Ilya Safro
Clemson University
AGATHA: Automatic Graph-mining And Transformer based Hypothesis generation Approach
Feb 13, 2020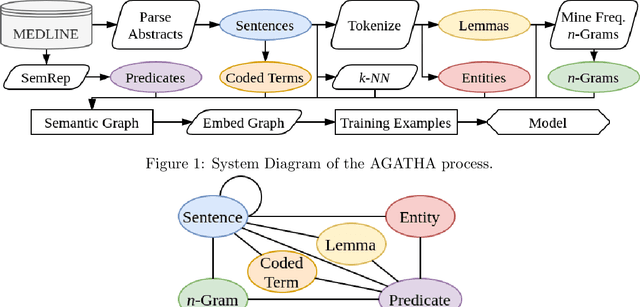
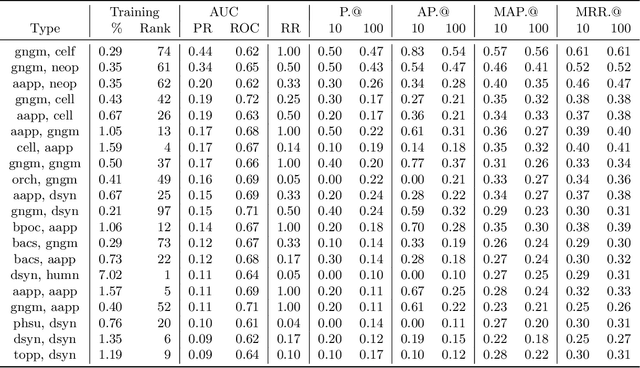
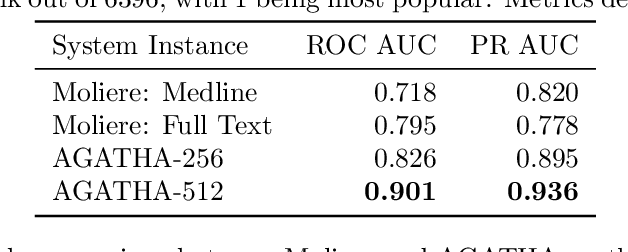
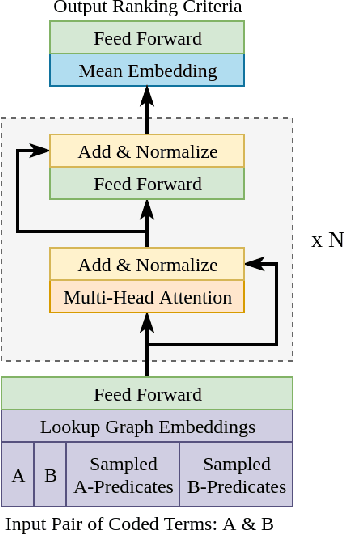
Abstract:Medical research is risky and expensive. Drug discovery, as an example, requires that researchers efficiently winnow thousands of potential targets to a small candidate set for more thorough evaluation. However, research groups spend significant time and money to perform the experiments necessary to determine this candidate set long before seeing intermediate results. Hypothesis generation systems address this challenge by mining the wealth of publicly available scientific information to predict plausible research directions. We present AGATHA, a deep-learning hypothesis generation system that can introduce data-driven insights earlier in the discovery process. Through a learned ranking criteria, this system quickly prioritizes plausible term-pairs among entity sets, allowing us to recommend new research directions. We massively validate our system with a temporal holdout wherein we predict connections first introduced after 2015 using data published beforehand. We additionally explore biomedical sub-domains, and demonstrate AGATHA's predictive capacity across the twenty most popular relationship types. This system achieves best-in-class performance on an established benchmark, and demonstrates high recommendation scores across subdomains. Reproducibility: All code, experimental data, and pre-trained models are available online: sybrandt.com/2020/agatha
Hypergraph Partitioning With Embeddings
Sep 16, 2019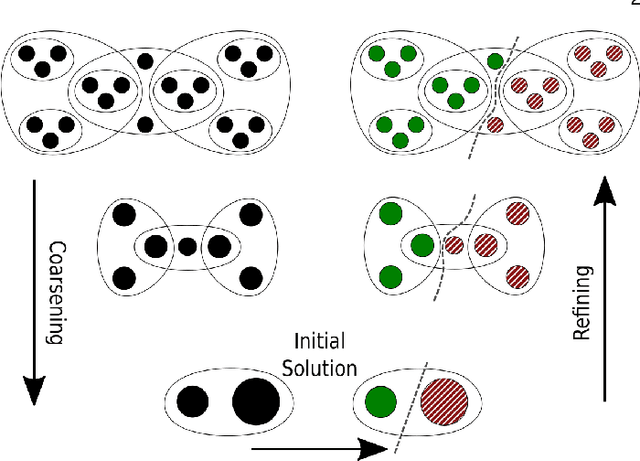
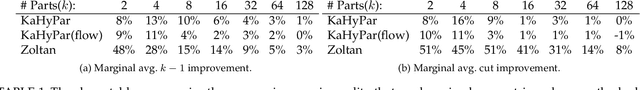
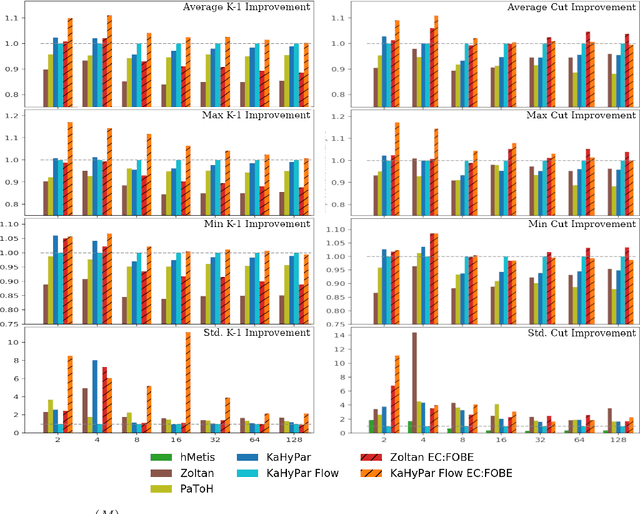
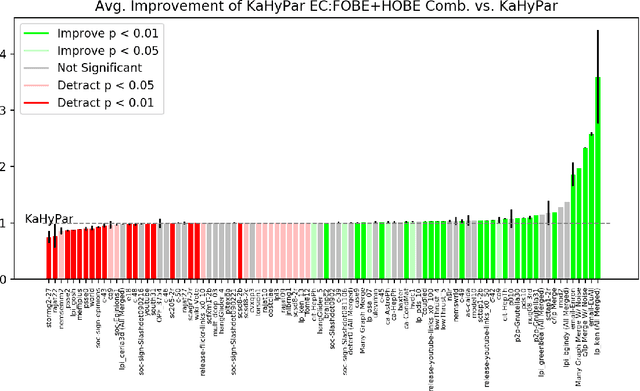
Abstract:The problem of placing circuits on a chip or distributing sparse matrix operations can be modeled as the hypergraph partitioning problem. A hypergraph is a generalization of the traditional graph wherein each "hyperedge" may connect any number of nodes. Hypergraph partitioning, therefore, is the NP-Hard problem of dividing nodes into $k$ similarly sized disjoint sets while minimizing the number of hyperedges that span multiple partitions. Due to this problem's complexity, many partitioners leverage the multilevel heuristic of iteratively "coarsening" their input to a smaller approximation until an inefficient algorithm becomes feasible. The initial solution is then propagated back to the original hypergraph, which produces a reasonably accurate result provided the coarse representation preserves structural properties of the original. The multilevel hypergraph partitioners are considered today as state-of-the-art solvers that achieve an excellent quality/running time trade-off on practical large-scale instances of different types. In order to improve the quality of multilevel hypergraph partitioners, we propose leveraging graph embeddings to better capture structural properties during the coarsening process. Our approach prioritizes dense subspaces found at the embedding, and contracts nodes according to both traditional and embedding-based similarity measures. Reproducibility: All source code, plots and experimental data are available at https://sybrandt.com/2019/partition.
FOBE and HOBE: First- and High-Order Bipartite Embeddings
May 27, 2019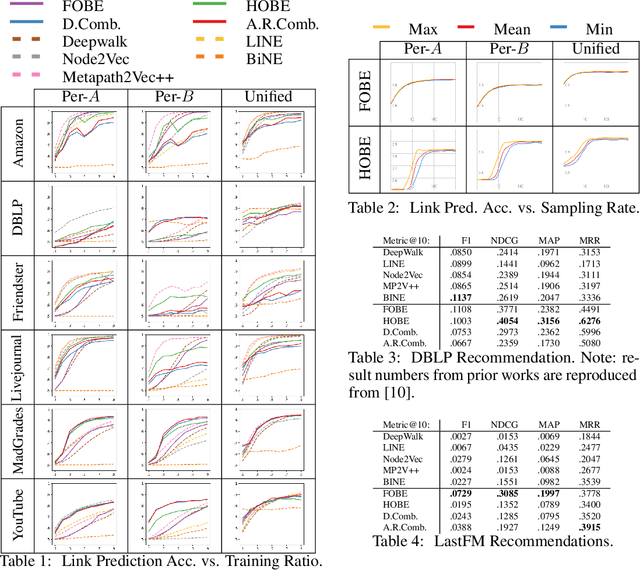
Abstract:Typical graph embeddings may not capture type-specific bipartite graph features that arise in such areas as recommender systems, data visualization, and drug discovery. Machine learning methods utilized in these applications would be better served with specialized embedding techniques. We propose two embeddings for bipartite graphs that decompose edges into sets of indirect relationships between node neighborhoods. When sampling higher-order relationships, we reinforce similarities through algebraic distance on graphs. We also introduce ensemble embeddings to combine both into a "best of both worlds" embedding. The proposed methods are evaluated on link prediction and recommendation tasks and compared with other state-of-the-art embeddings. While being all highly beneficial in applications, we demonstrate that none of the considered embeddings is clearly superior (in contrast to what is claimed in many papers), and discuss the trade offs present among them. Reproducibility: Our code, data sets, and results are all publicly available online at: http://bit.ly/fobe_hobe_code.
Large-Scale Validation of Hypothesis Generation Systems via Candidate Ranking
Oct 19, 2018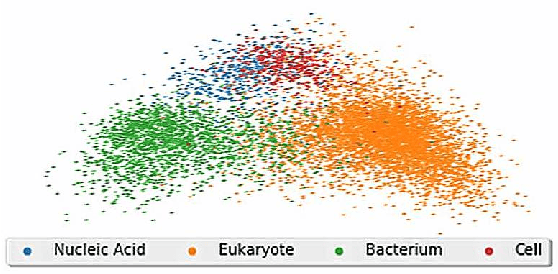
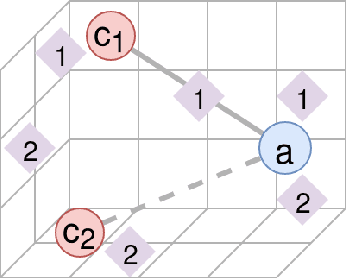
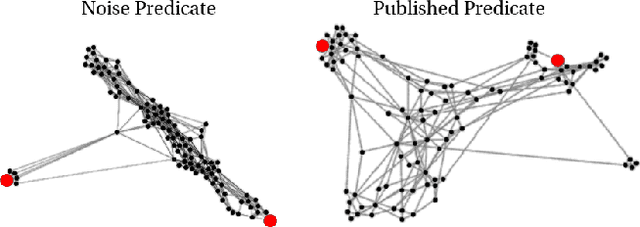
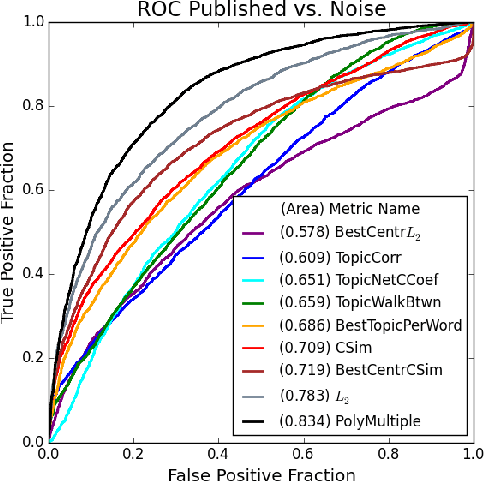
Abstract:The first step of many research projects is to define and rank a short list of candidates for study. In the modern rapidity of scientific progress, some turn to automated hypothesis generation (HG) systems to aid this process. These systems can identify implicit or overlooked connections within a large scientific corpus, and while their importance grows alongside the pace of science, they lack thorough validation. Without any standard numerical evaluation method, many validate general-purpose HG systems by rediscovering a handful of historical findings, and some wishing to be more thorough may run laboratory experiments based on automatic suggestions. These methods are expensive, time consuming, and cannot scale. Thus, we present a numerical evaluation framework for the purpose of validating HG systems that leverages thousands of validation hypotheses. This method evaluates a HG system by its ability to rank hypotheses by plausibility; a process reminiscent of human candidate selection. Because HG systems do not produce a ranking criteria, specifically those that produce topic models, we additionally present novel metrics to quantify the plausibility of hypotheses given topic model system output. Finally, we demonstrate that our proposed validation method aligns with real-world research goals by deploying our method within Moliere, our recent topic-driven HG system, in order to automatically generate a set of candidate genes related to HIV-associated neurodegenerative disease (HAND). By performing laboratory experiments based on this candidate set, we discover a new connection between HAND and Dead Box RNA Helicase 3 (DDX3). Reproducibility: code, validation data, and results can be found at sybrandt.com/2018/validation.
Scalable Dynamic Topic Modeling with Clustered Latent Dirichlet Allocation (CLDA)
Oct 15, 2017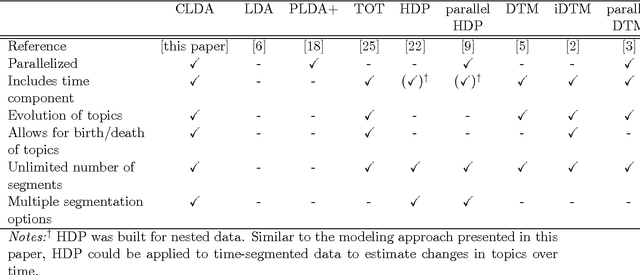
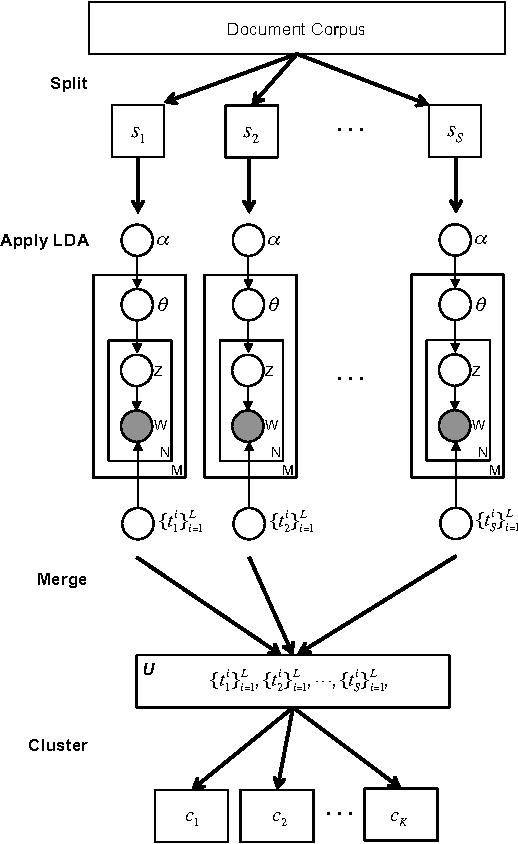
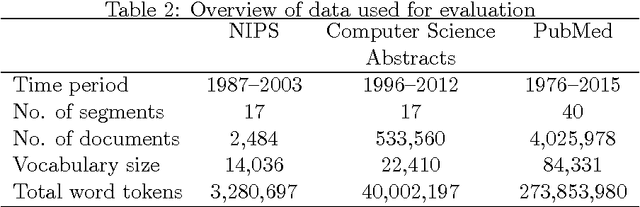
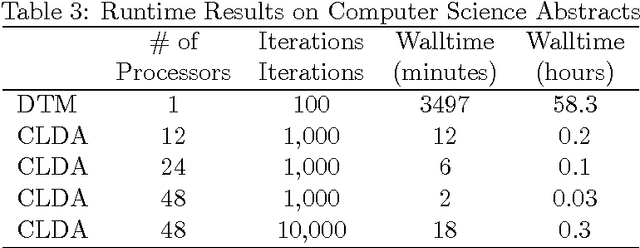
Abstract:Topic modeling, a method for extracting the underlying themes from a collection of documents, is an increasingly important component of the design of intelligent systems enabling the sense-making of highly dynamic and diverse streams of text data. Traditional methods such as Dynamic Topic Modeling (DTM) do not lend themselves well to direct parallelization because of dependencies from one time step to another. In this paper, we introduce and empirically analyze Clustered Latent Dirichlet Allocation (CLDA), a method for extracting dynamic latent topics from a collection of documents. Our approach is based on data decomposition in which the data is partitioned into segments, followed by topic modeling on the individual segments. The resulting local models are then combined into a global solution using clustering. The decomposition and resulting parallelization leads to very fast runtime even on very large datasets. Our approach furthermore provides insight into how the composition of topics changes over time and can also be applied using other data partitioning strategies over any discrete features of the data, such as geographic features or classes of users. In this paper CLDA is applied successfully to seventeen years of NIPS conference papers (2,484 documents and 3,280,697 words), seventeen years of computer science journal abstracts (533,560 documents and 32,551,540 words), and to forty years of the PubMed corpus (4,025,978 documents and 273,853,980 words).
Algebraic multigrid support vector machines
Nov 24, 2016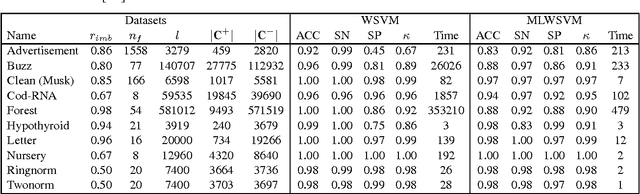
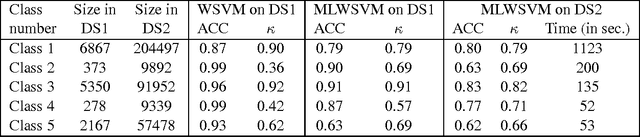
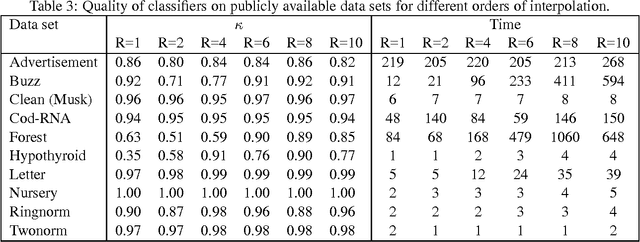
Abstract:The support vector machine is a flexible optimization-based technique widely used for classification problems. In practice, its training part becomes computationally expensive on large-scale data sets because of such reasons as the complexity and number of iterations in parameter fitting methods, underlying optimization solvers, and nonlinearity of kernels. We introduce a fast multilevel framework for solving support vector machine models that is inspired by the algebraic multigrid. Significant improvement in the running has been achieved without any loss in the quality. The proposed technique is highly beneficial on imbalanced sets. We demonstrate computational results on publicly available and industrial data sets.
Multilevel Weighted Support Vector Machine for Classification on Healthcare Data with Missing Values
Apr 07, 2016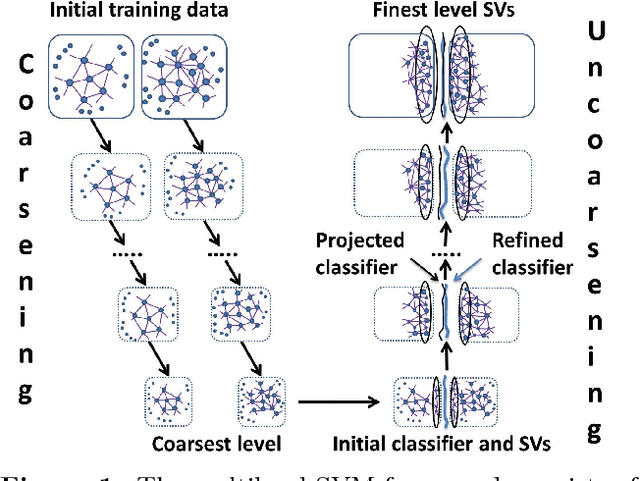
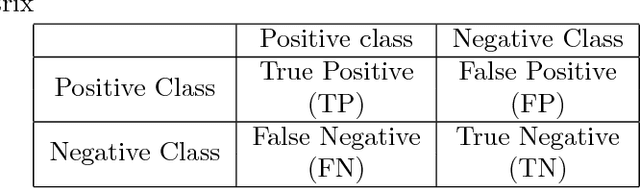
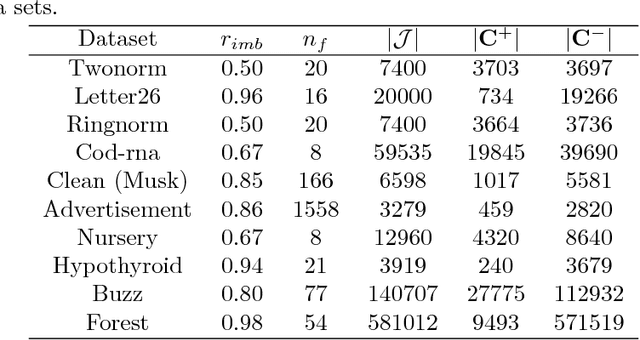
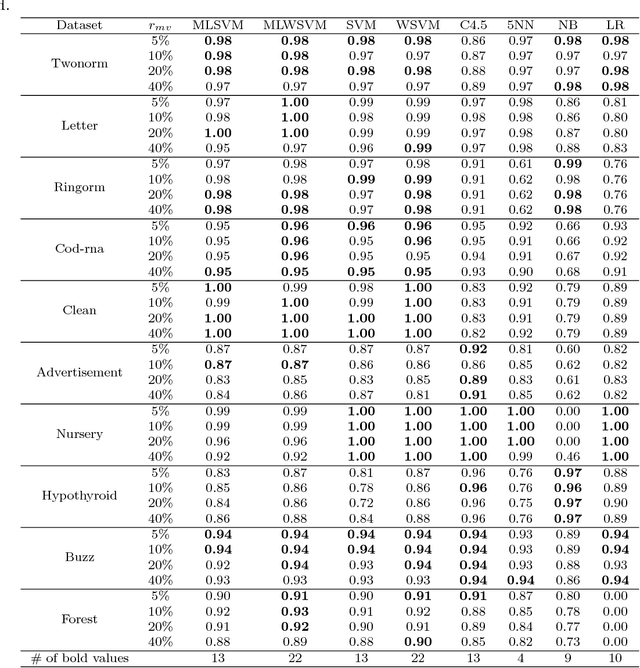
Abstract:This work is motivated by the needs of predictive analytics on healthcare data as represented by Electronic Medical Records. Such data is invariably problematic: noisy, with missing entries, with imbalance in classes of interests, leading to serious bias in predictive modeling. Since standard data mining methods often produce poor performance measures, we argue for development of specialized techniques of data-preprocessing and classification. In this paper, we propose a new method to simultaneously classify large datasets and reduce the effects of missing values. It is based on a multilevel framework of the cost-sensitive SVM and the expected maximization imputation method for missing values, which relies on iterated regression analyses. We compare classification results of multilevel SVM-based algorithms on public benchmark datasets with imbalanced classes and missing values as well as real data in health applications, and show that our multilevel SVM-based method produces fast, and more accurate and robust classification results.
Fast Imbalanced Classification of Healthcare Data with Missing Values
Mar 21, 2015
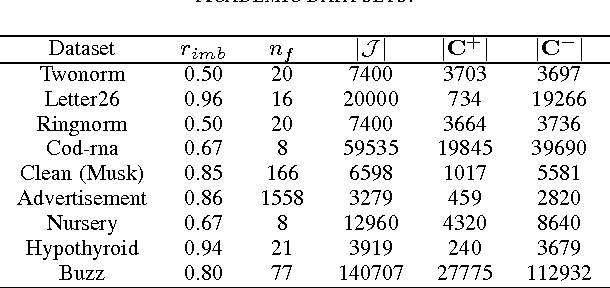
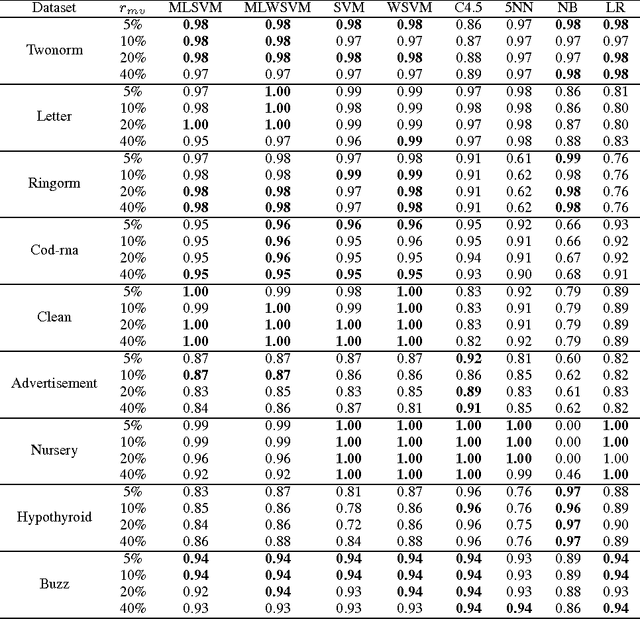
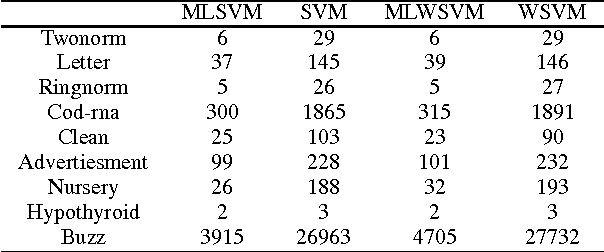
Abstract:In medical domain, data features often contain missing values. This can create serious bias in the predictive modeling. Typical standard data mining methods often produce poor performance measures. In this paper, we propose a new method to simultaneously classify large datasets and reduce the effects of missing values. The proposed method is based on a multilevel framework of the cost-sensitive SVM and the expected maximization imputation method for missing values, which relies on iterated regression analyses. We compare classification results of multilevel SVM-based algorithms on public benchmark datasets with imbalanced classes and missing values as well as real data in health applications, and show that our multilevel SVM-based method produces fast, and more accurate and robust classification results.
Fast Multilevel Support Vector Machines
Oct 13, 2014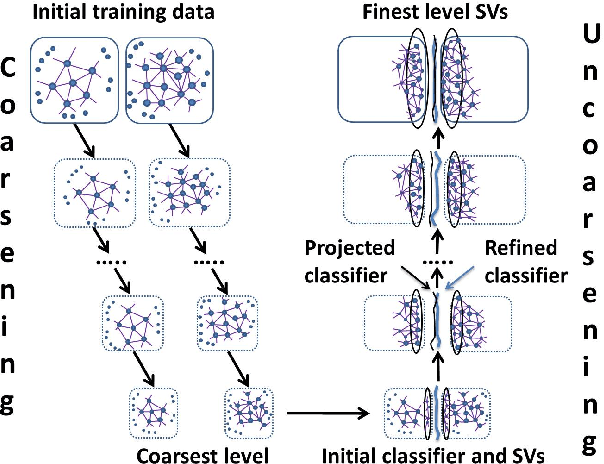
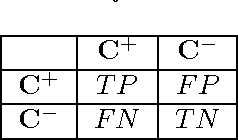
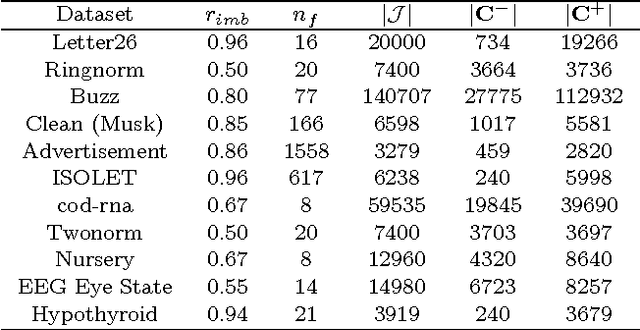
Abstract:Solving different types of optimization models (including parameters fitting) for support vector machines on large-scale training data is often an expensive computational task. This paper proposes a multilevel algorithmic framework that scales efficiently to very large data sets. Instead of solving the whole training set in one optimization process, the support vectors are obtained and gradually refined at multiple levels of coarseness of the data. The proposed framework includes: (a) construction of hierarchy of large-scale data coarse representations, and (b) a local processing of updating the hyperplane throughout this hierarchy. Our multilevel framework substantially improves the computational time without loosing the quality of classifiers. The algorithms are demonstrated for both regular and weighted support vector machines. Experimental results are presented for balanced and imbalanced classification problems. Quality improvement on several imbalanced data sets has been observed.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge