Hesham Elhalawani
Analysis of the MICCAI Brain Tumor Segmentation -- Metastases (BraTS-METS) 2025 Lighthouse Challenge: Brain Metastasis Segmentation on Pre- and Post-treatment MRI
Apr 16, 2025Abstract:Despite continuous advancements in cancer treatment, brain metastatic disease remains a significant complication of primary cancer and is associated with an unfavorable prognosis. One approach for improving diagnosis, management, and outcomes is to implement algorithms based on artificial intelligence for the automated segmentation of both pre- and post-treatment MRI brain images. Such algorithms rely on volumetric criteria for lesion identification and treatment response assessment, which are still not available in clinical practice. Therefore, it is critical to establish tools for rapid volumetric segmentations methods that can be translated to clinical practice and that are trained on high quality annotated data. The BraTS-METS 2025 Lighthouse Challenge aims to address this critical need by establishing inter-rater and intra-rater variability in dataset annotation by generating high quality annotated datasets from four individual instances of segmentation by neuroradiologists while being recorded on video (two instances doing "from scratch" and two instances after AI pre-segmentation). This high-quality annotated dataset will be used for testing phase in 2025 Lighthouse challenge and will be publicly released at the completion of the challenge. The 2025 Lighthouse challenge will also release the 2023 and 2024 segmented datasets that were annotated using an established pipeline of pre-segmentation, student annotation, two neuroradiologists checking, and one neuroradiologist finalizing the process. It builds upon its previous edition by including post-treatment cases in the dataset. Using these high-quality annotated datasets, the 2025 Lighthouse challenge plans to test benchmark algorithms for automated segmentation of pre-and post-treatment brain metastases (BM), trained on diverse and multi-institutional datasets of MRI images obtained from patients with brain metastases.
The impact of using an AI chatbot to respond to patient messages
Oct 26, 2023



Abstract:Documentation burden is a major contributor to clinician burnout, which is rising nationally and is an urgent threat to our ability to care for patients. Artificial intelligence (AI) chatbots, such as ChatGPT, could reduce clinician burden by assisting with documentation. Although many hospitals are actively integrating such systems into electronic medical record systems, AI chatbots utility and impact on clinical decision-making have not been studied for this intended use. We are the first to examine the utility of large language models in assisting clinicians draft responses to patient questions. In our two-stage cross-sectional study, 6 oncologists responded to 100 realistic synthetic cancer patient scenarios and portal messages developed to reflect common medical situations, first manually, then with AI assistance. We find AI-assisted responses were longer, less readable, but provided acceptable drafts without edits 58% of time. AI assistance improved efficiency 77% of time, with low harm risk (82% safe). However, 7.7% unedited AI responses could severely harm. In 31% cases, physicians thought AI drafts were human-written. AI assistance led to more patient education recommendations, fewer clinical actions than manual responses. Results show promise for AI to improve clinician efficiency and patient care through assisting documentation, if used judiciously. Monitoring model outputs and human-AI interaction remains crucial for safe implementation.
Overview of the HECKTOR Challenge at MICCAI 2021: Automatic Head and Neck Tumor Segmentation and Outcome Prediction in PET/CT Images
Jan 11, 2022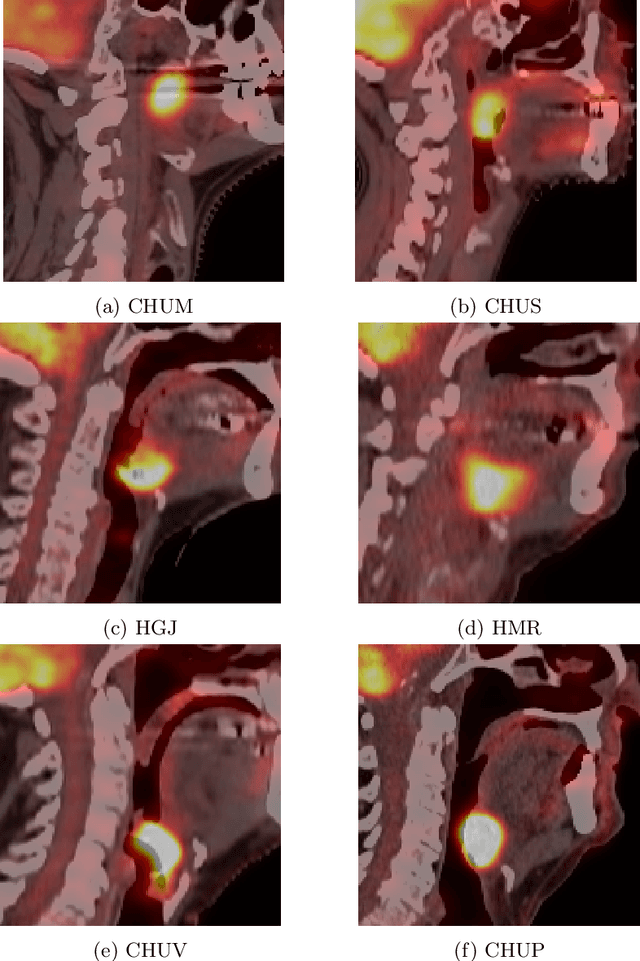
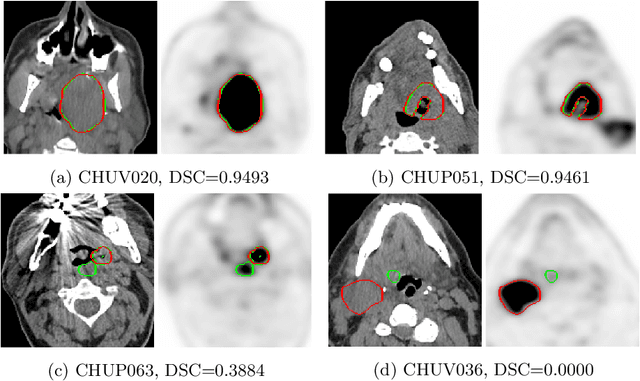
Abstract:This paper presents an overview of the second edition of the HEad and neCK TumOR (HECKTOR) challenge, organized as a satellite event of the 24th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) 2021. The challenge is composed of three tasks related to the automatic analysis of PET/CT images for patients with Head and Neck cancer (H&N), focusing on the oropharynx region. Task 1 is the automatic segmentation of H&N primary Gross Tumor Volume (GTVt) in FDG-PET/CT images. Task 2 is the automatic prediction of Progression Free Survival (PFS) from the same FDG-PET/CT. Finally, Task 3 is the same as Task 2 with ground truth GTVt annotations provided to the participants. The data were collected from six centers for a total of 325 images, split into 224 training and 101 testing cases. The interest in the challenge was highlighted by the important participation with 103 registered teams and 448 result submissions. The best methods obtained a Dice Similarity Coefficient (DSC) of 0.7591 in the first task, and a Concordance index (C-index) of 0.7196 and 0.6978 in Tasks 2 and 3, respectively. In all tasks, simplicity of the approach was found to be key to ensure generalization performance. The comparison of the PFS prediction performance in Tasks 2 and 3 suggests that providing the GTVt contour was not crucial to achieve best results, which indicates that fully automatic methods can be used. This potentially obviates the need for GTVt contouring, opening avenues for reproducible and large scale radiomics studies including thousands potential subjects.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge