Fons J. Verbeek
Using Machine Learning for Lunar Mineralogy-I: Hyperspectral Imaging of Volcanic Samples
Mar 28, 2025Abstract:This study examines the mineral composition of volcanic samples similar to lunar materials, focusing on olivine and pyroxene. Using hyperspectral imaging from 400 to 1000 nm, we created data cubes to analyze the reflectance characteristics of samples from samples from Vulcano, a volcanically active island in the Aeolian Archipelago, north of Sicily, Italy, categorizing them into nine regions of interest and analyzing spectral data for each. We applied various unsupervised clustering algorithms, including K-Means, Hierarchical Clustering, GMM, and Spectral Clustering, to classify the spectral profiles. Principal Component Analysis revealed distinct spectral signatures associated with specific minerals, facilitating precise identification. Clustering performance varied by region, with K-Means achieving the highest silhouette-score of 0.47, whereas GMM performed poorly with a score of only 0.25. Non-negative Matrix Factorization aided in identifying similarities among clusters across different methods and reference spectra for olivine and pyroxene. Hierarchical clustering emerged as the most reliable technique, achieving a 94\% similarity with the olivine spectrum in one sample, whereas GMM exhibited notable variability. Overall, the analysis indicated that both Hierarchical and K-Means methods yielded lower errors in total measurements, with K-Means demonstrating superior performance in estimated dispersion and clustering. Additionally, GMM showed a higher root mean square error compared to the other models. The RMSE analysis confirmed K-Means as the most consistent algorithm across all samples, suggesting a predominance of olivine in the Vulcano region relative to pyroxene. This predominance is likely linked to historical formation conditions similar to volcanic processes on the Moon, where olivine-rich compositions are common in ancient lava flows and impact melt rocks.
Advancing Machine Learning for Stellar Activity and Exoplanet Period Rotation
Sep 09, 2024



Abstract:This study applied machine learning models to estimate stellar rotation periods from corrected light curve data obtained by the NASA Kepler mission. Traditional methods often struggle to estimate rotation periods accurately due to noise and variability in the light curve data. The workflow involved using initial period estimates from the LS-Periodogram and Transit Least Squares techniques, followed by splitting the data into training, validation, and testing sets. We employed several machine learning algorithms, including Decision Tree, Random Forest, K-Nearest Neighbors, and Gradient Boosting, and also utilized a Voting Ensemble approach to improve prediction accuracy and robustness. The analysis included data from multiple Kepler IDs, providing detailed metrics on orbital periods and planet radii. Performance evaluation showed that the Voting Ensemble model yielded the most accurate results, with an RMSE approximately 50\% lower than the Decision Tree model and 17\% better than the K-Nearest Neighbors model. The Random Forest model performed comparably to the Voting Ensemble, indicating high accuracy. In contrast, the Gradient Boosting model exhibited a worse RMSE compared to the other approaches. Comparisons of the predicted rotation periods to the photometric reference periods showed close alignment, suggesting the machine learning models achieved high prediction accuracy. The results indicate that machine learning, particularly ensemble methods, can effectively solve the problem of accurately estimating stellar rotation periods, with significant implications for advancing the study of exoplanets and stellar astrophysics.
Advances in Kidney Biopsy Structural Assessment through Dense Instance Segmentation
Sep 29, 2023



Abstract:The kidney biopsy is the gold standard for the diagnosis of kidney diseases. Lesion scores made by expert renal pathologists are semi-quantitative and suffer from high inter-observer variability. Automatically obtaining statistics per segmented anatomical object, therefore, can bring significant benefits in reducing labor and this inter-observer variability. Instance segmentation for a biopsy, however, has been a challenging problem due to (a) the on average large number (around 300 to 1000) of densely touching anatomical structures, (b) with multiple classes (at least 3) and (c) in different sizes and shapes. The currently used instance segmentation models cannot simultaneously deal with these challenges in an efficient yet generic manner. In this paper, we propose the first anchor-free instance segmentation model that combines diffusion models, transformer modules, and RCNNs (regional convolution neural networks). Our model is trained on just one NVIDIA GeForce RTX 3090 GPU, but can efficiently recognize more than 500 objects with 3 common anatomical object classes in renal biopsies, i.e., glomeruli, tubuli, and arteries. Our data set consisted of 303 patches extracted from 148 Jones' silver-stained renal whole slide images (WSIs), where 249 patches were used for training and 54 patches for evaluation. In addition, without adjustment or retraining, the model can directly transfer its domain to generate decent instance segmentation results from PAS-stained WSIs. Importantly, it outperforms other baseline models and reaches an AP 51.7% in detection as the new state-of-the-art.
Detection of anomalously emitting ships through deviations from predicted TROPOMI NO2 retrievals
Feb 24, 2023



Abstract:Starting from 2021, more demanding $\text{NO}_\text{x}$ emission restrictions were introduced for ships operating in the North and Baltic Sea waters. Since all methods currently used for ship compliance monitoring are financially and time demanding, it is important to prioritize the inspection of ships that have high chances of being non-compliant. The current state-of-the-art approach for a large-scale ship $\text{NO}_\text{2}$ estimation is a supervised machine learning-based segmentation of ship plumes on TROPOMI images. However, challenging data annotation and insufficiently complex ship emission proxy used for the validation limit the applicability of the model for ship compliance monitoring. In this study, we present a method for the automated selection of potentially non-compliant ships using a combination of machine learning models on TROPOMI/S5P satellite data. It is based on a proposed regression model predicting the amount of $\text{NO}_\text{2}$ that is expected to be produced by a ship with certain properties operating in the given atmospheric conditions. The model does not require manual labeling and is validated with TROPOMI data directly. The differences between the predicted and actual amount of produced $\text{NO}_\text{2}$ are integrated over different observations of the same ship in time and are used as a measure of the inspection worthiness of a ship. To assure the robustness of the results, we compare the obtained results with the results of the previously developed segmentation-based method. Ships that are also highly deviating in accordance with the segmentation method require further attention. If no other explanations can be found by checking the TROPOMI data, the respective ships are advised to be the candidates for inspection.
Supervised segmentation of NO2 plumes from individual ships using TROPOMI satellite data
Mar 14, 2022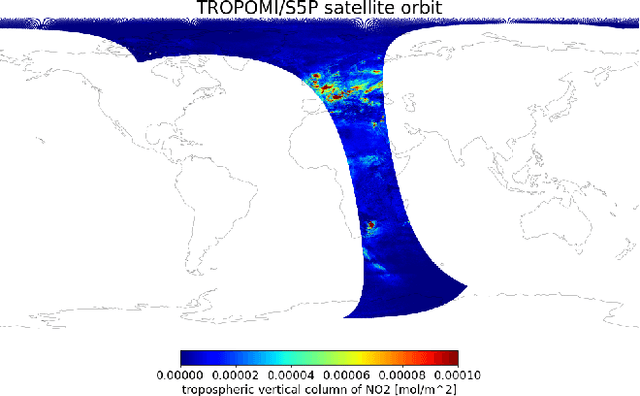
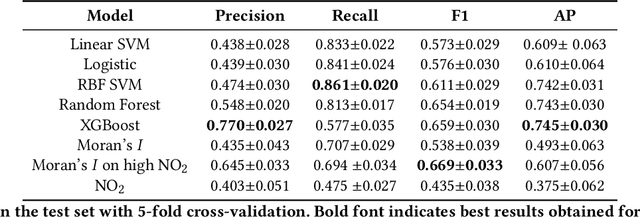
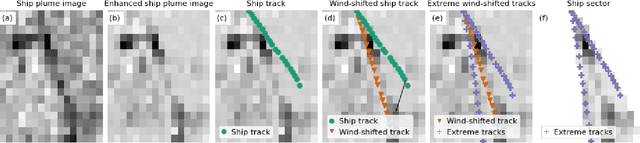

Abstract:Starting from 2021, the International Maritime Organization significantly tightened the $\text{NO}_\text{x}$ emission requirements for ships entering the Baltic and North Sea waters. Since all methods currently used for the ships' compliance monitoring are costly and require proximity to the ship, the performance of a global and continuous monitoring of the emission standards' fulfillment has been impossible up to now. A promising approach is the use of remote sensing with the recently launched TROPOMI/S5P satellite. Due to its unprecedentedly high spatial resolution, it allows for the visual distinction of $\text{NO}_\text{2}$ plumes of individual ships. To successfully deploy a compliance monitoring system that is based on TROPOMI data, an automated procedure for the attribution of $\text{NO}_\text{2}$ to individual ships has to be developed. However, due to the extremely low signal-to-noise ratio, interference with the signal from other - often stronger - sources, and the absence of ground truth, the task is very challenging. In this study, we present an automated method for segmentation of plumes produced by individual ships using TROPOMI satellite data - a first step towards the automated procedure for global ship compliance monitoring. We develop a multivariate plume segmentation method based on various ships', wind's and spatial properties. For this, we propose to automatically define a region of interest - a ship sector that we normalize with respect to scale and orientation. We create a dataset, where each pixel has a label for belonging to the respective ship plume or not. We train five linear and nonlinear classifiers. The results show a significant improvement over the threshold-based baselines. Moreover, the aggregated $\text{NO}_\text{2}$ levels of the segmented plumes show high correlation with the theoretically derived measure of ship's emission potential.
Probabilistic Inference for Camera Calibration in Light Microscopy under Circular Motion
Oct 30, 2019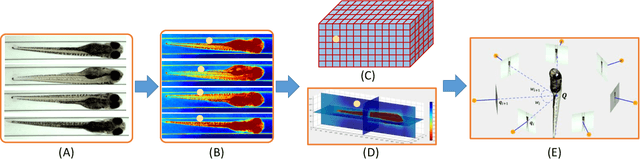
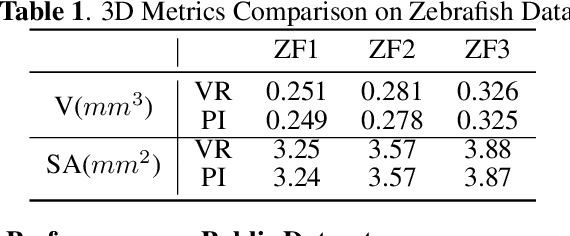
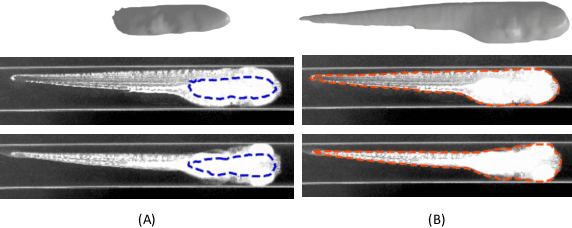
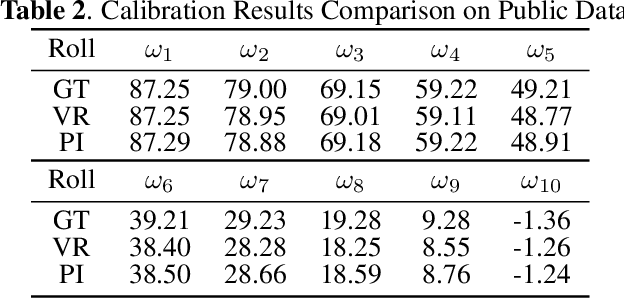
Abstract:Robust and accurate camera calibration is essential for 3D reconstruction in light microscopy under circular motion. Conventional methods require either accurate key point matching or precise segmentation of the axial-view images. Both remain challenging because specimens often exhibit transparency/translucency in a light microscope. To address those issues, we propose a probabilistic inference based method for the camera calibration that does not require sophisticated image pre-processing. Based on 3D projective geometry, our method assigns a probability on each of a range of voxels that cover the whole object. The probability indicates the likelihood of a voxel belonging to the object to be reconstructed. Our method maximizes a joint probability that distinguishes the object from the background. Experimental results show that the proposed method can accurately recover camera configurations in both light microscopy and natural scene imaging. Furthermore, the method can be used to produce high-fidelity 3D reconstructions and accurate 3D measurements.
Creating a new Ontology: a Modular Approach
Dec 08, 2010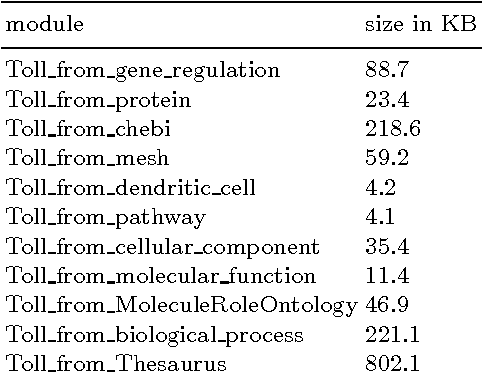
Abstract:Creating a new Ontology: a Modular Approach
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge