Claus O. Wilke
Caltech
Mining Patents with Large Language Models Demonstrates Congruence of Functional Labels and Chemical Structures
Sep 15, 2023



Abstract:Predicting chemical function from structure is a major goal of the chemical sciences, from the discovery and repurposing of novel drugs to the creation of new materials. Recently, new machine learning algorithms are opening up the possibility of general predictive models spanning many different chemical functions. Here, we consider the challenge of applying large language models to chemical patents in order to consolidate and leverage the information about chemical functionality captured by these resources. Chemical patents contain vast knowledge on chemical function, but their usefulness as a dataset has historically been neglected due to the impracticality of extracting high-quality functional labels. Using a scalable ChatGPT-assisted patent summarization and word-embedding label cleaning pipeline, we derive a Chemical Function (CheF) dataset, containing 100K molecules and their patent-derived functional labels. The functional labels were validated to be of high quality, allowing us to detect a strong relationship between functional label and chemical structural spaces. Further, we find that the co-occurrence graph of the functional labels contains a robust semantic structure, which allowed us in turn to examine functional relatedness among the compounds. We then trained a model on the CheF dataset, allowing us to assign new functional labels to compounds. Using this model, we were able to retrodict approved Hepatitis C antivirals, uncover an antiviral mechanism undisclosed in the patent, and identify plausible serotonin-related drugs. The CheF dataset and associated model offers a promising new approach to predict chemical functionality.
Prompt Engineering for Transformer-based Chemical Similarity Search Identifies Structurally Distinct Functional Analogues
May 17, 2023



Abstract:Chemical similarity searches are widely used in-silico methods for identifying new drug-like molecules. These methods have historically relied on structure-based comparisons to compute molecular similarity. Here, we use a chemical language model to create a vector-based chemical search. We extend implementations by creating a prompt engineering strategy that utilizes two different chemical string representation algorithms: one for the query and the other for the database. We explore this method by reviewing the search results from five drug-like query molecules (penicillin G, nirmatrelvir, zidovudine, lysergic acid diethylamide, and fentanyl) and three dye-like query molecules (acid blue 25, avobenzone, and 2-diphenylaminocarbazole). We find that this novel method identifies molecules that are functionally similar to the query, indicated by the associated patent literature, and that many of these molecules are structurally distinct from the query, making them unlikely to be found with traditional chemical similarity search methods. This method may aid in the discovery of novel structural classes of molecules that achieve target functionality.
Adaptive evolution on neutral networks
Jan 03, 2001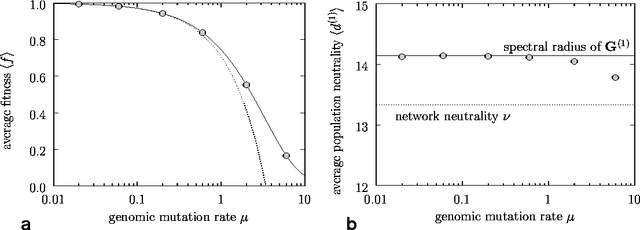
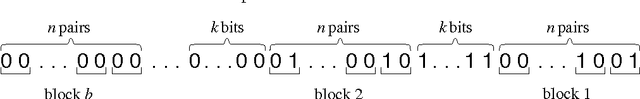
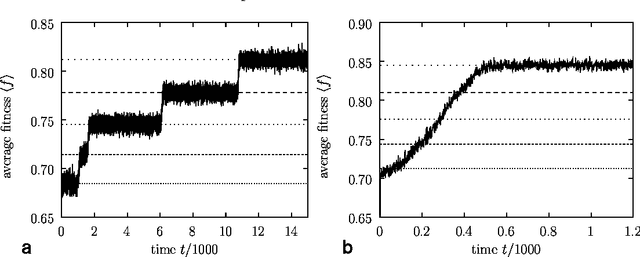
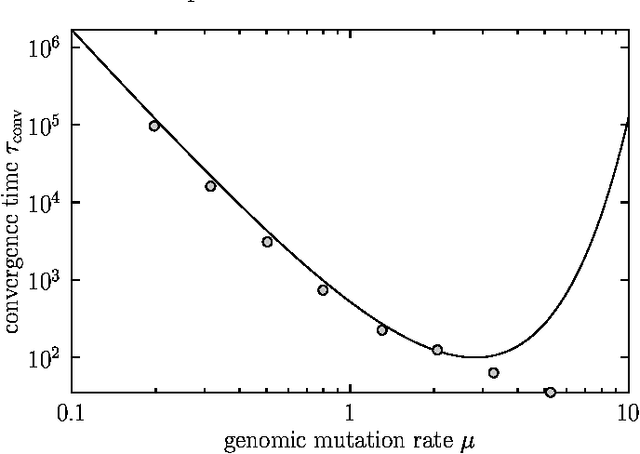
Abstract:We study the evolution of large but finite asexual populations evolving in fitness landscapes in which all mutations are either neutral or strongly deleterious. We demonstrate that despite the absence of higher fitness genotypes, adaptation takes place as regions with more advantageous distributions of neutral genotypes are discovered. Since these discoveries are typically rare events, the population dynamics can be subdivided into separate epochs, with rapid transitions between them. Within one epoch, the average fitness in the population is approximately constant. The transitions between epochs, however, are generally accompanied by a significant increase in the average fitness. We verify our theoretical considerations with two analytically tractable bitstring models.
* 16 pages, 4 eps figures, Latex (academic press style file), submitted to the Bulletin of Mathematical Biology
Genetic Algorithms in Time-Dependent Environments
Nov 04, 1999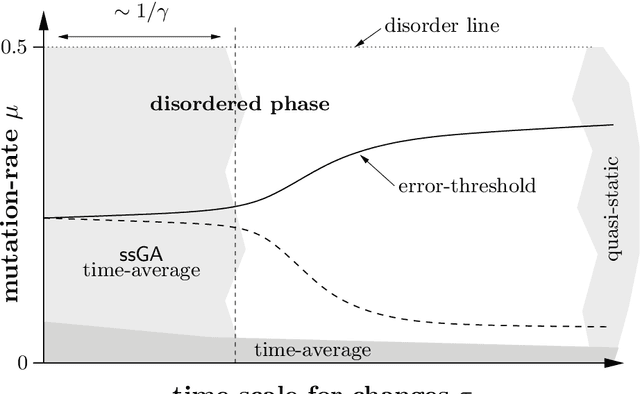
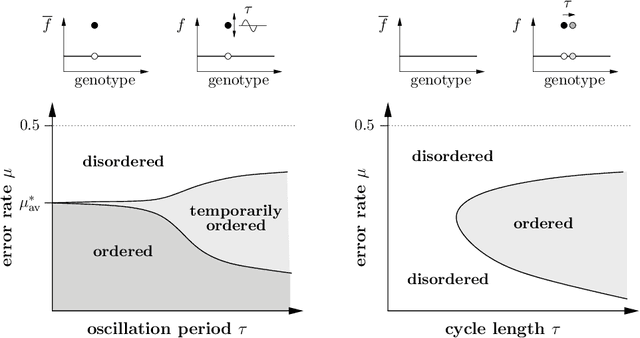
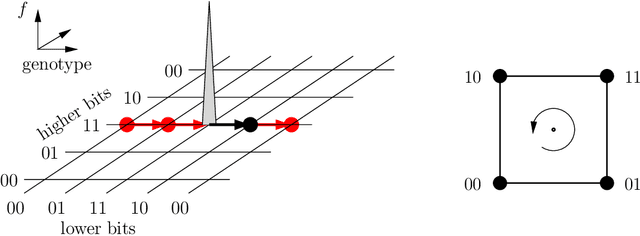
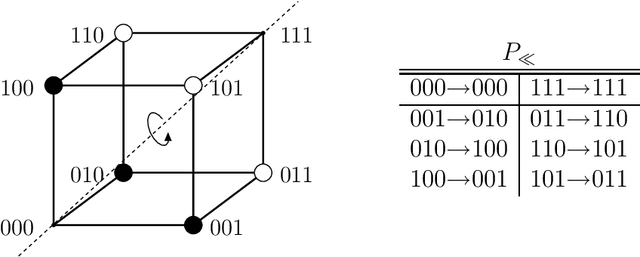
Abstract:The influence of time-dependent fitnesses on the infinite population dynamics of simple genetic algorithms (without crossover) is analyzed. Based on general arguments, a schematic phase diagram is constructed that allows one to characterize the asymptotic states in dependence on the mutation rate and the time scale of changes. Furthermore, the notion of regular changes is raised for which the population can be shown to converge towards a generalized quasispecies. Based on this, error thresholds and an optimal mutation rate are approximately calculated for a generational genetic algorithm with a moving needle-in-the-haystack landscape. The so found phase diagram is fully consistent with our general considerations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge