Chinasa Kalaiwo
Towards Trustworthy Breast Tumor Segmentation in Ultrasound using Monte Carlo Dropout and Deep Ensembles for Epistemic Uncertainty Estimation
Aug 25, 2025
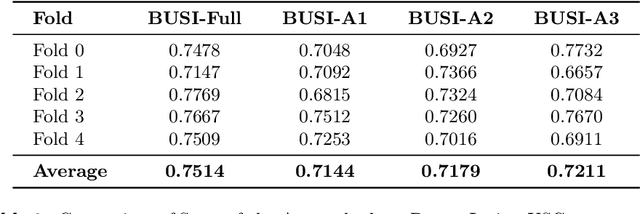
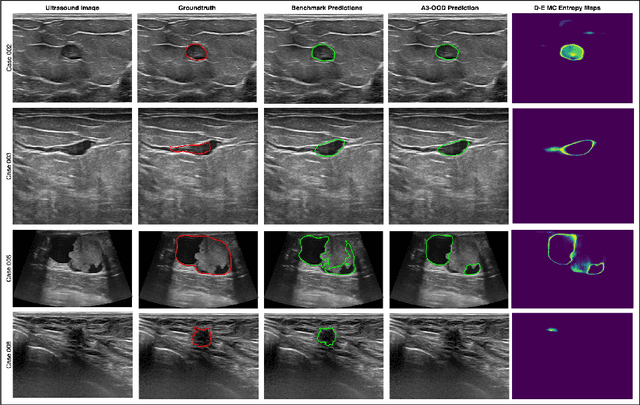
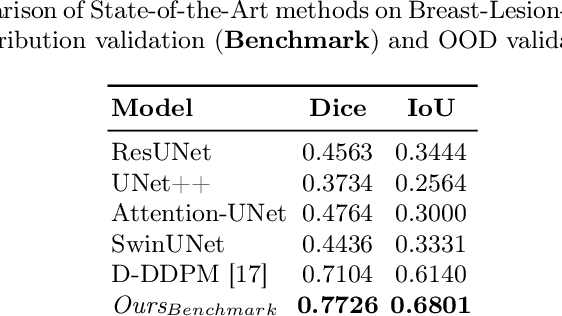
Abstract:Automated segmentation of BUS images is important for precise lesion delineation and tumor characterization, but is challenged by inherent artifacts and dataset inconsistencies. In this work, we evaluate the use of a modified Residual Encoder U-Net for breast ultrasound segmentation, with a focus on uncertainty quantification. We identify and correct for data duplication in the BUSI dataset, and use a deduplicated subset for more reliable estimates of generalization performance. Epistemic uncertainty is quantified using Monte Carlo dropout, deep ensembles, and their combination. Models are benchmarked on both in-distribution and out-of-distribution datasets to demonstrate how they generalize to unseen cross-domain data. Our approach achieves state-of-the-art segmentation accuracy on the Breast-Lesion-USG dataset with in-distribution validation, and provides calibrated uncertainty estimates that effectively signal regions of low model confidence. Performance declines and increased uncertainty observed in out-of-distribution evaluation highlight the persistent challenge of domain shift in medical imaging, and the importance of integrated uncertainty modeling for trustworthy clinical deployment. \footnote{Code available at: https://github.com/toufiqmusah/nn-uncertainty.git}
The Brain Tumor Segmentation Challenge 2023: Glioma Segmentation in Sub-Saharan Africa Patient Population
May 30, 2023

Abstract:Gliomas are the most common type of primary brain tumors. Although gliomas are relatively rare, they are among the deadliest types of cancer, with a survival rate of less than 2 years after diagnosis. Gliomas are challenging to diagnose, hard to treat and inherently resistant to conventional therapy. Years of extensive research to improve diagnosis and treatment of gliomas have decreased mortality rates across the Global North, while chances of survival among individuals in low- and middle-income countries (LMICs) remain unchanged and are significantly worse in Sub-Saharan Africa (SSA) populations. Long-term survival with glioma is associated with the identification of appropriate pathological features on brain MRI and confirmation by histopathology. Since 2012, the Brain Tumor Segmentation (BraTS) Challenge have evaluated state-of-the-art machine learning methods to detect, characterize, and classify gliomas. However, it is unclear if the state-of-the-art methods can be widely implemented in SSA given the extensive use of lower-quality MRI technology, which produces poor image contrast and resolution and more importantly, the propensity for late presentation of disease at advanced stages as well as the unique characteristics of gliomas in SSA (i.e., suspected higher rates of gliomatosis cerebri). Thus, the BraTS-Africa Challenge provides a unique opportunity to include brain MRI glioma cases from SSA in global efforts through the BraTS Challenge to develop and evaluate computer-aided-diagnostic (CAD) methods for the detection and characterization of glioma in resource-limited settings, where the potential for CAD tools to transform healthcare are more likely.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge