Jack Baker
Large-Scale Stochastic Sampling from the Probability Simplex
Oct 26, 2018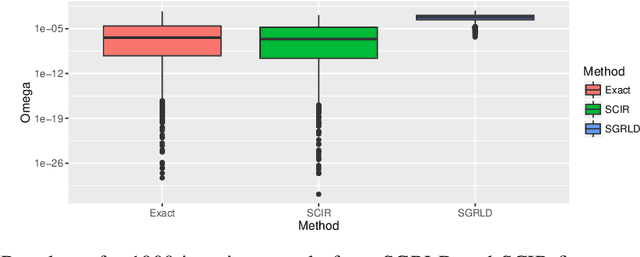
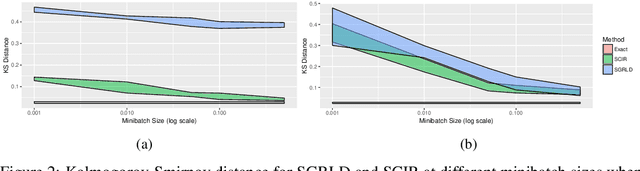
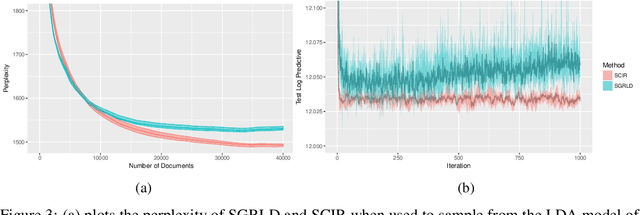
Abstract:Stochastic gradient Markov chain Monte Carlo (SGMCMC) has become a popular method for scalable Bayesian inference. These methods are based on sampling a discrete-time approximation to a continuous time process, such as the Langevin diffusion. When applied to distributions defined on a constrained space the time-discretization error can dominate when we are near the boundary of the space. We demonstrate that because of this, current SGMCMC methods for the simplex struggle with sparse simplex spaces; when many of the components are close to zero. Unfortunately, many popular large-scale Bayesian models, such as network or topic models, require inference on sparse simplex spaces. To avoid the biases caused by this discretization error, we propose the stochastic Cox-Ingersoll-Ross process (SCIR), which removes all discretization error and we prove that samples from the SCIR process are asymptotically unbiased. We discuss how this idea can be extended to target other constrained spaces. Use of the SCIR process within a SGMCMC algorithm is shown to give substantially better performance for a topic model and a Dirichlet process mixture model than existing SGMCMC approaches.
sgmcmc: An R Package for Stochastic Gradient Markov Chain Monte Carlo
Apr 13, 2018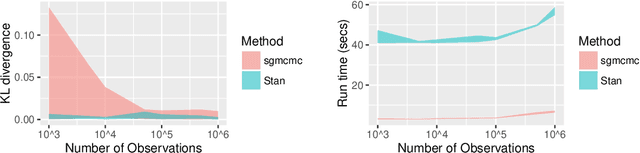
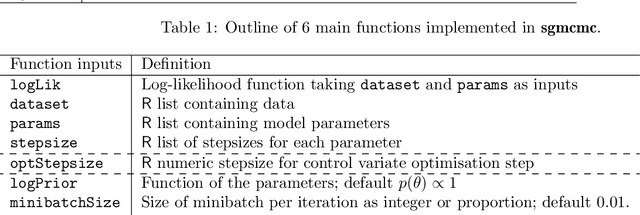


Abstract:This paper introduces the R package sgmcmc; which can be used for Bayesian inference on problems with large datasets using stochastic gradient Markov chain Monte Carlo (SGMCMC). Traditional Markov chain Monte Carlo (MCMC) methods, such as Metropolis-Hastings, are known to run prohibitively slowly as the dataset size increases. SGMCMC solves this issue by only using a subset of data at each iteration. SGMCMC requires calculating gradients of the log likelihood and log priors, which can be time consuming and error prone to perform by hand. The sgmcmc package calculates these gradients itself using automatic differentiation, making the implementation of these methods much easier. To do this, the package uses the software library TensorFlow, which has a variety of statistical distributions and mathematical operations as standard, meaning a wide class of models can be built using this framework. SGMCMC has become widely adopted in the machine learning literature, but less so in the statistics community. We believe this may be partly due to lack of software; this package aims to bridge this gap.
Control Variates for Stochastic Gradient MCMC
Dec 14, 2017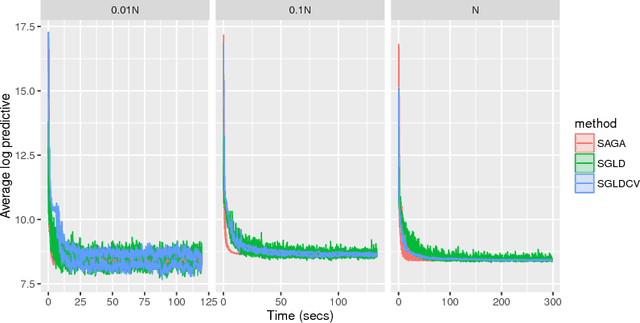
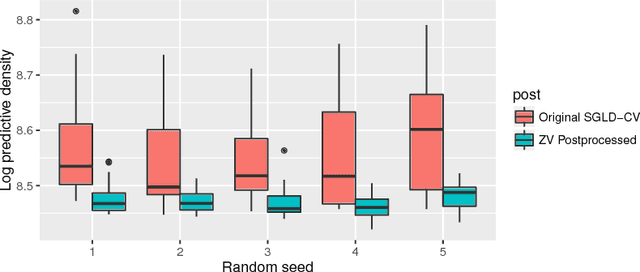
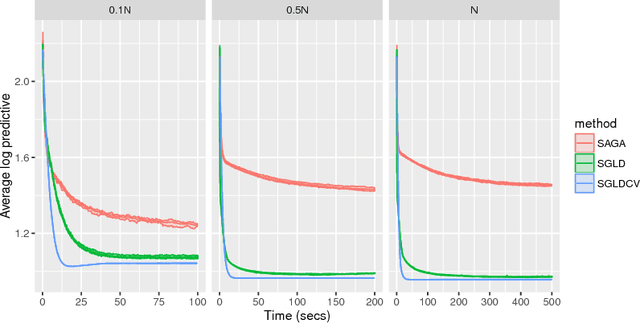
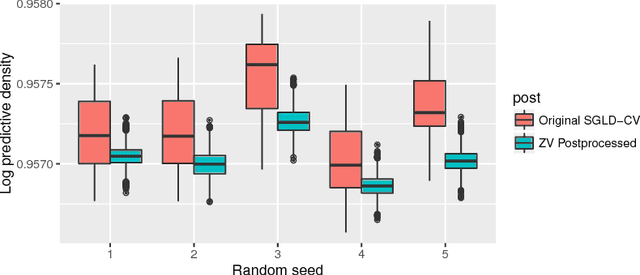
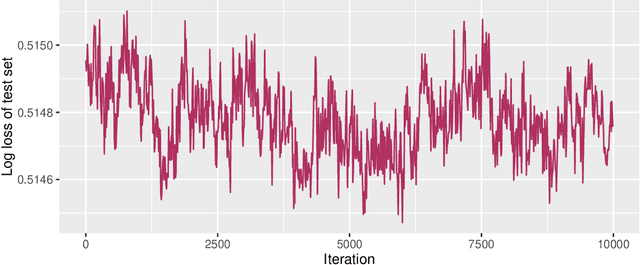
Abstract:It is well known that Markov chain Monte Carlo (MCMC) methods scale poorly with dataset size. A popular class of methods for solving this issue is stochastic gradient MCMC. These methods use a noisy estimate of the gradient of the log posterior, which reduces the per iteration computational cost of the algorithm. Despite this, there are a number of results suggesting that stochastic gradient Langevin dynamics (SGLD), probably the most popular of these methods, still has computational cost proportional to the dataset size. We suggest an alternative log posterior gradient estimate for stochastic gradient MCMC, which uses control variates to reduce the variance. We analyse SGLD using this gradient estimate, and show that, under log-concavity assumptions on the target distribution, the computational cost required for a given level of accuracy is independent of the dataset size. Next we show that a different control variate technique, known as zero variance control variates can be applied to SGMCMC algorithms for free. This post-processing step improves the inference of the algorithm by reducing the variance of the MCMC output. Zero variance control variates rely on the gradient of the log posterior; we explore how the variance reduction is affected by replacing this with the noisy gradient estimate calculated by SGMCMC.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge