Giorgio Maria Di Nunzio
A Domain-Specific Curated Benchmark for Entity and Document-Level Relation Extraction
Feb 04, 2026Abstract:Information Extraction (IE), encompassing Named Entity Recognition (NER), Named Entity Linking (NEL), and Relation Extraction (RE), is critical for transforming the rapidly growing volume of scientific publications into structured, actionable knowledge. This need is especially evident in fast-evolving biomedical fields such as the gut-brain axis, where research investigates complex interactions between the gut microbiota and brain-related disorders. Existing biomedical IE benchmarks, however, are often narrow in scope and rely heavily on distantly supervised or automatically generated annotations, limiting their utility for advancing robust IE methods. We introduce GutBrainIE, a benchmark based on more than 1,600 PubMed abstracts, manually annotated by biomedical and terminological experts with fine-grained entities, concept-level links, and relations. While grounded in the gut-brain axis, the benchmark's rich schema, multiple tasks, and combination of highly curated and weakly supervised data make it broadly applicable to the development and evaluation of biomedical IE systems across domains.
Towards an automatic recognition of mixed languages: The Ukrainian-Russian hybrid language Surzhyk
Dec 18, 2019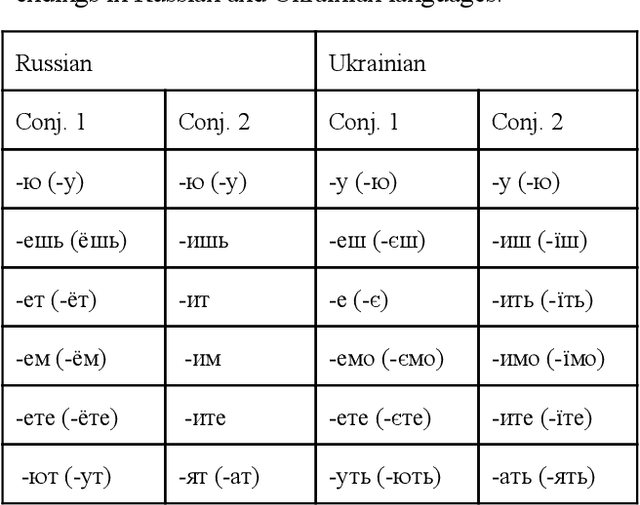


Abstract:Language interference is common in today's multilingual societies where more languages are being in contact and as a global final result leads to the creation of hybrid languages. These, together with doubts on their right to be officially recognised made emerge in the area of computational linguistics the problem of their automatic identification and further elaboration. In this paper, we propose a first attempt to identify the elements of a Ukrainian-Russian hybrid language, Surzhyk, through the adoption of the example-based rules created with the instruments of programming language R. Our example-based study consists of: 1) analysis of spoken samples of Surzhyk registered by Del Gaudio (2010) in Kyiv area and creation of the written corpus; 2) production of specific rules on the identification of Surzhyk patterns and their implementation; 3) testing the code and analysing the effectiveness.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge