D. S. Magruder
MIOFlow 2.0: A unified framework for inferring cellular stochastic dynamics from single cell and spatial transcriptomics data
Mar 23, 2026Abstract:Understanding cellular trajectories via time-resolved single-cell transcriptomics is vital for studying development, regeneration, and disease. A key challenge is inferring continuous trajectories from discrete snapshots. Biological complexity stems from stochastic cell fate decisions, temporal proliferation changes, and spatial environmental influences. Current methods often use deterministic interpolations treating cells in isolation, failing to capture the probabilistic branching, population shifts, and niche-dependent signaling driving real biological processes. We introduce Manifold Interpolating Optimal-Transport Flow (MIOFlow) 2.0. This framework learns biologically informed cellular trajectories by integrating manifold learning, optimal transport, and neural differential equations. It models three core processes: (1) stochasticity and branching via Neural Stochastic Differential Equations; (2) non-conservative population changes using a learned growth-rate model initialized with unbalanced optimal transport; and (3) environmental influence through a joint latent space unifying gene expression with spatial features like local cell type composition and signaling. By operating in a PHATE-distance matching autoencoder latent space, MIOFlow 2.0 ensures trajectories respect the data's intrinsic geometry. Empirical comparisons show expressive trajectory learning via neural differential equations outperforms existing generative models, including simulation-free flow matching. Validated on synthetic datasets, embryoid body differentiation, and spatially resolved axolotl brain regeneration, MIOFlow 2.0 improves trajectory accuracy and reveals hidden drivers of cellular transitions, like specific signaling niches. MIOFlow 2.0 thus bridges single-cell and spatial transcriptomics to uncover tissue-scale trajectories.
Manifold Interpolating Optimal-Transport Flows for Trajectory Inference
Jun 29, 2022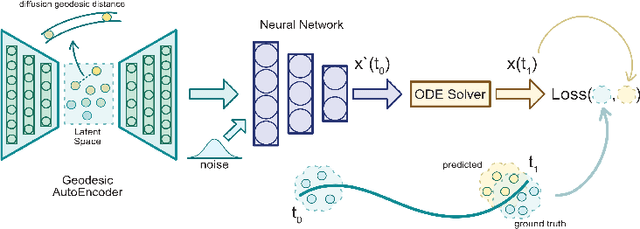
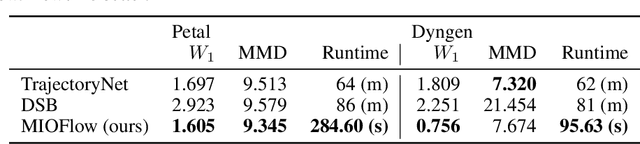
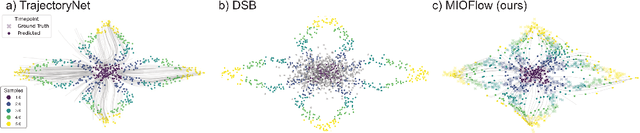
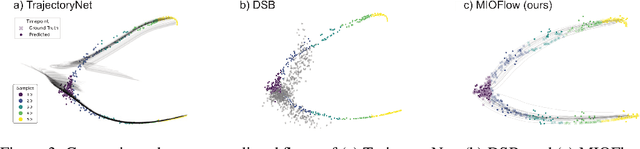
Abstract:Here, we present a method called Manifold Interpolating Optimal-Transport Flow (MIOFlow) that learns stochastic, continuous population dynamics from static snapshot samples taken at sporadic timepoints. MIOFlow combines dynamic models, manifold learning, and optimal transport by training neural ordinary differential equations (Neural ODE) to interpolate between static population snapshots as penalized by optimal transport with manifold ground distance. Further, we ensure that the flow follows the geometry by operating in the latent space of an autoencoder that we call a geodesic autoencoder (GAE). In GAE the latent space distance between points is regularized to match a novel multiscale geodesic distance on the data manifold that we define. We show that this method is superior to normalizing flows, Schr\"odinger bridges and other generative models that are designed to flow from noise to data in terms of interpolating between populations. Theoretically, we link these trajectories with dynamic optimal transport. We evaluate our method on simulated data with bifurcations and merges, as well as scRNA-seq data from embryoid body differentiation, and acute myeloid leukemia treatment.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge