Lucile Ter-Minassian
Is merging worth it? Securely evaluating the information gain for causal dataset acquisition
Sep 11, 2024



Abstract:Merging datasets across institutions is a lengthy and costly procedure, especially when it involves private information. Data hosts may therefore want to prospectively gauge which datasets are most beneficial to merge with, without revealing sensitive information. For causal estimation this is particularly challenging as the value of a merge will depend not only on the reduction in epistemic uncertainty but also the improvement in overlap. To address this challenge, we introduce the first cryptographically secure information-theoretic approach for quantifying the value of a merge in the context of heterogeneous treatment effect estimation. We do this by evaluating the Expected Information Gain (EIG) and utilising multi-party computation to ensure it can be securely computed without revealing any raw data. As we demonstrate, this can be used with differential privacy (DP) to ensure privacy requirements whilst preserving more accurate computation than naive DP alone. To the best of our knowledge, this work presents the first privacy-preserving method for dataset acquisition tailored to causal estimation. We demonstrate the effectiveness and reliability of our method on a range of simulated and realistic benchmarks. The code is available anonymously.
Extracting Paragraphs from LLM Token Activations
Sep 10, 2024



Abstract:Generative large language models (LLMs) excel in natural language processing tasks, yet their inner workings remain underexplored beyond token-level predictions. This study investigates the degree to which these models decide the content of a paragraph at its onset, shedding light on their contextual understanding. By examining the information encoded in single-token activations, specifically the "\textbackslash n\textbackslash n" double newline token, we demonstrate that patching these activations can transfer significant information about the context of the following paragraph, providing further insights into the model's capacity to plan ahead.
Hierarchical Bias-Driven Stratification for Interpretable Causal Effect Estimation
Jan 31, 2024Abstract:Interpretability and transparency are essential for incorporating causal effect models from observational data into policy decision-making. They can provide trust for the model in the absence of ground truth labels to evaluate the accuracy of such models. To date, attempts at transparent causal effect estimation consist of applying post hoc explanation methods to black-box models, which are not interpretable. Here, we present BICauseTree: an interpretable balancing method that identifies clusters where natural experiments occur locally. Our approach builds on decision trees with a customized objective function to improve balancing and reduce treatment allocation bias. Consequently, it can additionally detect subgroups presenting positivity violations, exclude them, and provide a covariate-based definition of the target population we can infer from and generalize to. We evaluate the method's performance using synthetic and realistic datasets, explore its bias-interpretability tradeoff, and show that it is comparable with existing approaches.
Explainable AI for survival analysis: a median-SHAP approach
Jan 30, 2024Abstract:With the adoption of machine learning into routine clinical practice comes the need for Explainable AI methods tailored to medical applications. Shapley values have sparked wide interest for locally explaining models. Here, we demonstrate their interpretation strongly depends on both the summary statistic and the estimator for it, which in turn define what we identify as an 'anchor point'. We show that the convention of using a mean anchor point may generate misleading interpretations for survival analysis and introduce median-SHAP, a method for explaining black-box models predicting individual survival times.
PWSHAP: A Path-Wise Explanation Model for Targeted Variables
Jun 26, 2023



Abstract:Predictive black-box models can exhibit high accuracy but their opaque nature hinders their uptake in safety-critical deployment environments. Explanation methods (XAI) can provide confidence for decision-making through increased transparency. However, existing XAI methods are not tailored towards models in sensitive domains where one predictor is of special interest, such as a treatment effect in a clinical model, or ethnicity in policy models. We introduce Path-Wise Shapley effects (PWSHAP), a framework for assessing the targeted effect of a binary (e.g.~treatment) variable from a complex outcome model. Our approach augments the predictive model with a user-defined directed acyclic graph (DAG). The method then uses the graph alongside on-manifold Shapley values to identify effects along causal pathways whilst maintaining robustness to adversarial attacks. We establish error bounds for the identified path-wise Shapley effects and for Shapley values. We show PWSHAP can perform local bias and mediation analyses with faithfulness to the model. Further, if the targeted variable is randomised we can quantify local effect modification. We demonstrate the resolution, interpretability, and true locality of our approach on examples and a real-world experiment.
On Locality of Local Explanation Models
Jun 24, 2021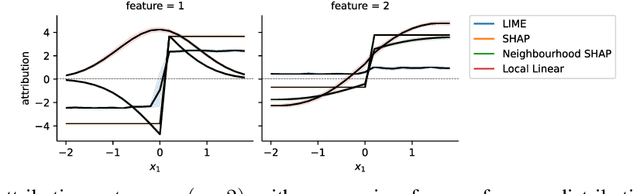
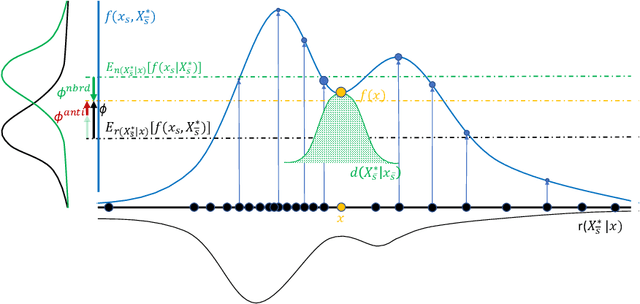

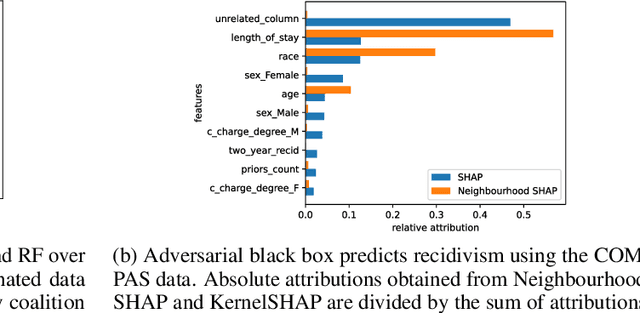
Abstract:Shapley values provide model agnostic feature attributions for model outcome at a particular instance by simulating feature absence under a global population distribution. The use of a global population can lead to potentially misleading results when local model behaviour is of interest. Hence we consider the formulation of neighbourhood reference distributions that improve the local interpretability of Shapley values. By doing so, we find that the Nadaraya-Watson estimator, a well-studied kernel regressor, can be expressed as a self-normalised importance sampling estimator. Empirically, we observe that Neighbourhood Shapley values identify meaningful sparse feature relevance attributions that provide insight into local model behaviour, complimenting conventional Shapley analysis. They also increase on-manifold explainability and robustness to the construction of adversarial classifiers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge