Han Joo Chae
A Guideline-Aware AI Agent for Zero-Shot Target Volume Auto-Delineation
Mar 10, 2026Abstract:Delineating the clinical target volume (CTV) in radiotherapy involves complex margins constrained by tumor location and anatomical barriers. While deep learning models automate this process, their rigid reliance on expert-annotated data requires costly retraining whenever clinical guidelines update. To overcome this limitation, we introduce OncoAgent, a novel guideline-aware AI agent framework that seamlessly converts textual clinical guidelines into three-dimensional target contours in a training-free manner. Evaluated on esophageal cancer cases, the agent achieves a zero-shot Dice similarity coefficient of 0.842 for the CTV and 0.880 for the planning target volume, demonstrating performance highly comparable to a fully supervised nnU-Net baseline. Notably, in a blinded clinical evaluation, physicians strongly preferred OncoAgent over the supervised baseline, rating it higher in guideline compliance, modification effort, and clinical acceptability. Furthermore, the framework generalizes zero-shot to alternative esophageal guidelines and other anatomical sites (e.g., prostate) without any retraining. Beyond mere volumetric overlap, our agent-based paradigm offers near-instantaneous adaptability to alternative guidelines, providing a scalable and transparent pathway toward interpretability in radiotherapy treatment planning.
FastSurf: Fast Neural RGB-D Surface Reconstruction using Per-Frame Intrinsic Refinement and TSDF Fusion Prior Learning
Mar 08, 2023



Abstract:We introduce FastSurf, an accelerated neural radiance field (NeRF) framework that incorporates depth information for 3D reconstruction. A dense feature grid and shallow multi-layer perceptron are used for fast and accurate surface optimization of the entire scene. Our per-frame intrinsic refinement scheme corrects the frame-specific errors that cannot be handled by global optimization. Furthermore, FastSurf utilizes a classical real-time 3D surface reconstruction method, the truncated signed distance field (TSDF) Fusion, as prior knowledge to pretrain the feature grid to accelerate the training. The quantitative and qualitative experiments comparing the performances of FastSurf against prior work indicate that our method is capable of quickly and accurately reconstructing a scene with high-frequency details. We also demonstrate the effectiveness of our per-frame intrinsic refinement and TSDF Fusion prior learning techniques via an ablation study.
Generating 3D Bio-Printable Patches Using Wound Segmentation and Reconstruction to Treat Diabetic Foot Ulcers
Mar 08, 2022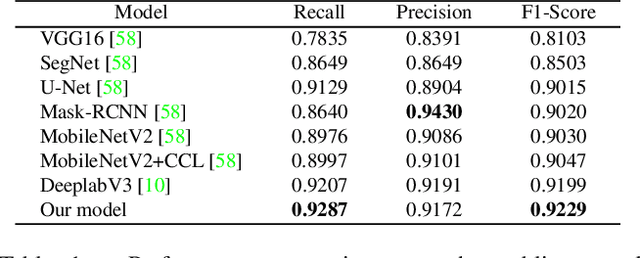
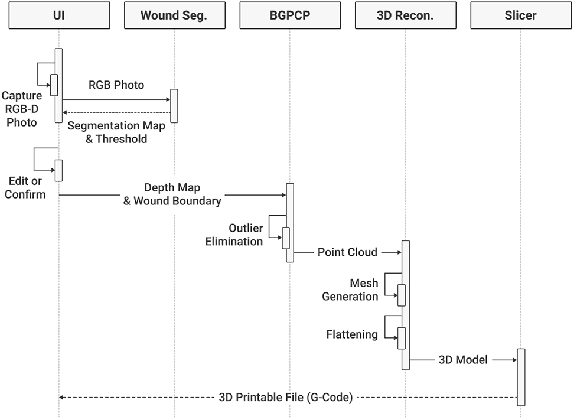
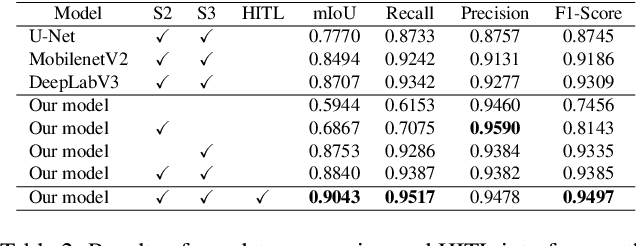
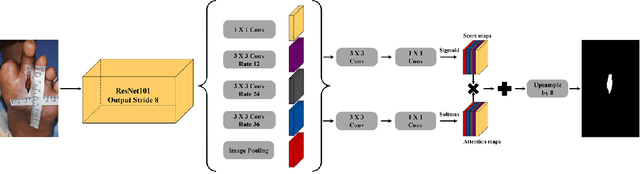
Abstract:We introduce AiD Regen, a novel system that generates 3D wound models combining 2D semantic segmentation with 3D reconstruction so that they can be printed via 3D bio-printers during the surgery to treat diabetic foot ulcers (DFUs). AiD Regen seamlessly binds the full pipeline, which includes RGB-D image capturing, semantic segmentation, boundary-guided point-cloud processing, 3D model reconstruction, and 3D printable G-code generation, into a single system that can be used out of the box. We developed a multi-stage data preprocessing method to handle small and unbalanced DFU image datasets. AiD Regen's human-in-the-loop machine learning interface enables clinicians to not only create 3D regenerative patches with just a few touch interactions but also customize and confirm wound boundaries. As evidenced by our experiments, our model outperforms prior wound segmentation models and our reconstruction algorithm is capable of generating 3D wound models with compelling accuracy. We further conducted a case study on a real DFU patient and demonstrated the effectiveness of AiD Regen in treating DFU wounds.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge