Christian A. Naesseth
A Variational Perspective on Generative Flow Networks
Oct 14, 2022
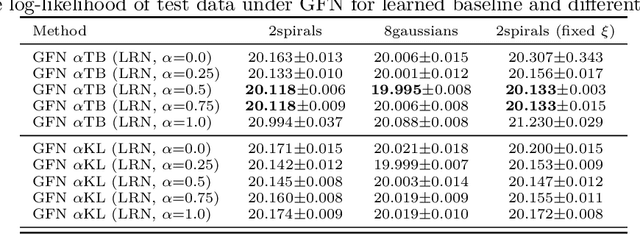
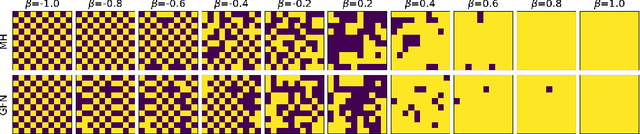
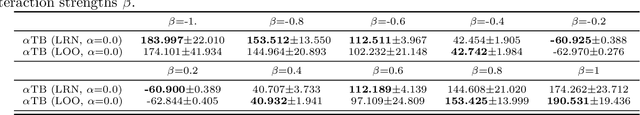
Abstract:Generative flow networks (GFNs) are a class of models for sequential sampling of composite objects, which approximate a target distribution that is defined in terms of an energy function or a reward. GFNs are typically trained using a flow matching or trajectory balance objective, which matches forward and backward transition models over trajectories. In this work, we define variational objectives for GFNs in terms of the Kullback-Leibler (KL) divergences between the forward and backward distribution. We show that variational inference in GFNs is equivalent to minimizing the trajectory balance objective when sampling trajectories from the forward model. We generalize this approach by optimizing a convex combination of the reverse- and forward KL divergence. This insight suggests variational inference methods can serve as a means to define a more general family of objectives for training generative flow networks, for example by incorporating control variates, which are commonly used in variational inference, to reduce the variance of the gradients of the trajectory balance objective. We evaluate our findings and the performance of the proposed variational objective numerically by comparing it to the trajectory balance objective on two synthetic tasks.
Transport Score Climbing: Variational Inference Using Forward KL and Adaptive Neural Transport
Feb 07, 2022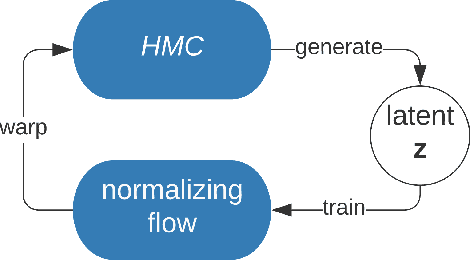
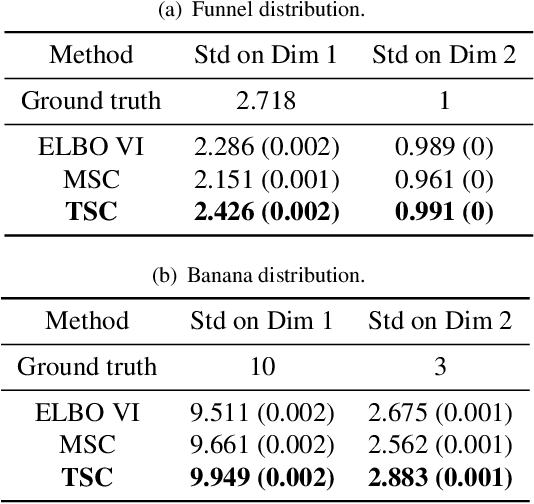
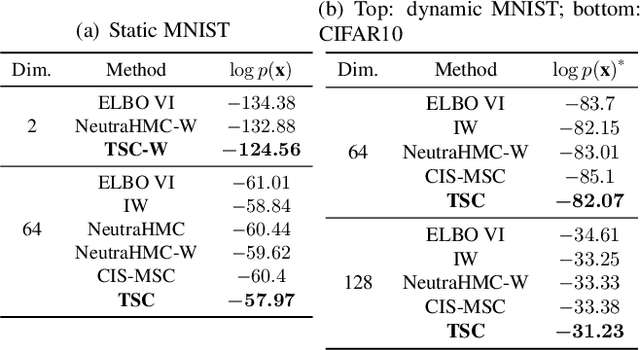
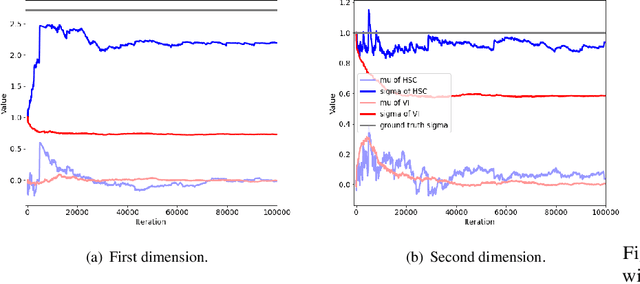
Abstract:Variational inference often minimizes the "reverse" Kullbeck-Leibler (KL) KL(q||p) from the approximate distribution q to the posterior p. Recent work studies the "forward" KL KL(p||q), which unlike reverse KL does not lead to variational approximations that underestimate uncertainty. This paper introduces Transport Score Climbing (TSC), a method that optimizes KL(p||q) by using Hamiltonian Monte Carlo (HMC) and a novel adaptive transport map. The transport map improves the trajectory of HMC by acting as a change of variable between the latent variable space and a warped space. TSC uses HMC samples to dynamically train the transport map while optimizing KL(p||q). TSC leverages synergies, where better transport maps lead to better HMC sampling, which then leads to better transport maps. We demonstrate TSC on synthetic and real data. We find that TSC achieves competitive performance when training variational autoencoders on large-scale data.
Variational Combinatorial Sequential Monte Carlo Methods for Bayesian Phylogenetic Inference
Jun 17, 2021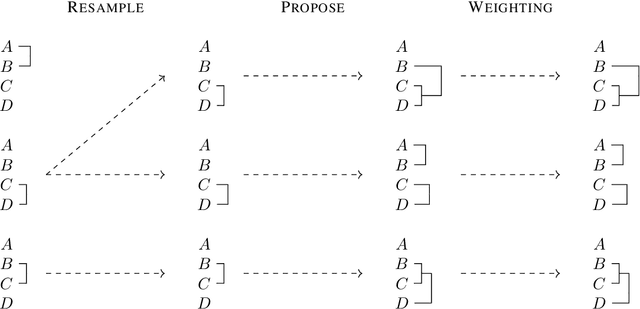
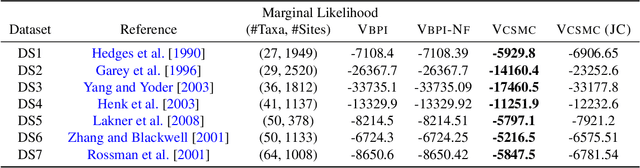
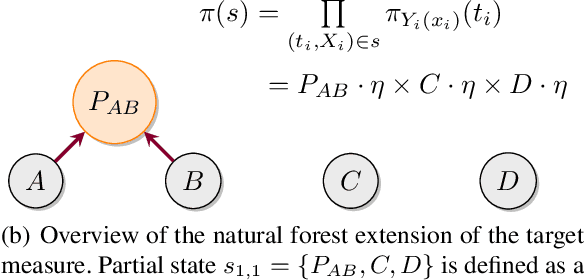
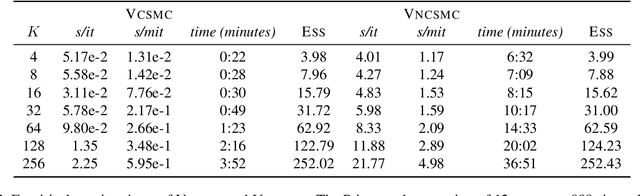
Abstract:Bayesian phylogenetic inference is often conducted via local or sequential search over topologies and branch lengths using algorithms such as random-walk Markov chain Monte Carlo (MCMC) or Combinatorial Sequential Monte Carlo (CSMC). However, when MCMC is used for evolutionary parameter learning, convergence requires long runs with inefficient exploration of the state space. We introduce Variational Combinatorial Sequential Monte Carlo (VCSMC), a powerful framework that establishes variational sequential search to learn distributions over intricate combinatorial structures. We then develop nested CSMC, an efficient proposal distribution for CSMC and prove that nested CSMC is an exact approximation to the (intractable) locally optimal proposal. We use nested CSMC to define a second objective, VNCSMC which yields tighter lower bounds than VCSMC. We show that VCSMC and VNCSMC are computationally efficient and explore higher probability spaces than existing methods on a range of tasks.
Markovian Score Climbing: Variational Inference with KL(p||q)
Mar 23, 2020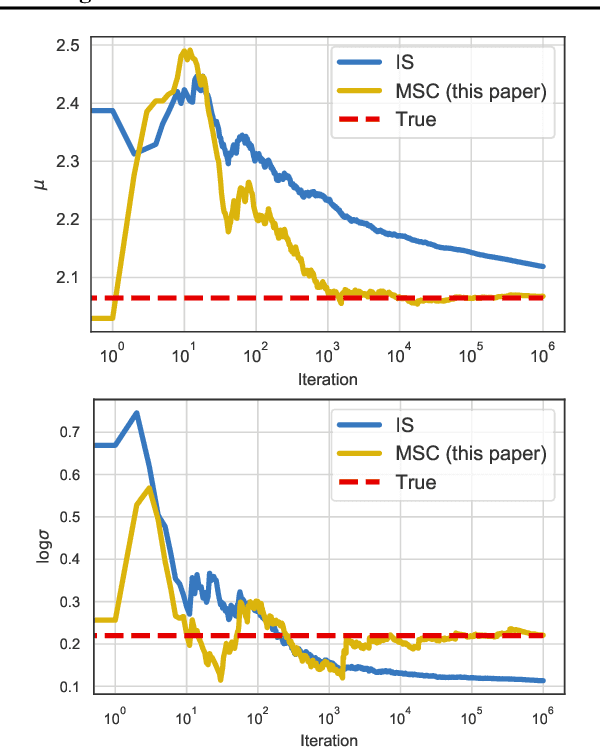
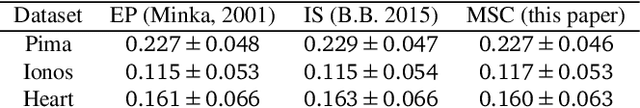

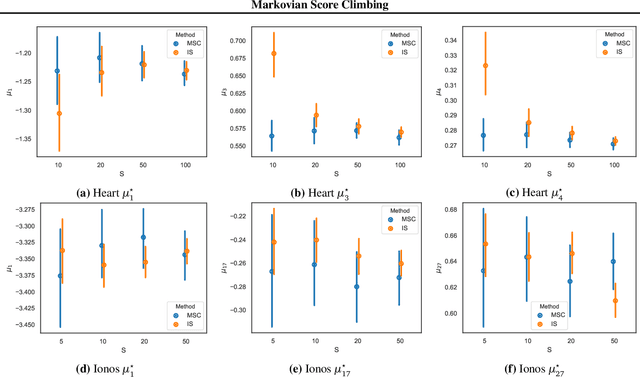
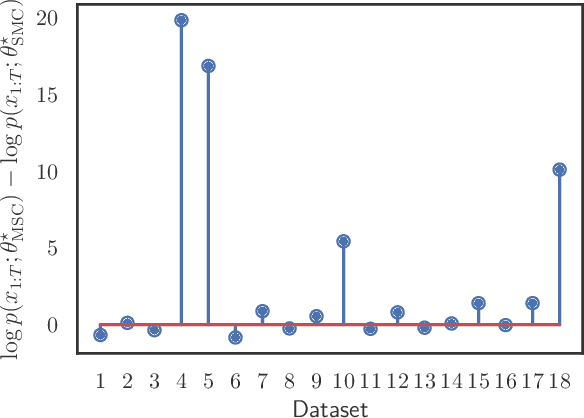
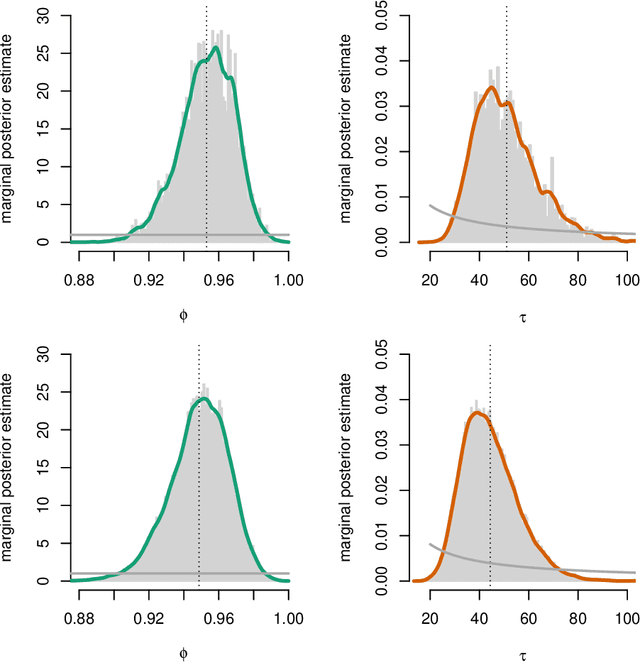
Abstract:Modern variational inference (VI) uses stochastic gradients to avoid intractable expectations, enabling large-scale probabilistic inference in complex models. VI posits a family of approximating distributions $q$ and then finds the member of that family that is closest to the exact posterior $p$. Traditionally, VI algorithms minimize the "exclusive KL" KL$(q\|p)$, often for computational convenience. Recent research, however, has also focused on the "inclusive KL" KL$(p\|q)$, which has good statistical properties that makes it more appropriate for certain inference problems. This paper develops a simple algorithm for reliably minimizing the inclusive KL. Consider a valid MCMC method, a Markov chain whose stationary distribution is $p$. The algorithm we develop iteratively samples the chain $z[k]$, and then uses those samples to follow the score function of the variational approximation, $\nabla \log q(z[k])$ with a Robbins-Monro step-size schedule. This method, which we call Markovian score climbing (MSC), converges to a local optimum of the inclusive KL. It does not suffer from the systematic errors inherent in existing methods, such as Reweighted Wake-Sleep and Neural Adaptive Sequential Monte Carlo, which lead to bias in their final estimates. In a variant that ties the variational approximation directly to the Markov chain, MSC further provides a new algorithm that melds VI and MCMC. We illustrate convergence on a toy model and demonstrate the utility of MSC on Bayesian probit regression for classification as well as a stochastic volatility model for financial data.
Elements of Sequential Monte Carlo
Mar 12, 2019
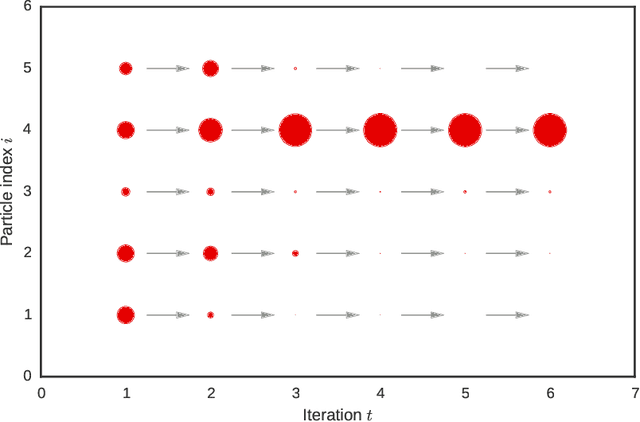
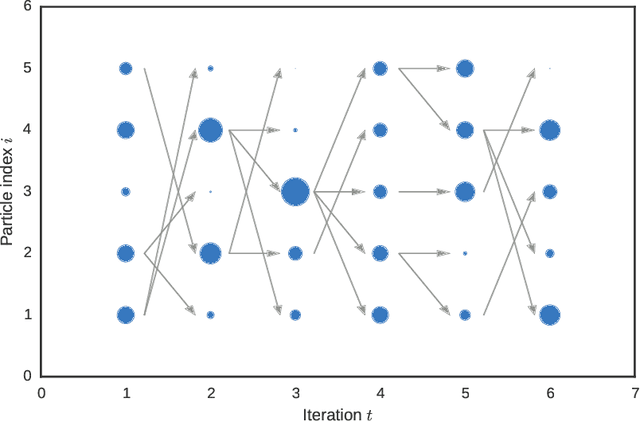
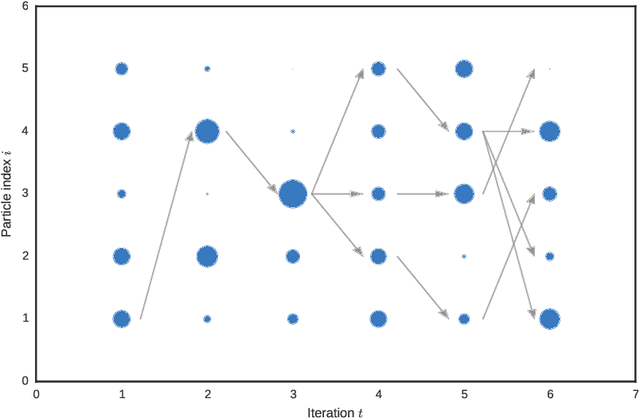
Abstract:A core problem in statistics and probabilistic machine learning is to compute probability distributions and expectations. This is the fundamental problem of Bayesian statistics and machine learning, which frames all inference as expectations with respect to the posterior distribution. The key challenge is to approximate these intractable expectations. In this tutorial, we review sequential Monte Carlo (SMC), a random-sampling-based class of methods for approximate inference. First, we explain the basics of SMC, discuss practical issues, and review theoretical results. We then examine two of the main user design choices: the proposal distributions and the so called intermediate target distributions. We review recent results on how variational inference and amortization can be used to learn efficient proposals and target distributions. Next, we discuss the SMC estimate of the normalizing constant, how this can be used for pseudo-marginal inference and inference evaluation. Throughout the tutorial we illustrate the use of SMC on various models commonly used in machine learning, such as stochastic recurrent neural networks, probabilistic graphical models, and probabilistic programs.
Variational Sequential Monte Carlo
Feb 21, 2018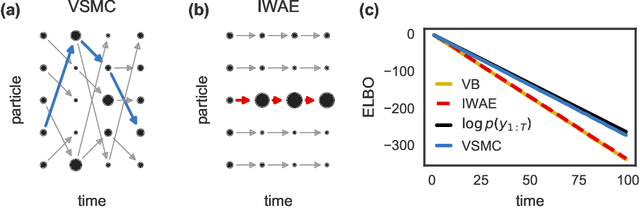
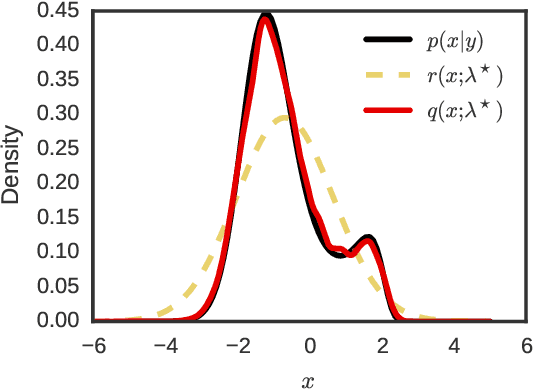
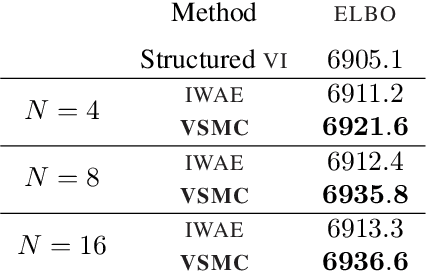
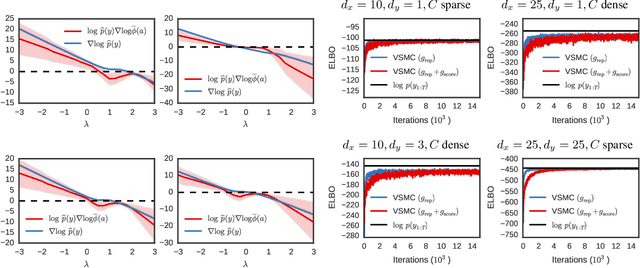
Abstract:Many recent advances in large scale probabilistic inference rely on variational methods. The success of variational approaches depends on (i) formulating a flexible parametric family of distributions, and (ii) optimizing the parameters to find the member of this family that most closely approximates the exact posterior. In this paper we present a new approximating family of distributions, the variational sequential Monte Carlo (VSMC) family, and show how to optimize it in variational inference. VSMC melds variational inference (VI) and sequential Monte Carlo (SMC), providing practitioners with flexible, accurate, and powerful Bayesian inference. The VSMC family is a variational family that can approximate the posterior arbitrarily well, while still allowing for efficient optimization of its parameters. We demonstrate its utility on state space models, stochastic volatility models for financial data, and deep Markov models of brain neural circuits.
Interacting Particle Markov Chain Monte Carlo
Apr 12, 2017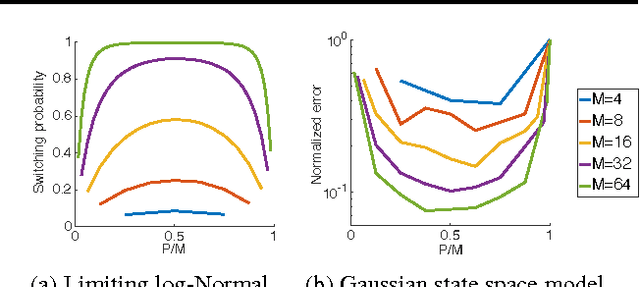
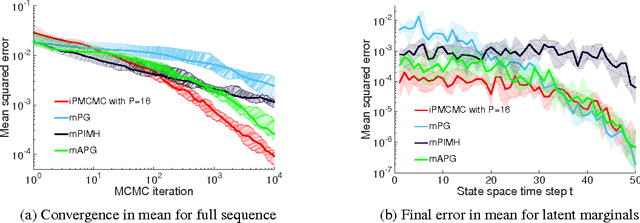
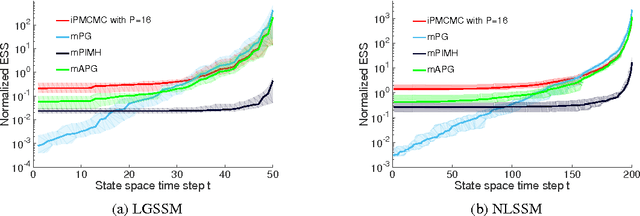
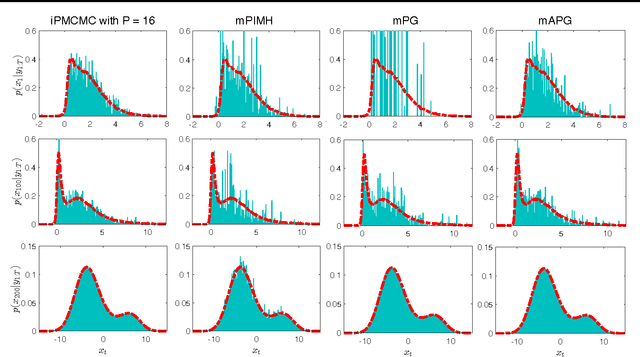
Abstract:We introduce interacting particle Markov chain Monte Carlo (iPMCMC), a PMCMC method based on an interacting pool of standard and conditional sequential Monte Carlo samplers. Like related methods, iPMCMC is a Markov chain Monte Carlo sampler on an extended space. We present empirical results that show significant improvements in mixing rates relative to both non-interacting PMCMC samplers, and a single PMCMC sampler with an equivalent memory and computational budget. An additional advantage of the iPMCMC method is that it is suitable for distributed and multi-core architectures.
Reparameterization Gradients through Acceptance-Rejection Sampling Algorithms
Mar 10, 2017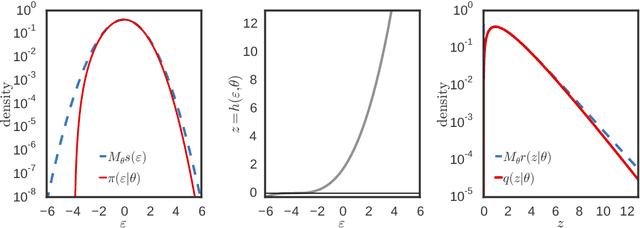
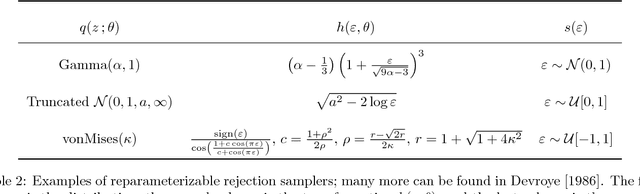

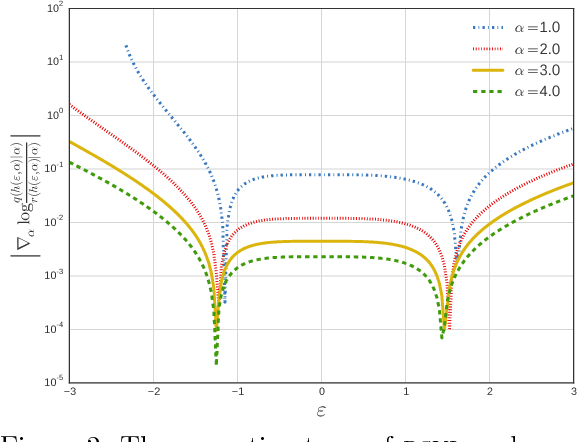

Abstract:Variational inference using the reparameterization trick has enabled large-scale approximate Bayesian inference in complex probabilistic models, leveraging stochastic optimization to sidestep intractable expectations. The reparameterization trick is applicable when we can simulate a random variable by applying a differentiable deterministic function on an auxiliary random variable whose distribution is fixed. For many distributions of interest (such as the gamma or Dirichlet), simulation of random variables relies on acceptance-rejection sampling. The discontinuity introduced by the accept-reject step means that standard reparameterization tricks are not applicable. We propose a new method that lets us leverage reparameterization gradients even when variables are outputs of a acceptance-rejection sampling algorithm. Our approach enables reparameterization on a larger class of variational distributions. In several studies of real and synthetic data, we show that the variance of the estimator of the gradient is significantly lower than other state-of-the-art methods. This leads to faster convergence of stochastic gradient variational inference.
High-dimensional Filtering using Nested Sequential Monte Carlo
Dec 29, 2016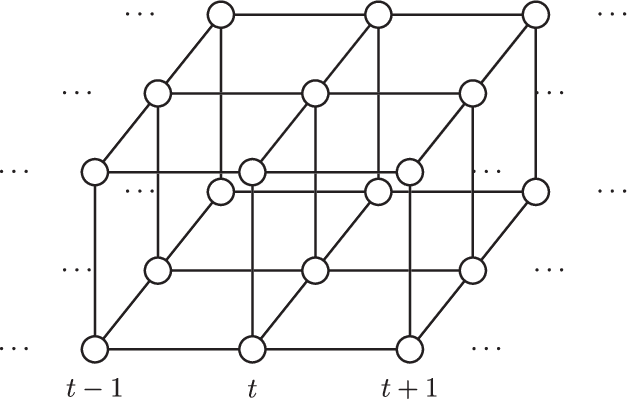
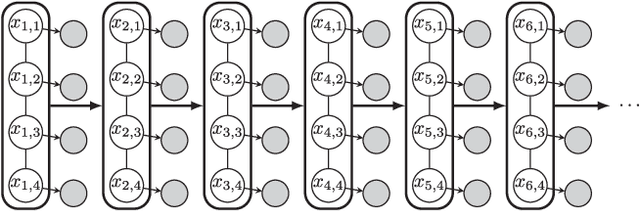
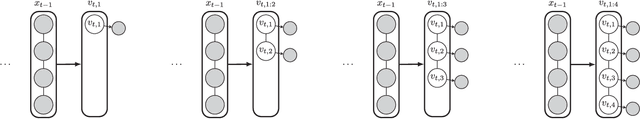
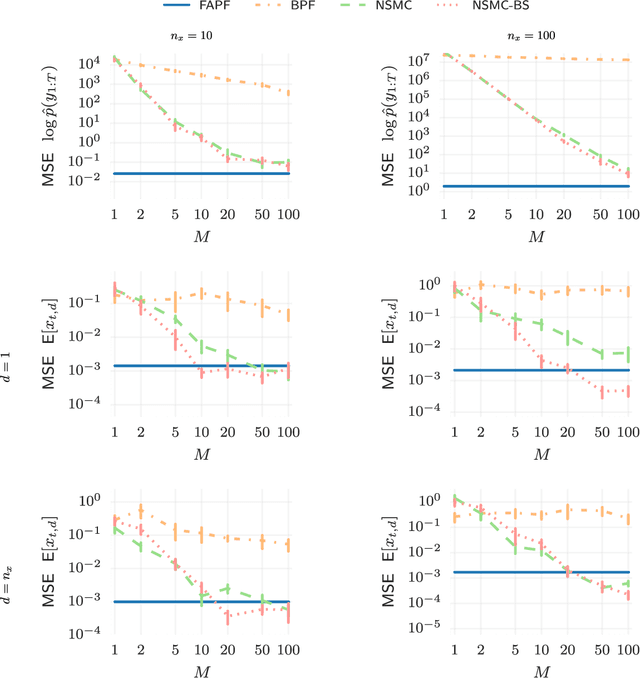
Abstract:Sequential Monte Carlo (SMC) methods comprise one of the most successful approaches to approximate Bayesian filtering. However, SMC without good proposal distributions struggle in high dimensions. We propose nested sequential Monte Carlo (NSMC), a methodology that generalises the SMC framework by requiring only approximate, properly weighted, samples from the SMC proposal distribution, while still resulting in a correct SMC algorithm. This way we can exactly approximate the locally optimal proposal, and extend the class of models for which we can perform efficient inference using SMC. We show improved accuracy over other state-of-the-art methods on several spatio-temporal state space models.
Sequential Monte Carlo Methods for System Identification
Mar 10, 2016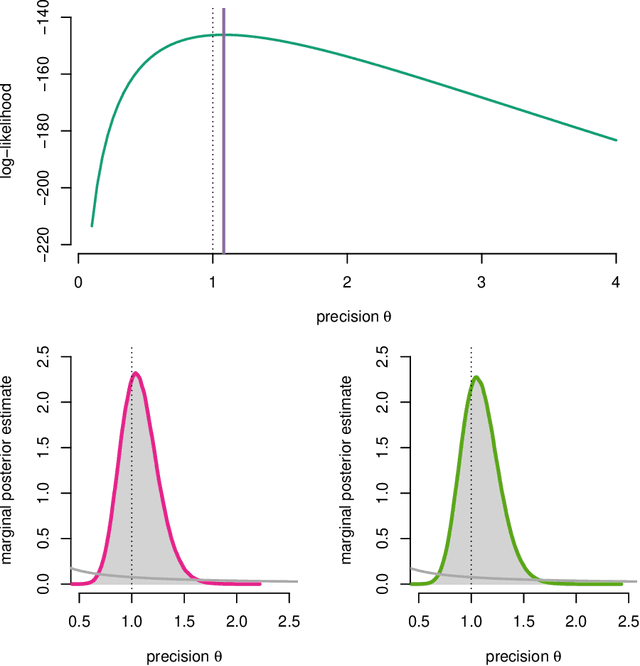
Abstract:One of the key challenges in identifying nonlinear and possibly non-Gaussian state space models (SSMs) is the intractability of estimating the system state. Sequential Monte Carlo (SMC) methods, such as the particle filter (introduced more than two decades ago), provide numerical solutions to the nonlinear state estimation problems arising in SSMs. When combined with additional identification techniques, these algorithms provide solid solutions to the nonlinear system identification problem. We describe two general strategies for creating such combinations and discuss why SMC is a natural tool for implementing these strategies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge