Evan M. Yu
A robust and interpretable deep learning framework for multi-modal registration via keypoints
Apr 19, 2023Abstract:We present KeyMorph, a deep learning-based image registration framework that relies on automatically detecting corresponding keypoints. State-of-the-art deep learning methods for registration often are not robust to large misalignments, are not interpretable, and do not incorporate the symmetries of the problem. In addition, most models produce only a single prediction at test-time. Our core insight which addresses these shortcomings is that corresponding keypoints between images can be used to obtain the optimal transformation via a differentiable closed-form expression. We use this observation to drive the end-to-end learning of keypoints tailored for the registration task, and without knowledge of ground-truth keypoints. This framework not only leads to substantially more robust registration but also yields better interpretability, since the keypoints reveal which parts of the image are driving the final alignment. Moreover, KeyMorph can be designed to be equivariant under image translations and/or symmetric with respect to the input image ordering. Finally, we show how multiple deformation fields can be computed efficiently and in closed-form at test time corresponding to different transformation variants. We demonstrate the proposed framework in solving 3D affine and spline-based registration of multi-modal brain MRI scans. In particular, we show registration accuracy that surpasses current state-of-the-art methods, especially in the context of large displacements. Our code is available at https://github.com/evanmy/keymorph.
An Auto-Encoder Strategy for Adaptive Image Segmentation
Apr 29, 2020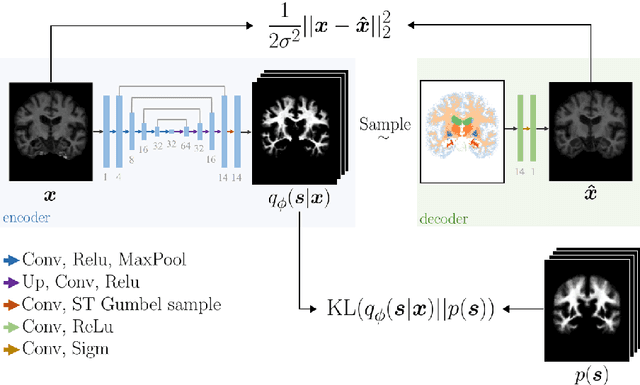
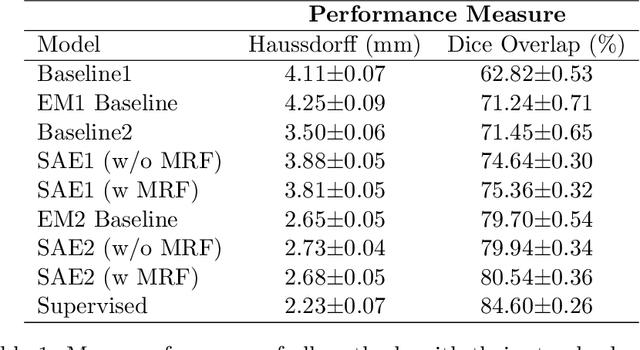
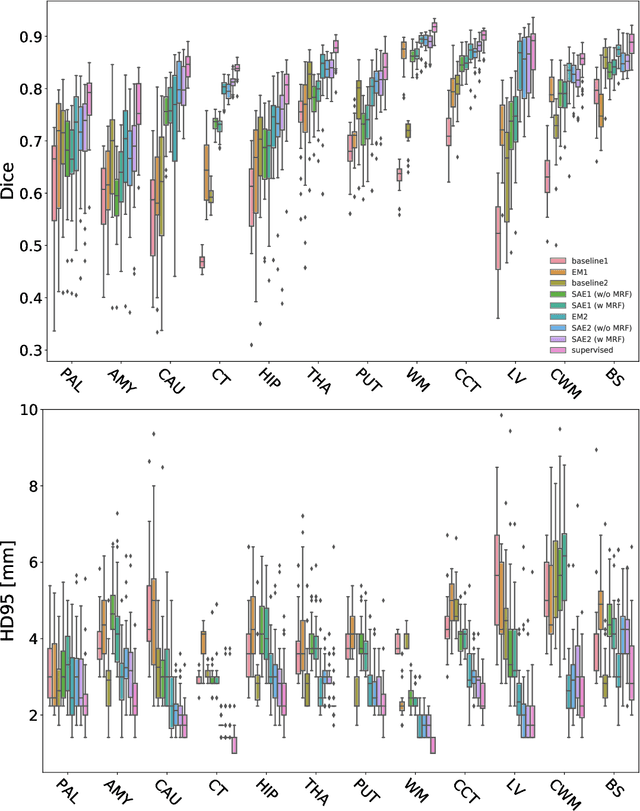
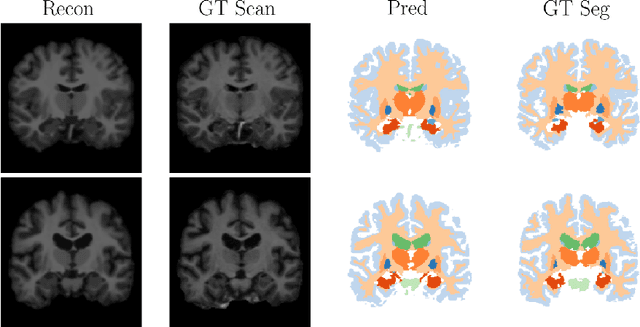
Abstract:Deep neural networks are powerful tools for biomedical image segmentation. These models are often trained with heavy supervision, relying on pairs of images and corresponding voxel-level labels. However, obtaining segmentations of anatomical regions on a large number of cases can be prohibitively expensive. Thus there is a strong need for deep learning-based segmentation tools that do not require heavy supervision and can continuously adapt. In this paper, we propose a novel perspective of segmentation as a discrete representation learning problem, and present a variational autoencoder segmentation strategy that is flexible and adaptive. Our method, called Segmentation Auto-Encoder (SAE), leverages all available unlabeled scans and merely requires a segmentation prior, which can be a single unpaired segmentation image. In experiments, we apply SAE to brain MRI scans. Our results show that SAE can produce good quality segmentations, particularly when the prior is good. We demonstrate that a Markov Random Field prior can yield significantly better results than a spatially independent prior. Our code is freely available at https://github.com/evanmy/sae.
A Convolutional Autoencoder Approach to Learn Volumetric Shape Representations for Brain Structures
Oct 17, 2018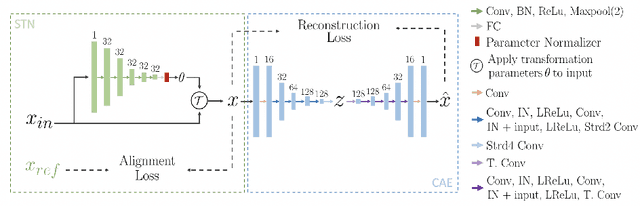
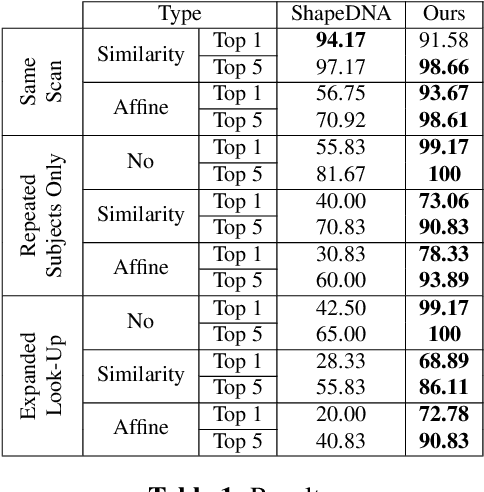
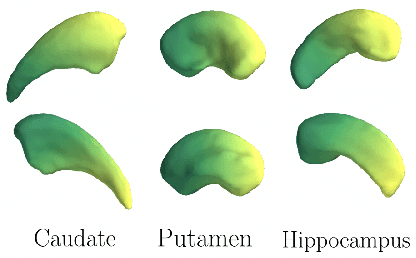
Abstract:We propose a novel machine learning strategy for studying neuroanatomical shape variation. Our model works with volumetric binary segmentation images, and requires no pre-processing such as the extraction of surface points or a mesh. The learned shape descriptor is invariant to affine transformations, including shifts, rotations and scaling. Thanks to the adopted autoencoder framework, inter-subject differences are automatically enhanced in the learned representation, while intra-subject variances are minimized. Our experimental results on a shape retrieval task showed that the proposed representation outperforms a state-of-the-art benchmark for brain structures extracted from MRI scans.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge