Christopher Snyder
Interpretable Factorization for Neural Network ECG Models
Jun 26, 2020
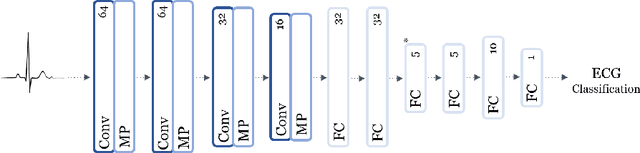
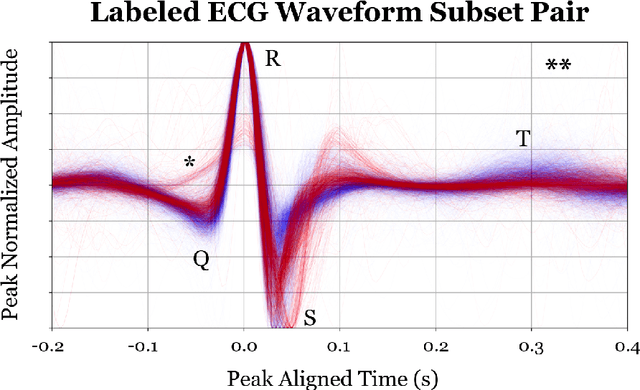
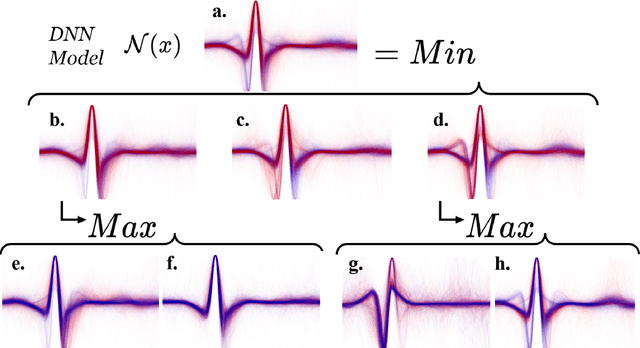
Abstract:The ability of deep learning (DL) to improve the practice of medicine and its clinical outcomes faces a looming obstacle: model interpretation. Without description of how outputs are generated, a collaborating physician can neither resolve when the model's conclusions are in conflict with his or her own, nor learn to anticipate model behavior. Current research aims to interpret networks that diagnose ECG recordings, which has great potential impact as recordings become more personalized and widely deployed. A generalizable impact beyond ECGs lies in the ability to provide a rich test-bed for the development of interpretive techniques in medicine. Interpretive techniques for Deep Neural Networks (DNNs), however, tend to be heuristic and observational in nature, lacking the mathematical rigor one might expect in the analysis of math equations. The motivation of this paper is to offer a third option, a scientific approach. We treat the model output itself as a phenomenon to be explained through component parts and equations governing their behavior. We argue that these component parts should also be "black boxes" --additional targets to interpret heuristically with clear functional connection to the original. We show how to rigorously factor a DNN into a hierarchical equation consisting of black box variables. This is not a subdivision into physical parts, like an organism into its cells; it is but one choice of an equation into a collection of abstract functions. Yet, for DNNs trained to identify normal ECG waveforms on PhysioNet 2017 Challenge data, we demonstrate this choice yields interpretable component models identified with visual composite sketches of ECG samples in corresponding input regions. Moreover, the recursion distills this interpretation: additional factorization of component black boxes corresponds to ECG partitions that are more morphologically pure.
Deep Networks as Logical Circuits: Generalization and Interpretation
Mar 25, 2020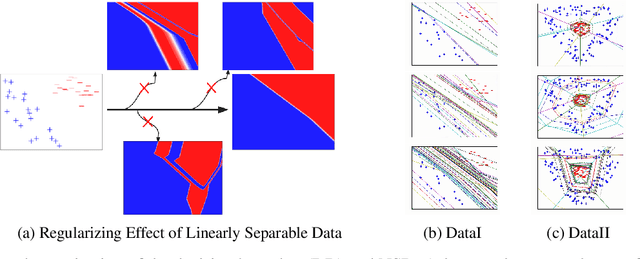
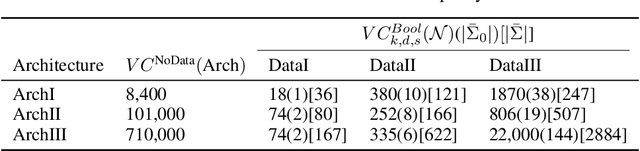

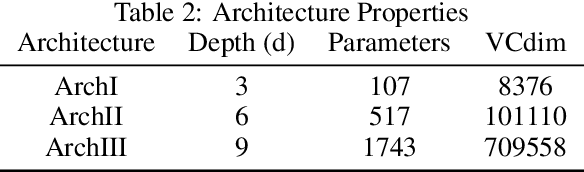
Abstract:Not only are Deep Neural Networks (DNNs) black box models, but also we frequently conceptualize them as such. We lack good interpretations of the mechanisms linking inputs to outputs. Therefore, we find it difficult to analyze in human-meaningful terms (1) what the network learned and (2) whether the network learned. We present a hierarchical decomposition of the DNN discrete classification map into logical (AND/OR) combinations of intermediate (True/False) classifiers of the input. Those classifiers that can not be further decomposed, called atoms, are (interpretable) linear classifiers. Taken together, we obtain a logical circuit with linear classifier inputs that computes the same label as the DNN. This circuit does not structurally resemble the network architecture, and it may require many fewer parameters, depending on the configuration of weights. In these cases, we obtain simultaneously an interpretation and generalization bound (for the original DNN), connecting two fronts which have historically been investigated separately. Unlike compression techniques, our representation is. We motivate the utility of this perspective by studying DNNs in simple, controlled settings, where we obtain superior generalization bounds despite using only combinatorial information (e.g. no margin information). We demonstrate how to "open the black box" on the MNIST dataset. We show that the learned, internal, logical computations correspond to semantically meaningful (unlabeled) categories that allow DNN descriptions in plain English. We improve the generalization of an already trained network by interpreting, diagnosing, and replacing components the logical circuit that is the DNN.
Generalization Bounds for Neural Networks: Kernels, Symmetry, and Sample Compression
Nov 05, 2018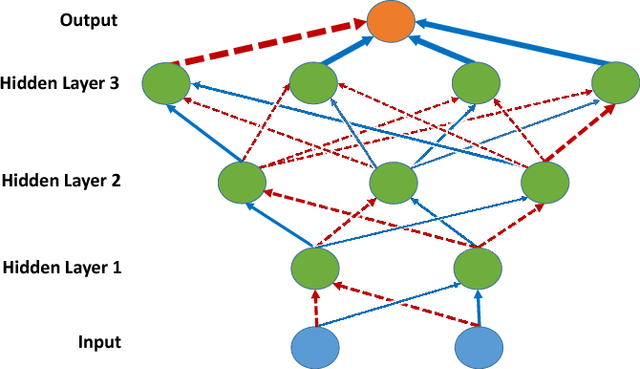
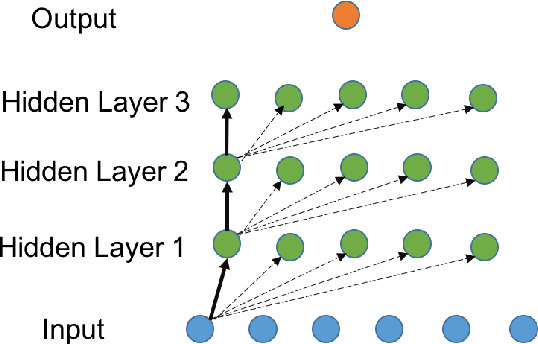
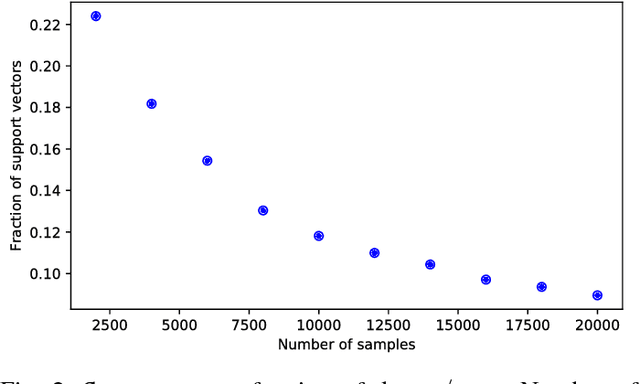
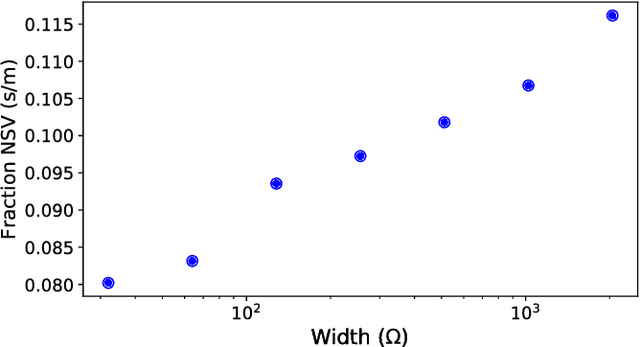
Abstract:Though Deep Neural Networks (DNNs) are widely celebrated for their practical performance, they demonstrate many intriguing phenomena related to depth that are difficult to explain both theoretically and intuitively. Understanding how weights in deep networks coordinate together across layers to form useful learners has proven somewhat intractable, in part because of the repeated composition of nonlinearities induced by depth. We present a reparameterization of DNNs as a linear function of a particular feature map that is locally independent of the weights. This feature map transforms depth-dependencies into simple {\em tensor} products and maps each input to a discrete subset of the feature space. Then, in analogy with logistic regression, we propose a max-margin assumption that enables us to present a so-called {\em sample compression} representation of the neural network in terms of the discrete activation state of neurons induced by s "support vectors". We show how the number of support vectors relate to learning guarantees for neural networks through sample compression bounds, yielding a sample complexity O(ns/\epsilon) for networks with n neurons. Additionally, this number of support vectors has monotonic dependence on width, depth, and label noise for simple networks trained on the MNIST dataset.
CausalGAN: Learning Causal Implicit Generative Models with Adversarial Training
Sep 14, 2017
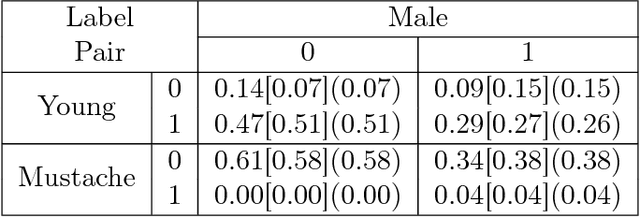

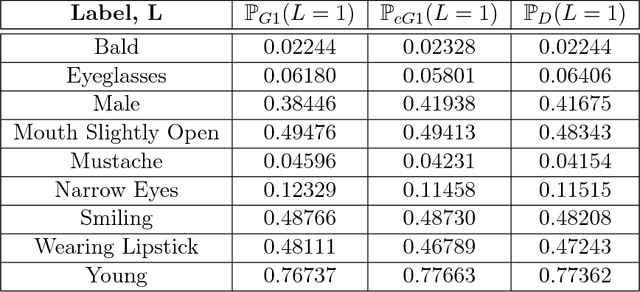
Abstract:We propose an adversarial training procedure for learning a causal implicit generative model for a given causal graph. We show that adversarial training can be used to learn a generative model with true observational and interventional distributions if the generator architecture is consistent with the given causal graph. We consider the application of generating faces based on given binary labels where the dependency structure between the labels is preserved with a causal graph. This problem can be seen as learning a causal implicit generative model for the image and labels. We devise a two-stage procedure for this problem. First we train a causal implicit generative model over binary labels using a neural network consistent with a causal graph as the generator. We empirically show that WassersteinGAN can be used to output discrete labels. Later, we propose two new conditional GAN architectures, which we call CausalGAN and CausalBEGAN. We show that the optimal generator of the CausalGAN, given the labels, samples from the image distributions conditioned on these labels. The conditional GAN combined with a trained causal implicit generative model for the labels is then a causal implicit generative model over the labels and the generated image. We show that the proposed architectures can be used to sample from observational and interventional image distributions, even for interventions which do not naturally occur in the dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge