Daniel S. Berman
Johns Hopkins Applied Physics Laboratory, Laurel, United States
What You Read is What You Classify: Highlighting Attributions to Text and Text-Like Inputs
Feb 27, 2026Abstract:At present, there are no easily understood explainable artificial intelligence (AI) methods for discrete token inputs, like text. Most explainable AI techniques do not extend well to token sequences, where both local and global features matter, because state-of-the-art models, like transformers, tend to focus on global connections. Therefore, existing explainable AI algorithms fail by (i) identifying disparate tokens of importance, or (ii) assigning a large number of tokens a low value of importance. This method for explainable AI for tokens-based classifiers generalizes a mask-based explainable AI algorithm for images. It starts with an Explainer neural network that is trained to create masks to hide information not relevant for classification. Then, the Hadamard product of the mask and the continuous values of the classifier's embedding layer is taken and passed through the classifier, changing the magnitude of the embedding vector but keeping the orientation unchanged. The Explainer is trained for a taxonomic classifier for nucleotide sequences and it is shown that the masked segments are less relevant to classification than the unmasked ones. This method focused on the importance the token as a whole (i.e., a segment of the input sequence), producing a human-readable explanation.
MutaGAN: A Seq2seq GAN Framework to Predict Mutations of Evolving Protein Populations
Aug 26, 2020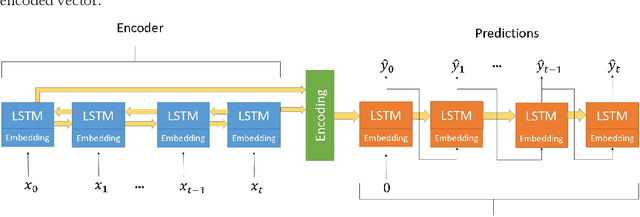
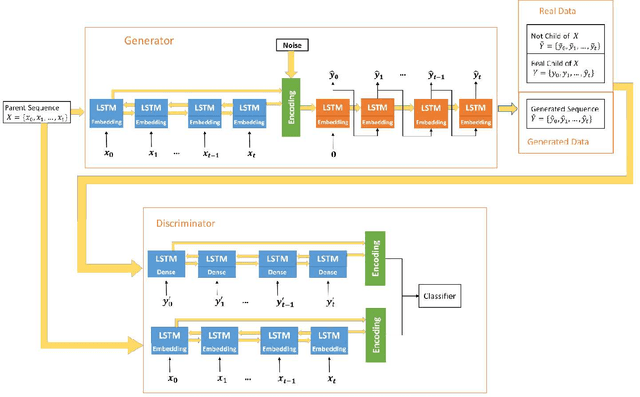
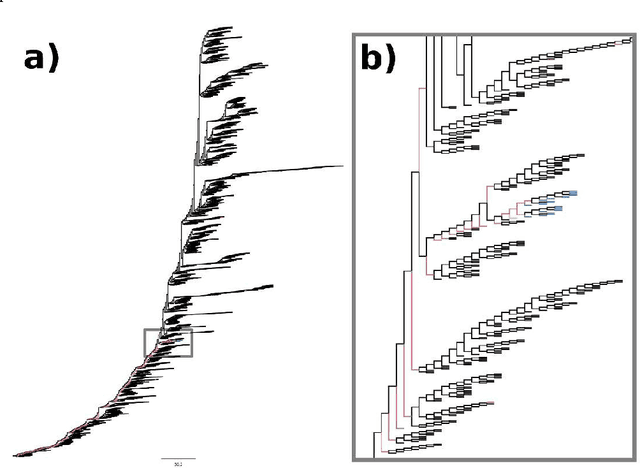
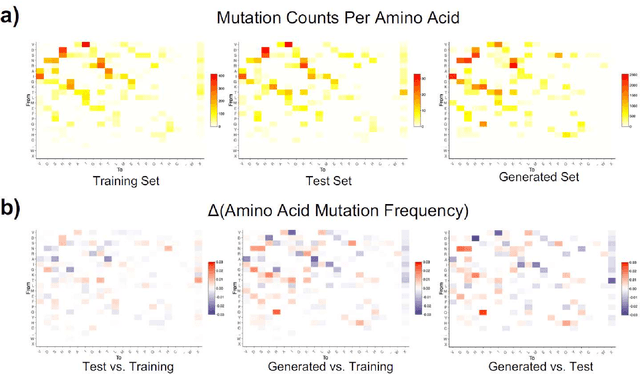
Abstract:The ability to predict the evolution of a pathogen would significantly improve the ability to control, prevent, and treat disease. Despite significant progress in other problem spaces, deep learning has yet to contribute to the issue of predicting mutations of evolving populations. To address this gap, we developed a novel machine learning framework using generative adversarial networks (GANs) with recurrent neural networks (RNNs) to accurately predict genetic mutations and evolution of future biological populations. Using a generalized time-reversible phylogenetic model of protein evolution with bootstrapped maximum likelihood tree estimation, we trained a sequence-to-sequence generator within an adversarial framework, named MutaGAN, to generate complete protein sequences augmented with possible mutations of future virus populations. Influenza virus sequences were identified as an ideal test case for this deep learning framework because it is a significant human pathogen with new strains emerging annually and global surveillance efforts have generated a large amount of publicly available data from the National Center for Biotechnology Information's (NCBI) Influenza Virus Resource (IVR). MutaGAN generated "child" sequences from a given "parent" protein sequence with a median Levenshtein distance of 2.00 amino acids. Additionally, the generator was able to augment the majority of parent proteins with at least one mutation identified within the global influenza virus population. These results demonstrate the power of the MutaGAN framework to aid in pathogen forecasting with implications for broad utility in evolutionary prediction for any protein population.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge