Daphne Koller
Stanford University
Discriminative Probabilistic Models for Relational Data
Dec 12, 2012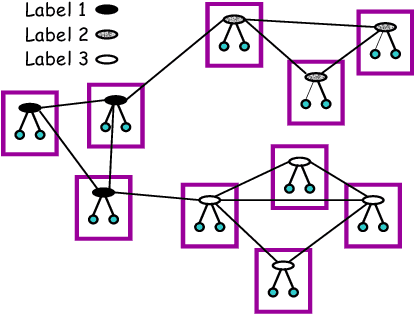

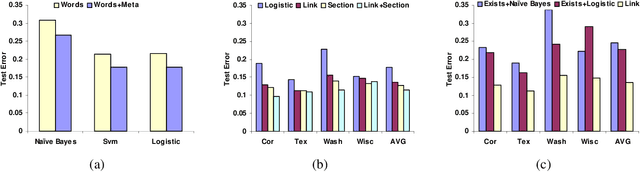
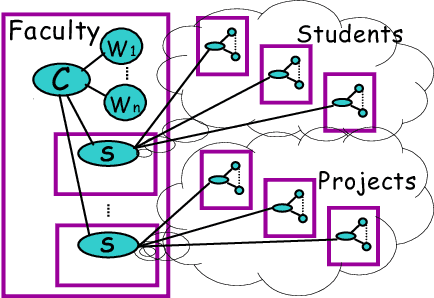
Abstract:In many supervised learning tasks, the entities to be labeled are related to each other in complex ways and their labels are not independent. For example, in hypertext classification, the labels of linked pages are highly correlated. A standard approach is to classify each entity independently, ignoring the correlations between them. Recently, Probabilistic Relational Models, a relational version of Bayesian networks, were used to define a joint probabilistic model for a collection of related entities. In this paper, we present an alternative framework that builds on (conditional) Markov networks and addresses two limitations of the previous approach. First, undirected models do not impose the acyclicity constraint that hinders representation of many important relational dependencies in directed models. Second, undirected models are well suited for discriminative training, where we optimize the conditional likelihood of the labels given the features, which generally improves classification accuracy. We show how to train these models effectively, and how to use approximate probabilistic inference over the learned model for collective classification of multiple related entities. We provide experimental results on a webpage classification task, showing that accuracy can be significantly improved by modeling relational dependencies.
Continuous Time Bayesian Networks
Dec 12, 2012
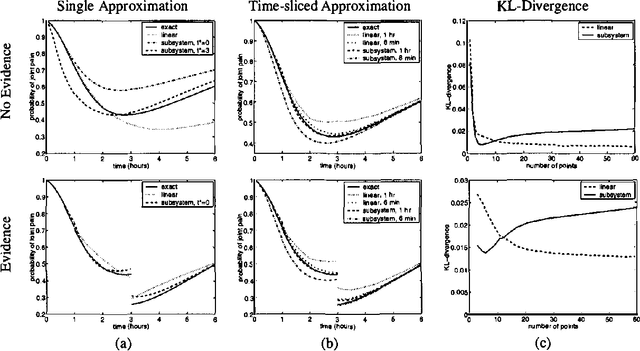
Abstract:In this paper we present a language for finite state continuous time Bayesian networks (CTBNs), which describe structured stochastic processes that evolve over continuous time. The state of the system is decomposed into a set of local variables whose values change over time. The dynamics of the system are described by specifying the behavior of each local variable as a function of its parents in a directed (possibly cyclic) graph. The model specifies, at any given point in time, the distribution over two aspects: when a local variable changes its value and the next value it takes. These distributions are determined by the variable s CURRENT value AND the CURRENT VALUES OF its parents IN the graph.More formally, each variable IS modelled AS a finite state continuous time Markov process whose transition intensities are functions OF its parents.We present a probabilistic semantics FOR the language IN terms OF the generative model a CTBN defines OVER sequences OF events.We list types OF queries one might ask OF a CTBN, discuss the conceptual AND computational difficulties associated WITH exact inference, AND provide an algorithm FOR approximate inference which takes advantage OF the structure within the process.
Monitoring a Complez Physical System using a Hybrid Dynamic Bayes Net
Dec 12, 2012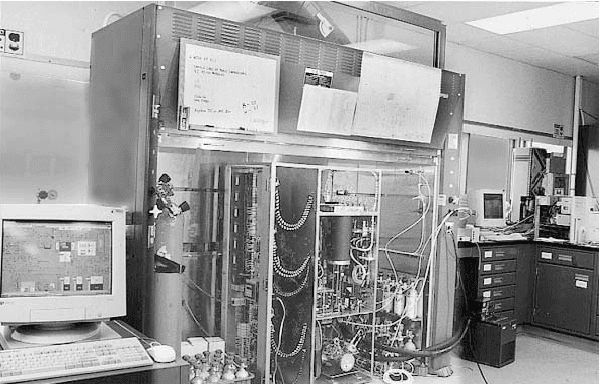
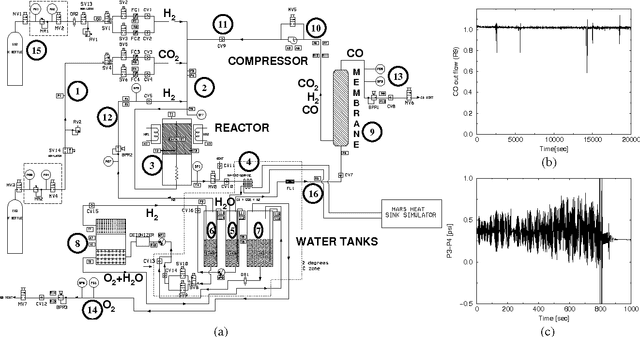
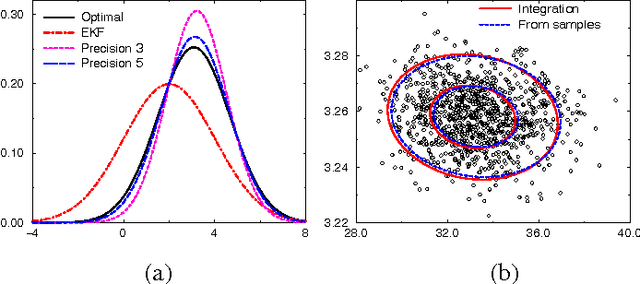
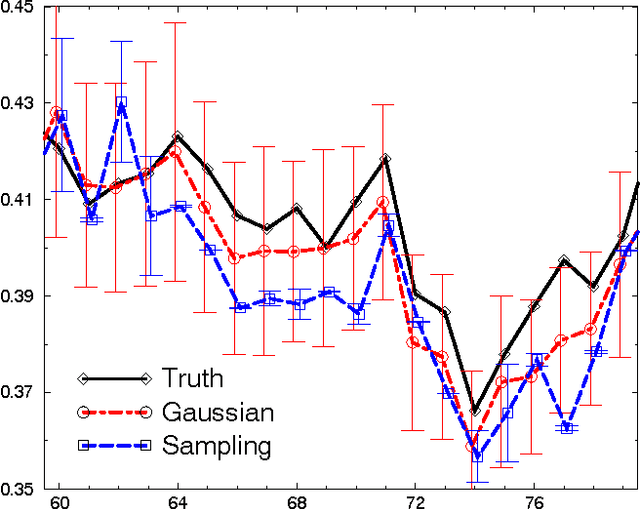
Abstract:The Reverse Water Gas Shift system (RWGS) is a complex physical system designed to produce oxygen from the carbon dioxide atmosphere on Mars. If sent to Mars, it would operate without human supervision, thus requiring a reliable automated system for monitoring and control. The RWGS presents many challenges typical of real-world systems, including: noisy and biased sensors, nonlinear behavior, effects that are manifested over different time granularities, and unobservability of many important quantities. In this paper we model the RWGS using a hybrid (discrete/continuous) Dynamic Bayesian Network (DBN), where the state at each time slice contains 33 discrete and 184 continuous variables. We show how the system state can be tracked using probabilistic inference over the model. We discuss how to deal with the various challenges presented by the RWGS, providing a suite of techniques that are likely to be useful in a wide range of applications. In particular, we describe a general framework for dealing with nonlinear behavior using numerical integration techniques, extending the successful Unscented Filter. We also show how to use a fixed-point computation to deal with effects that develop at different time scales, specifically rapid changes occurring during slowly changing processes. We test our model using real data collected from the RWGS, demonstrating the feasibility of hybrid DBNs for monitoring complex real-world physical systems.
Learning Hierarchical Object Maps Of Non-Stationary Environments with mobile robots
Dec 12, 2012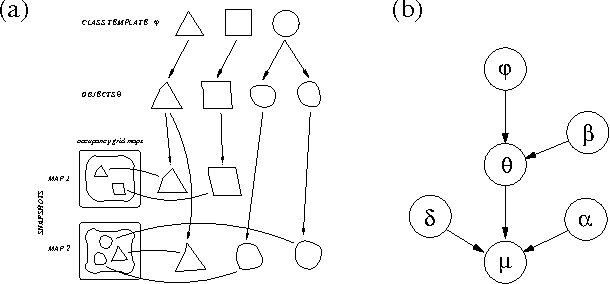
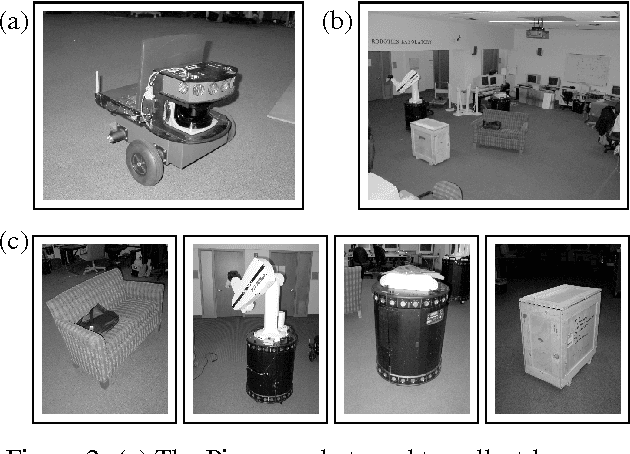
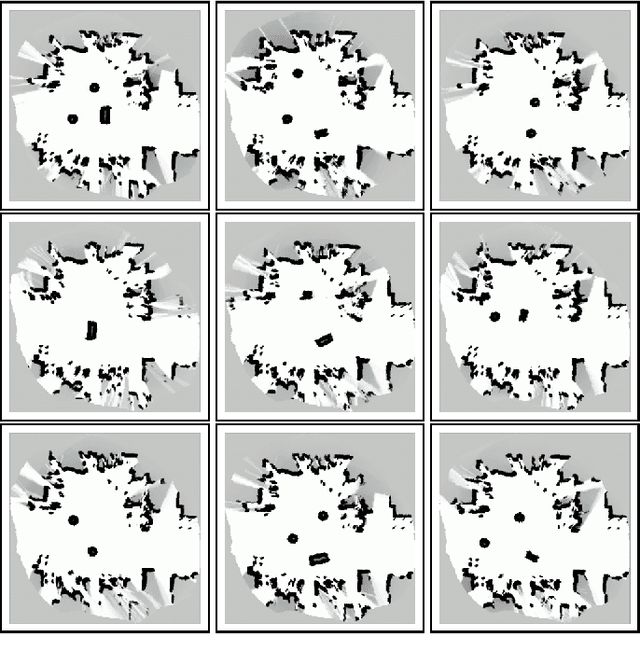

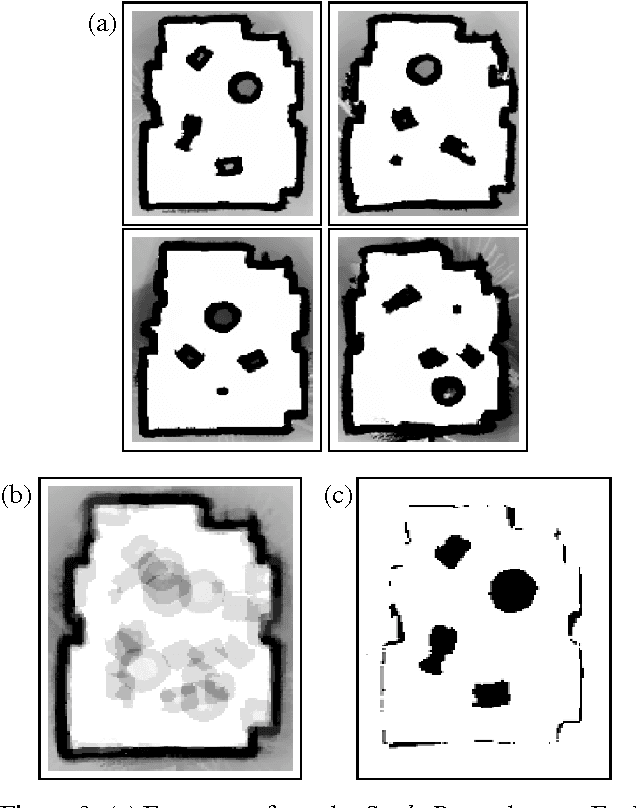

Abstract:Building models, or maps, of robot environments is a highly active research area; however, most existing techniques construct unstructured maps and assume static environments. In this paper, we present an algorithm for learning object models of non-stationary objects found in office-type environments. Our algorithm exploits the fact that many objects found in office environments look alike (e.g., chairs, recycling bins). It does so through a two-level hierarchical representation, which links individual objects with generic shape templates of object classes. We derive an approximate EM algorithm for learning shape parameters at both levels of the hierarchy, using local occupancy grid maps for representing shape. Additionally, we develop a Bayesian model selection algorithm that enables the robot to estimate the total number of objects and object templates in the environment. Experimental results using a real robot equipped with a laser range finder indicate that our approach performs well at learning object-based maps of simple office environments. The approach outperforms a previously developed non-hierarchical algorithm that models objects but lacks class templates.
Learning Module Networks
Oct 19, 2012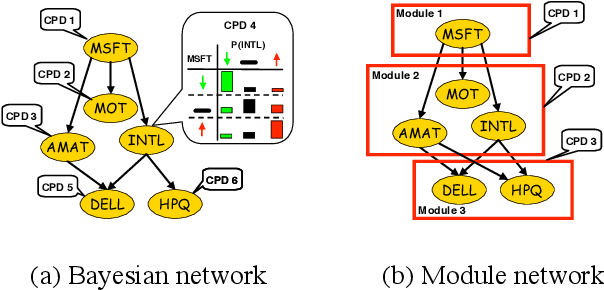
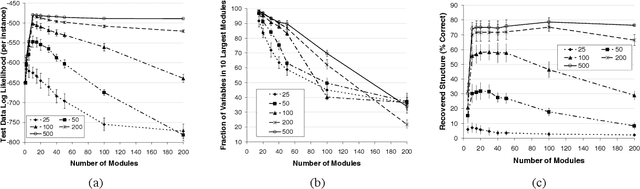
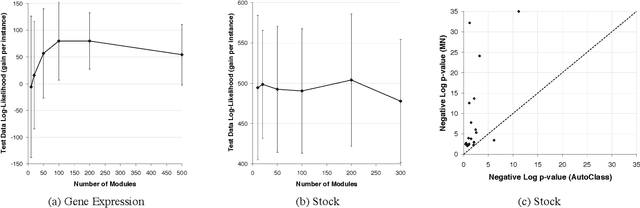
Abstract:Methods for learning Bayesian network structure can discover dependency structure between observed variables, and have been shown to be useful in many applications. However, in domains that involve a large number of variables, the space of possible network structures is enormous, making it difficult, for both computational and statistical reasons, to identify a good model. In this paper, we consider a solution to this problem, suitable for domains where many variables have similar behavior. Our method is based on a new class of models, which we call module networks. A module network explicitly represents the notion of a module - a set of variables that have the same parents in the network and share the same conditional probability distribution. We define the semantics of module networks, and describe an algorithm that learns a module network from data. The algorithm learns both the partitioning of the variables into modules and the dependency structure between the variables. We evaluate our algorithm on synthetic data, and on real data in the domains of gene expression and the stock market. Our results show that module networks generalize better than Bayesian networks, and that the learned module network structure reveals regularities that are obscured in learned Bayesian networks.
Learning Continuous Time Bayesian Networks
Oct 19, 2012
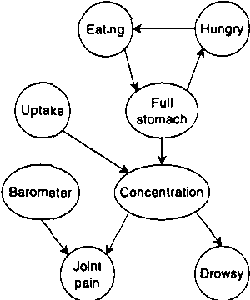
Abstract:Continuous time Bayesian networks (CTBNs) describe structured stochastic processes with finitely many states that evolve over continuous time. A CTBN is a directed (possibly cyclic) dependency graph over a set of variables, each of which represents a finite state continuous time Markov process whose transition model is a function of its parents. We address the problem of learning parameters and structure of a CTBN from fully observed data. We define a conjugate prior for CTBNs, and show how it can be used both for Bayesian parameter estimation and as the basis of a Bayesian score for structure learning. Because acyclicity is not a constraint in CTBNs, we can show that the structure learning problem is significantly easier, both in theory and in practice, than structure learning for dynamic Bayesian networks (DBNs). Furthermore, as CTBNs can tailor the parameters and dependency structure to the different time granularities of the evolution of different variables, they can provide a better fit to continuous-time processes than DBNs with a fixed time granularity.
Recovering Articulated Object Models from 3D Range Data
Jul 11, 2012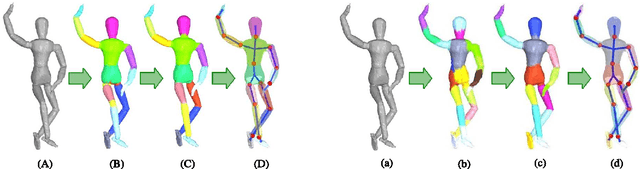
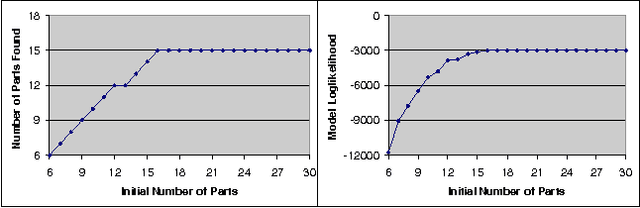
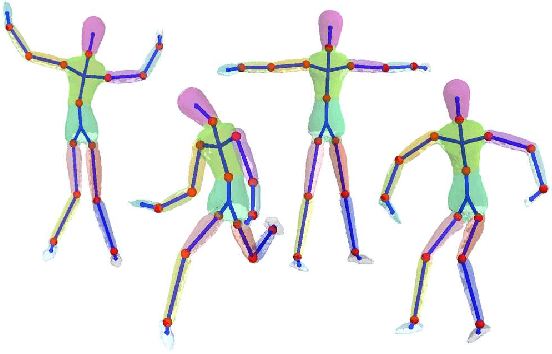
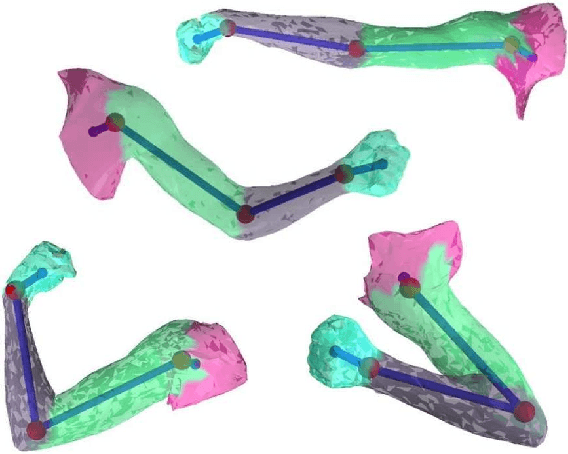
Abstract:We address the problem of unsupervised learning of complex articulated object models from 3D range data. We describe an algorithm whose input is a set of meshes corresponding to different configurations of an articulated object. The algorithm automatically recovers a decomposition of the object into approximately rigid parts, the location of the parts in the different object instances, and the articulated object skeleton linking the parts. Our algorithm first registers allthe meshes using an unsupervised non-rigid technique described in a companion paper. It then segments the meshes using a graphical model that captures the spatial contiguity of parts. The segmentation is done using the EM algorithm, iterating between finding a decomposition of the object into rigid parts, and finding the location of the parts in the object instances. Although the graphical model is densely connected, the object decomposition step can be performed optimally and efficiently, allowing us to identify a large number of object parts while avoiding local maxima. We demonstrate the algorithm on real world datasets, recovering a 15-part articulated model of a human puppet from just 7 different puppet configurations, as well as a 4 part model of a fiexing arm where significant non-rigid deformation was present.
Ordering-Based Search: A Simple and Effective Algorithm for Learning Bayesian Networks
Jul 04, 2012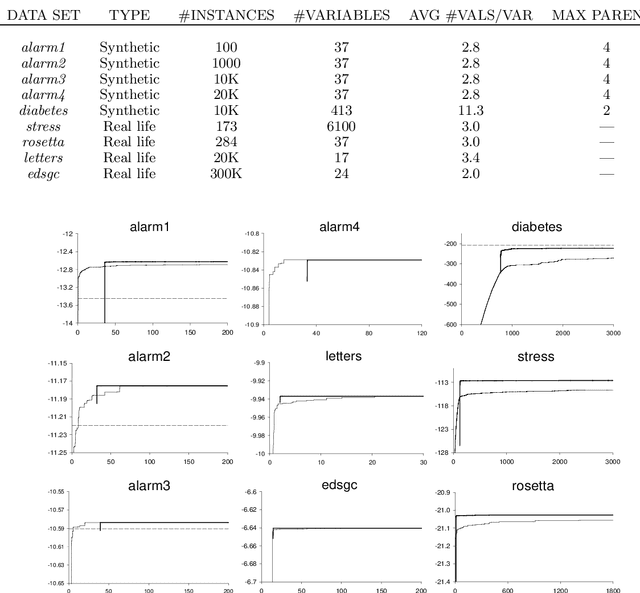
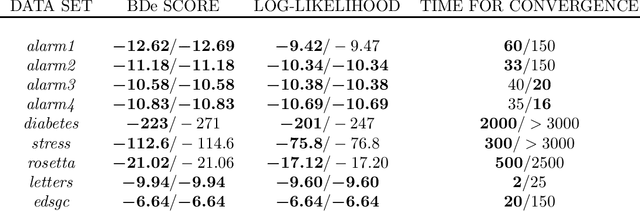


Abstract:One of the basic tasks for Bayesian networks (BNs) is that of learning a network structure from data. The BN-learning problem is NP-hard, so the standard solution is heuristic search. Many approaches have been proposed for this task, but only a very small number outperform the baseline of greedy hill-climbing with tabu lists; moreover, many of the proposed algorithms are quite complex and hard to implement. In this paper, we propose a very simple and easy-to-implement method for addressing this task. Our approach is based on the well-known fact that the best network (of bounded in-degree) consistent with a given node ordering can be found very efficiently. We therefore propose a search not over the space of structures, but over the space of orderings, selecting for each ordering the best network consistent with it. This search space is much smaller, makes more global search steps, has a lower branching factor, and avoids costly acyclicity checks. We present results for this algorithm on both synthetic and real data sets, evaluating both the score of the network found and in the running time. We show that ordering-based search outperforms the standard baseline, and is competitive with recent algorithms that are much harder to implement.
Expectation Maximization and Complex Duration Distributions for Continuous Time Bayesian Networks
Jul 04, 2012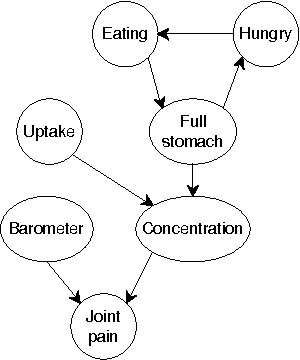
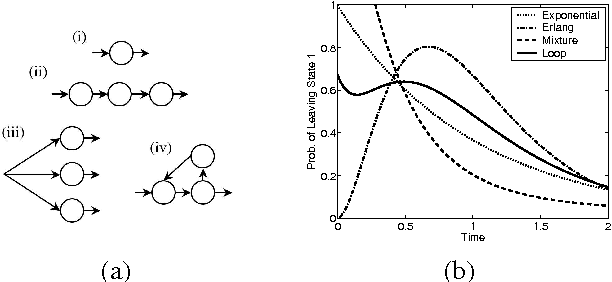
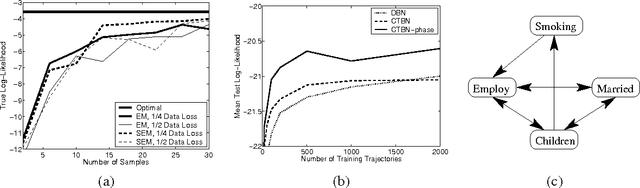
Abstract:Continuous time Bayesian networks (CTBNs) describe structured stochastic processes with finitely many states that evolve over continuous time. A CTBN is a directed (possibly cyclic) dependency graph over a set of variables, each of which represents a finite state continuous time Markov process whose transition model is a function of its parents. We address the problem of learning the parameters and structure of a CTBN from partially observed data. We show how to apply expectation maximization (EM) and structural expectation maximization (SEM) to CTBNs. The availability of the EM algorithm allows us to extend the representation of CTBNs to allow a much richer class of transition durations distributions, known as phase distributions. This class is a highly expressive semi-parametric representation, which can approximate any duration distribution arbitrarily closely. This extension to the CTBN framework addresses one of the main limitations of both CTBNs and DBNs - the restriction to exponentially / geometrically distributed duration. We present experimental results on a real data set of people's life spans, showing that our algorithm learns reasonable models - structure and parameters - from partially observed data, and, with the use of phase distributions, achieves better performance than DBNs.
Expectation Propagation for Continuous Time Bayesian Networks
Jul 04, 2012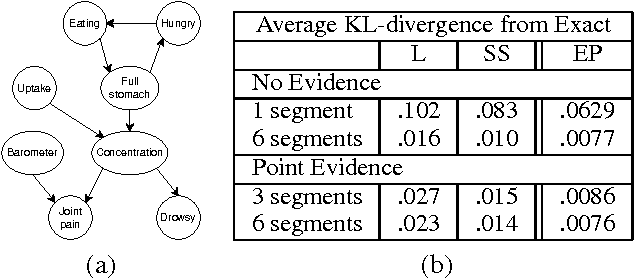
Abstract:Continuous time Bayesian networks (CTBNs) describe structured stochastic processes with finitely many states that evolve over continuous time. A CTBN is a directed (possibly cyclic) dependency graph over a set of variables, each of which represents a finite state continuous time Markov process whose transition model is a function of its parents. As shown previously, exact inference in CTBNs is intractable. We address the problem of approximate inference, allowing for general queries conditioned on evidence over continuous time intervals and at discrete time points. We show how CTBNs can be parameterized within the exponential family, and use that insight to develop a message passing scheme in cluster graphs and allows us to apply expectation propagation to CTBNs. The clusters in our cluster graph do not contain distributions over the cluster variables at individual time points, but distributions over trajectories of the variables throughout a duration. Thus, unlike discrete time temporal models such as dynamic Bayesian networks, we can adapt the time granularity at which we reason for different variables and in different conditions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge