D. Rubin
A Probabilistic Autoencoder for Type Ia Supernovae Spectral Time Series
Jul 15, 2022
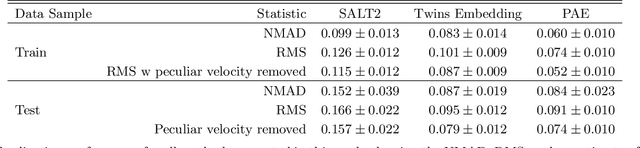
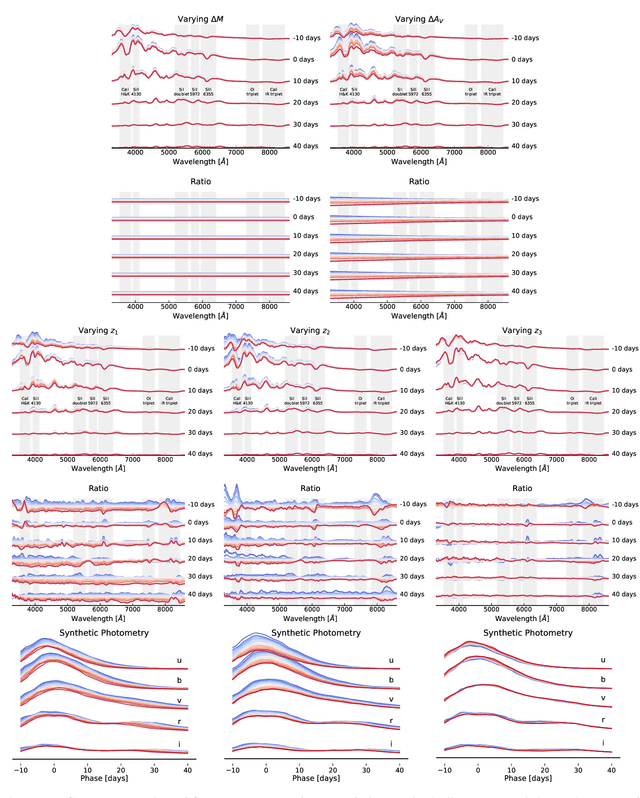
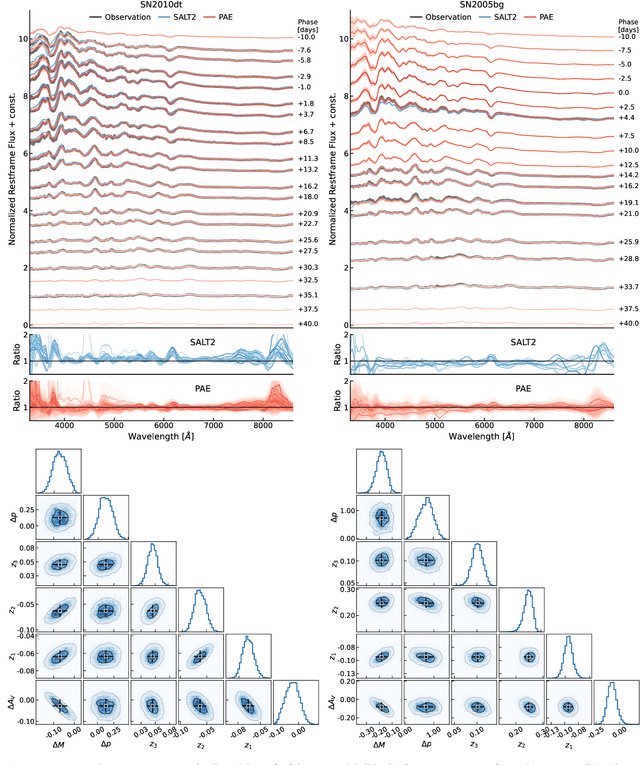
Abstract:We construct a physically-parameterized probabilistic autoencoder (PAE) to learn the intrinsic diversity of type Ia supernovae (SNe Ia) from a sparse set of spectral time series. The PAE is a two-stage generative model, composed of an Auto-Encoder (AE) which is interpreted probabilistically after training using a Normalizing Flow (NF). We demonstrate that the PAE learns a low-dimensional latent space that captures the nonlinear range of features that exists within the population, and can accurately model the spectral evolution of SNe Ia across the full range of wavelength and observation times directly from the data. By introducing a correlation penalty term and multi-stage training setup alongside our physically-parameterized network we show that intrinsic and extrinsic modes of variability can be separated during training, removing the need for the additional models to perform magnitude standardization. We then use our PAE in a number of downstream tasks on SNe Ia for increasingly precise cosmological analyses, including automatic detection of SN outliers, the generation of samples consistent with the data distribution, and solving the inverse problem in the presence of noisy and incomplete data to constrain cosmological distance measurements. We find that the optimal number of intrinsic model parameters appears to be three, in line with previous studies, and show that we can standardize our test sample of SNe Ia with an RMS of $0.091 \pm 0.010$ mag, which corresponds to $0.074 \pm 0.010$ mag if peculiar velocity contributions are removed. Trained models and codes are released at \href{https://github.com/georgestein/suPAErnova}{github.com/georgestein/suPAErnova}
An Adaptive Strategy for the Classification of G-Protein Coupled Receptors
Apr 25, 2007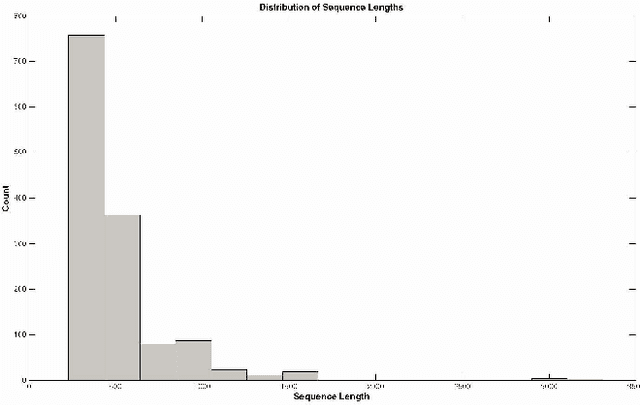
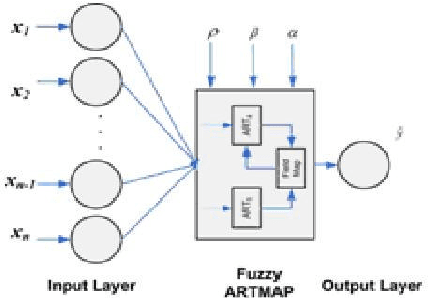
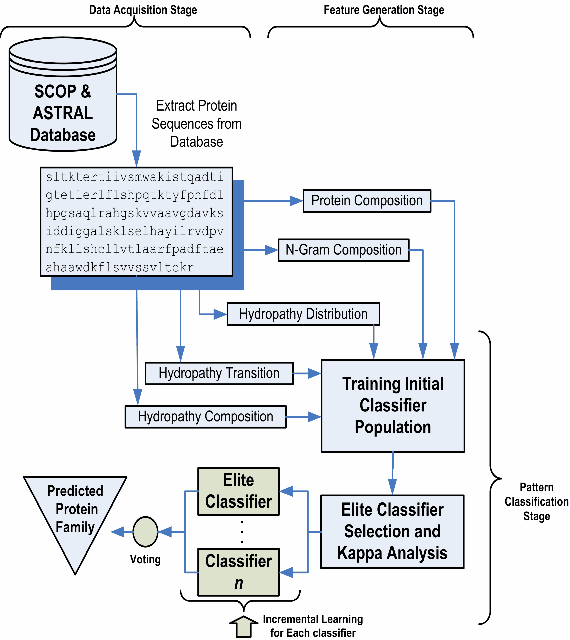
Abstract:One of the major problems in computational biology is the inability of existing classification models to incorporate expanding and new domain knowledge. This problem of static classification models is addressed in this paper by the introduction of incremental learning for problems in bioinformatics. Many machine learning tools have been applied to this problem using static machine learning structures such as neural networks or support vector machines that are unable to accommodate new information into their existing models. We utilize the fuzzy ARTMAP as an alternate machine learning system that has the ability of incrementally learning new data as it becomes available. The fuzzy ARTMAP is found to be comparable to many of the widespread machine learning systems. The use of an evolutionary strategy in the selection and combination of individual classifiers into an ensemble system, coupled with the incremental learning ability of the fuzzy ARTMAP is proven to be suitable as a pattern classifier. The algorithm presented is tested using data from the G-Coupled Protein Receptors Database and shows good accuracy of 83%. The system presented is also generally applicable, and can be used in problems in genomics and proteomics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge