Chris Holmes
Explainable AI for survival analysis: a median-SHAP approach
Jan 30, 2024Abstract:With the adoption of machine learning into routine clinical practice comes the need for Explainable AI methods tailored to medical applications. Shapley values have sparked wide interest for locally explaining models. Here, we demonstrate their interpretation strongly depends on both the summary statistic and the estimator for it, which in turn define what we identify as an 'anchor point'. We show that the convention of using a mean anchor point may generate misleading interpretations for survival analysis and introduce median-SHAP, a method for explaining black-box models predicting individual survival times.
Targeting Relative Risk Heterogeneity with Causal Forests
Sep 26, 2023



Abstract:Treatment effect heterogeneity (TEH), or variability in treatment effect for different subgroups within a population, is of significant interest in clinical trial analysis. Causal forests (Wager and Athey, 2018) is a highly popular method for this problem, but like many other methods for detecting TEH, its criterion for separating subgroups focuses on differences in absolute risk. This can dilute statistical power by masking nuance in the relative risk, which is often a more appropriate quantity of clinical interest. In this work, we propose and implement a methodology for modifying causal forests to target relative risk using a novel node-splitting procedure based on generalized linear model (GLM) comparison. We present results on simulated and real-world data that suggest relative risk causal forests can capture otherwise unobserved sources of heterogeneity.
Differentially Private Statistical Inference through $β$-Divergence One Posterior Sampling
Jul 11, 2023



Abstract:Differential privacy guarantees allow the results of a statistical analysis involving sensitive data to be released without compromising the privacy of any individual taking part. Achieving such guarantees generally requires the injection of noise, either directly into parameter estimates or into the estimation process. Instead of artificially introducing perturbations, sampling from Bayesian posterior distributions has been shown to be a special case of the exponential mechanism, producing consistent, and efficient private estimates without altering the data generative process. The application of current approaches has, however, been limited by their strong bounding assumptions which do not hold for basic models, such as simple linear regressors. To ameliorate this, we propose $\beta$D-Bayes, a posterior sampling scheme from a generalised posterior targeting the minimisation of the $\beta$-divergence between the model and the data generating process. This provides private estimation that is generally applicable without requiring changes to the underlying model and consistently learns the data generating parameter. We show that $\beta$D-Bayes produces more precise inference estimation for the same privacy guarantees, and further facilitates differentially private estimation via posterior sampling for complex classifiers and continuous regression models such as neural networks for the first time.
PWSHAP: A Path-Wise Explanation Model for Targeted Variables
Jun 26, 2023



Abstract:Predictive black-box models can exhibit high accuracy but their opaque nature hinders their uptake in safety-critical deployment environments. Explanation methods (XAI) can provide confidence for decision-making through increased transparency. However, existing XAI methods are not tailored towards models in sensitive domains where one predictor is of special interest, such as a treatment effect in a clinical model, or ethnicity in policy models. We introduce Path-Wise Shapley effects (PWSHAP), a framework for assessing the targeted effect of a binary (e.g.~treatment) variable from a complex outcome model. Our approach augments the predictive model with a user-defined directed acyclic graph (DAG). The method then uses the graph alongside on-manifold Shapley values to identify effects along causal pathways whilst maintaining robustness to adversarial attacks. We establish error bounds for the identified path-wise Shapley effects and for Shapley values. We show PWSHAP can perform local bias and mediation analyses with faithfulness to the model. Further, if the targeted variable is randomised we can quantify local effect modification. We demonstrate the resolution, interpretability, and true locality of our approach on examples and a real-world experiment.
A Unified Framework for U-Net Design and Analysis
May 31, 2023



Abstract:U-Nets are a go-to, state-of-the-art neural architecture across numerous tasks for continuous signals on a square such as images and Partial Differential Equations (PDE), however their design and architecture is understudied. In this paper, we provide a framework for designing and analysing general U-Net architectures. We present theoretical results which characterise the role of the encoder and decoder in a U-Net, their high-resolution scaling limits and their conjugacy to ResNets via preconditioning. We propose Multi-ResNets, U-Nets with a simplified, wavelet-based encoder without learnable parameters. Further, we show how to design novel U-Net architectures which encode function constraints, natural bases, or the geometry of the data. In diffusion models, our framework enables us to identify that high-frequency information is dominated by noise exponentially faster, and show how U-Nets with average pooling exploit this. In our experiments, we demonstrate how Multi-ResNets achieve competitive and often superior performance compared to classical U-Nets in image segmentation, PDE surrogate modelling, and generative modelling with diffusion models. Our U-Net framework paves the way to study the theoretical properties of U-Nets and design natural, scalable neural architectures for a multitude of problems beyond the square.
Learning from data with structured missingness
Apr 04, 2023Abstract:Missing data are an unavoidable complication in many machine learning tasks. When data are `missing at random' there exist a range of tools and techniques to deal with the issue. However, as machine learning studies become more ambitious, and seek to learn from ever-larger volumes of heterogeneous data, an increasingly encountered problem arises in which missing values exhibit an association or structure, either explicitly or implicitly. Such `structured missingness' raises a range of challenges that have not yet been systematically addressed, and presents a fundamental hindrance to machine learning at scale. Here, we outline the current literature and propose a set of grand challenges in learning from data with structured missingness.
Causal Falsification of Digital Twins
Jan 19, 2023Abstract:Digital twins hold substantial promise in many applications, but rigorous procedures for assessing their accuracy are essential for their widespread deployment in safety-critical settings. By formulating this task within the framework of causal inference, we show it is not possible to certify that a twin is "correct" using real-world observational data unless potentially tenuous assumptions are made about the data-generating process. To avoid these assumptions, we propose an assessment strategy that instead aims to find cases where the twin is not correct, and present a general-purpose statistical procedure for doing so that may be used across a wide variety of applications and twin models. Our approach yields reliable and actionable information about the twin under only the assumption of an i.i.d. dataset of real-world observations, and in particular remains sound even in the presence of arbitrary unmeasured confounding. We demonstrate the effectiveness of our methodology via a large-scale case study involving sepsis modelling within the Pulse Physiology Engine, which we assess using the MIMIC-III dataset of ICU patients.
A Multi-Resolution Framework for U-Nets with Applications to Hierarchical VAEs
Jan 19, 2023Abstract:U-Net architectures are ubiquitous in state-of-the-art deep learning, however their regularisation properties and relationship to wavelets are understudied. In this paper, we formulate a multi-resolution framework which identifies U-Nets as finite-dimensional truncations of models on an infinite-dimensional function space. We provide theoretical results which prove that average pooling corresponds to projection within the space of square-integrable functions and show that U-Nets with average pooling implicitly learn a Haar wavelet basis representation of the data. We then leverage our framework to identify state-of-the-art hierarchical VAEs (HVAEs), which have a U-Net architecture, as a type of two-step forward Euler discretisation of multi-resolution diffusion processes which flow from a point mass, introducing sampling instabilities. We also demonstrate that HVAEs learn a representation of time which allows for improved parameter efficiency through weight-sharing. We use this observation to achieve state-of-the-art HVAE performance with half the number of parameters of existing models, exploiting the properties of our continuous-time formulation.
Statistical Design and Analysis for Robust Machine Learning: A Case Study from COVID-19
Dec 15, 2022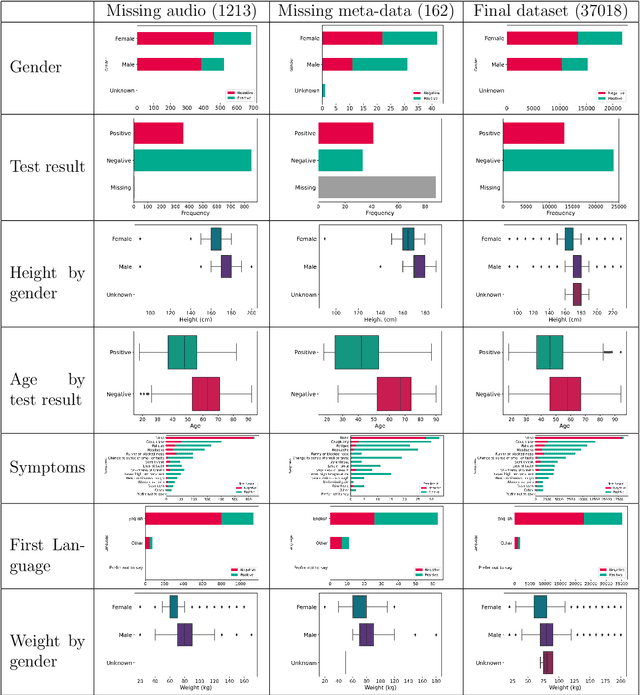
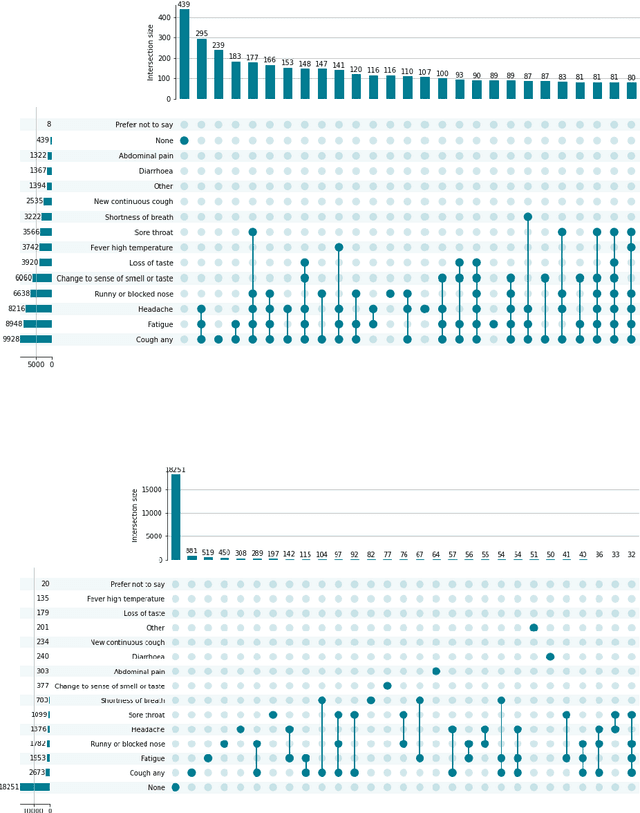

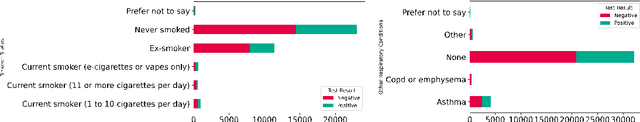
Abstract:Since early in the coronavirus disease 2019 (COVID-19) pandemic, there has been interest in using artificial intelligence methods to predict COVID-19 infection status based on vocal audio signals, for example cough recordings. However, existing studies have limitations in terms of data collection and of the assessment of the performances of the proposed predictive models. This paper rigorously assesses state-of-the-art machine learning techniques used to predict COVID-19 infection status based on vocal audio signals, using a dataset collected by the UK Health Security Agency. This dataset includes acoustic recordings and extensive study participant meta-data. We provide guidelines on testing the performance of methods to classify COVID-19 infection status based on acoustic features and we discuss how these can be extended more generally to the development and assessment of predictive methods based on public health datasets.
Audio-based AI classifiers show no evidence of improved COVID-19 screening over simple symptoms checkers
Dec 15, 2022



Abstract:Recent work has reported that AI classifiers trained on audio recordings can accurately predict severe acute respiratory syndrome coronavirus 2 (SARSCoV2) infection status. Here, we undertake a large scale study of audio-based deep learning classifiers, as part of the UK governments pandemic response. We collect and analyse a dataset of audio recordings from 67,842 individuals with linked metadata, including reverse transcription polymerase chain reaction (PCR) test outcomes, of whom 23,514 tested positive for SARS CoV 2. Subjects were recruited via the UK governments National Health Service Test-and-Trace programme and the REal-time Assessment of Community Transmission (REACT) randomised surveillance survey. In an unadjusted analysis of our dataset AI classifiers predict SARS-CoV-2 infection status with high accuracy (Receiver Operating Characteristic Area Under the Curve (ROCAUC) 0.846 [0.838, 0.854]) consistent with the findings of previous studies. However, after matching on measured confounders, such as age, gender, and self reported symptoms, our classifiers performance is much weaker (ROC-AUC 0.619 [0.594, 0.644]). Upon quantifying the utility of audio based classifiers in practical settings, we find them to be outperformed by simple predictive scores based on user reported symptoms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge