Eric Kelsic
Variational auto-encoding of protein sequences
Jan 03, 2018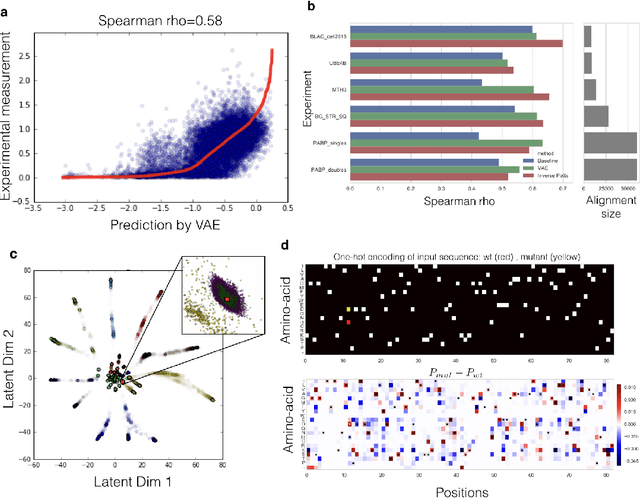
Abstract:Proteins are responsible for the most diverse set of functions in biology. The ability to extract information from protein sequences and to predict the effects of mutations is extremely valuable in many domains of biology and medicine. However the mapping between protein sequence and function is complex and poorly understood. Here we present an embedding of natural protein sequences using a Variational Auto-Encoder and use it to predict how mutations affect protein function. We use this unsupervised approach to cluster natural variants and learn interactions between sets of positions within a protein. This approach generally performs better than baseline methods that consider no interactions within sequences, and in some cases better than the state-of-the-art approaches that use the inverse-Potts model. This generative model can be used to computationally guide exploration of protein sequence space and to better inform rational and automatic protein design.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge