual effort using active learning. In this work, we build upon some such existing techniques, and validate by experimenting on a larger and comprehensive dataset than has been attempted until now. Our experiments provide insights on the use of different feature extraction models for different disciplines. More importantly, we identify that a naive active learning based screening process is biased in favour of selecting similar documents. We aimed to improve the performance of the screening process using a novel active learning algorithm with success. Additionally, we propose a mechanism to choose the best feature extraction method for a given review.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge