Christoph Dellago
Enhanced Sampling of Configuration and Path Space in a Generalized Ensemble by Shooting Point Exchange
Feb 17, 2023Abstract:The computer simulation of many molecular processes is complicated by long time scales caused by rare transitions between long-lived states. Here, we propose a new approach to simulate such rare events, which combines transition path sampling with enhanced exploration of configuration space. The method relies on exchange moves between configuration and trajectory space, carried out based on a generalized ensemble. This scheme substantially enhances the efficiency of the transition path sampling simulations, particularly for systems with multiple transition channels, and yields information on thermodynamics, kinetics and reaction coordinates of molecular processes without distorting their dynamics. The method is illustrated using the isomerization of proline in the KPTP tetrapeptide.
Conditioning Normalizing Flows for Rare Event Sampling
Jul 29, 2022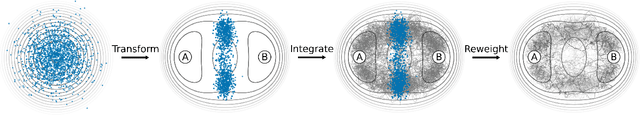
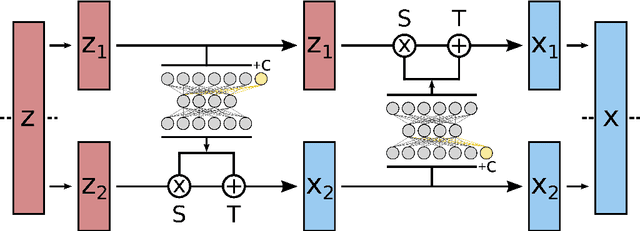
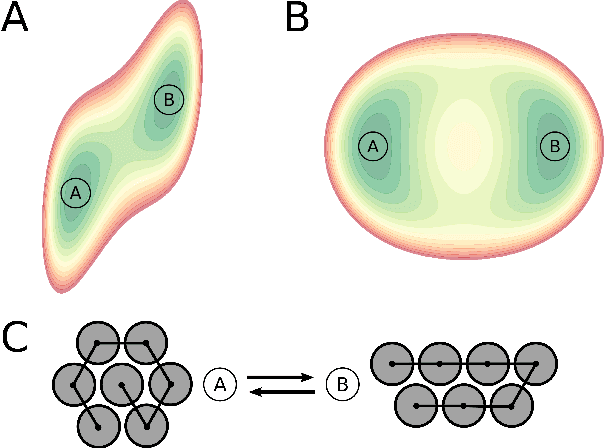
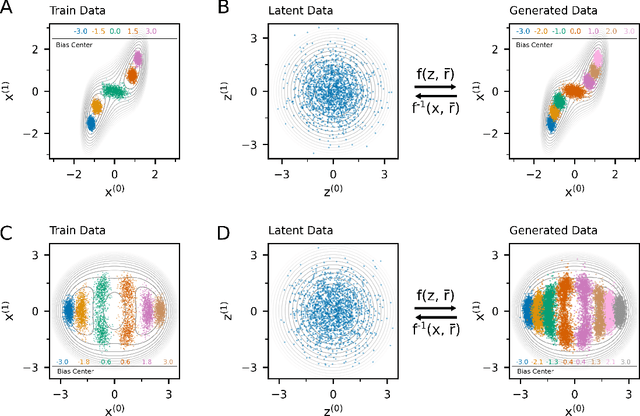
Abstract:Understanding the dynamics of complex molecular processes is often linked to the study of infrequent transitions between long-lived stable states. The standard approach to the sampling of such rare events is to generate an ensemble of transition paths using a random walk in trajectory space. This, however, comes with the drawback of strong correlation between subsequently visited paths and with an intrinsic difficulty in parallelizing the sampling process. We propose a transition path sampling scheme based on neural-network generated configurations. These are obtained employing normalizing flows, a neural network class able to generate decorrelated samples from a given distribution. With this approach, not only are correlations between visited paths removed, but the sampling process becomes easily parallelizable. Moreover, by conditioning the normalizing flow, the sampling of configurations can be steered towards the regions of interest. We show that this allows for resolving both the thermodynamics and kinetics of the transition region.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge