Chee-Ming Ting
FLAG-4D: Flow-Guided Local-Global Dual-Deformation Model for 4D Reconstruction
Feb 09, 2026Abstract:We introduce FLAG-4D, a novel framework for generating novel views of dynamic scenes by reconstructing how 3D Gaussian primitives evolve through space and time. Existing methods typically rely on a single Multilayer Perceptron (MLP) to model temporal deformations, and they often struggle to capture complex point motions and fine-grained dynamic details consistently over time, especially from sparse input views. Our approach, FLAG-4D, overcomes this by employing a dual-deformation network that dynamically warps a canonical set of 3D Gaussians over time into new positions and anisotropic shapes. This dual-deformation network consists of an Instantaneous Deformation Network (IDN) for modeling fine-grained, local deformations and a Global Motion Network (GMN) for capturing long-range dynamics, refined through mutual learning. To ensure these deformations are both accurate and temporally smooth, FLAG-4D incorporates dense motion features from a pretrained optical flow backbone. We fuse these motion cues from adjacent timeframes and use a deformation-guided attention mechanism to align this flow information with the current state of each evolving 3D Gaussian. Extensive experiments demonstrate that FLAG-4D achieves higher-fidelity and more temporally coherent reconstructions with finer detail preservation than state-of-the-art methods.
PanopMamba: Vision State Space Modeling for Nuclei Panoptic Segmentation
Jan 23, 2026Abstract:Nuclei panoptic segmentation supports cancer diagnostics by integrating both semantic and instance segmentation of different cell types to analyze overall tissue structure and individual nuclei in histopathology images. Major challenges include detecting small objects, handling ambiguous boundaries, and addressing class imbalance. To address these issues, we propose PanopMamba, a novel hybrid encoder-decoder architecture that integrates Mamba and Transformer with additional feature-enhanced fusion via state space modeling. We design a multiscale Mamba backbone and a State Space Model (SSM)-based fusion network to enable efficient long-range perception in pyramid features, thereby extending the pure encoder-decoder framework while facilitating information sharing across multiscale features of nuclei. The proposed SSM-based feature-enhanced fusion integrates pyramid feature networks and dynamic feature enhancement across different spatial scales, enhancing the feature representation of densely overlapping nuclei in both semantic and spatial dimensions. To the best of our knowledge, this is the first Mamba-based approach for panoptic segmentation. Additionally, we introduce alternative evaluation metrics, including image-level Panoptic Quality ($i$PQ), boundary-weighted PQ ($w$PQ), and frequency-weighted PQ ($fw$PQ), which are specifically designed to address the unique challenges of nuclei segmentation and thereby mitigate the potential bias inherent in vanilla PQ. Experimental evaluations on two multiclass nuclei segmentation benchmark datasets, MoNuSAC2020 and NuInsSeg, demonstrate the superiority of PanopMamba for nuclei panoptic segmentation over state-of-the-art methods. Consequently, the robustness of PanopMamba is validated across various metrics, while the distinctiveness of PQ variants is also demonstrated. Code is available at https://github.com/mkang315/PanopMamba.
Learning Energy-Based Generative Models via Potential Flow: A Variational Principle Approach to Probability Density Homotopy Matching
Apr 22, 2025Abstract:Energy-based models (EBMs) are a powerful class of probabilistic generative models due to their flexibility and interpretability. However, relationships between potential flows and explicit EBMs remain underexplored, while contrastive divergence training via implicit Markov chain Monte Carlo (MCMC) sampling is often unstable and expensive in high-dimensional settings. In this paper, we propose Variational Potential Flow Bayes (VPFB), a new energy-based generative framework that eliminates the need for implicit MCMC sampling and does not rely on auxiliary networks or cooperative training. VPFB learns an energy-parameterized potential flow by constructing a flow-driven density homotopy that is matched to the data distribution through a variational loss minimizing the Kullback-Leibler divergence between the flow-driven and marginal homotopies. This principled formulation enables robust and efficient generative modeling while preserving the interpretability of EBMs. Experimental results on image generation, interpolation, out-of-distribution detection, and compositional generation confirm the effectiveness of VPFB, showing that our method performs competitively with existing approaches in terms of sample quality and versatility across diverse generative modeling tasks.
SFC-GAN: A Generative Adversarial Network for Brain Functional and Structural Connectome Translation
Jan 13, 2025Abstract:Modern brain imaging technologies have enabled the detailed reconstruction of human brain connectomes, capturing structural connectivity (SC) from diffusion MRI and functional connectivity (FC) from functional MRI. Understanding the intricate relationships between SC and FC is vital for gaining deeper insights into the brain's functional and organizational mechanisms. However, obtaining both SC and FC modalities simultaneously remains challenging, hindering comprehensive analyses. Existing deep generative models typically focus on synthesizing a single modality or unidirectional translation between FC and SC, thereby missing the potential benefits of bi-directional translation, especially in scenarios where only one connectome is available. Therefore, we propose Structural-Functional Connectivity GAN (SFC-GAN), a novel framework for bidirectional translation between SC and FC. This approach leverages the CycleGAN architecture, incorporating convolutional layers to effectively capture the spatial structures of brain connectomes. To preserve the topological integrity of these connectomes, we employ a structure-preserving loss that guides the model in capturing both global and local connectome patterns while maintaining symmetry. Our framework demonstrates superior performance in translating between SC and FC, outperforming baseline models in similarity and graph property evaluations compared to ground truth data, each translated modality can be effectively utilized for downstream classification.
KANS: Knowledge Discovery Graph Attention Network for Soft Sensing in Multivariate Industrial Processes
Jan 02, 2025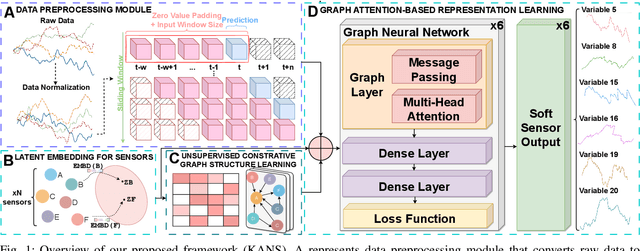
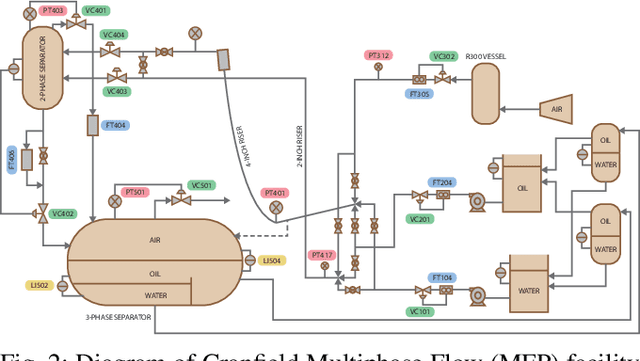
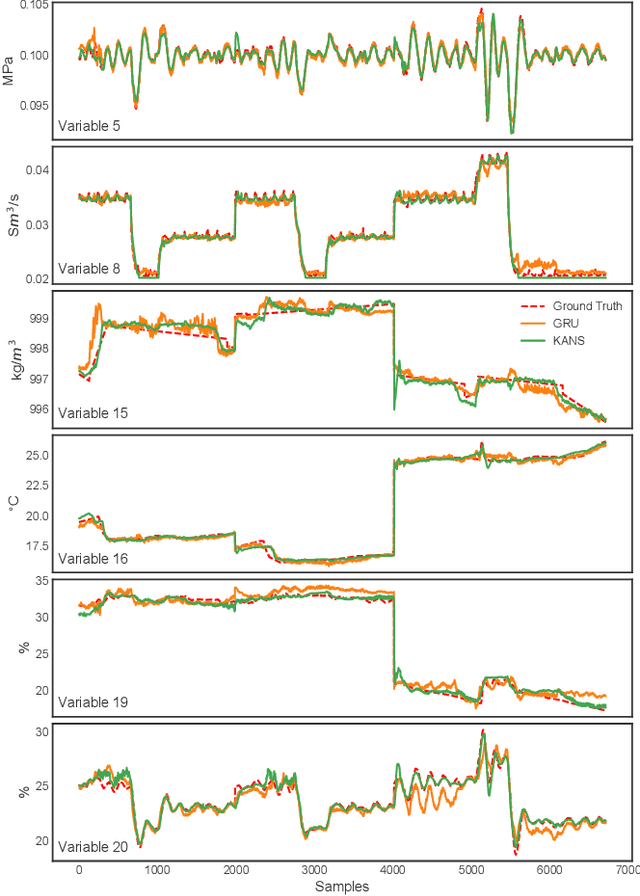
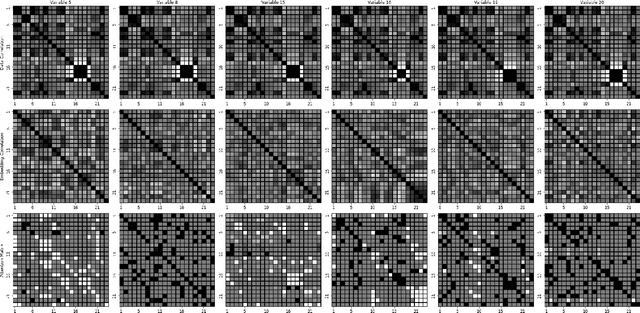
Abstract:Soft sensing of hard-to-measure variables is often crucial in industrial processes. Current practices rely heavily on conventional modeling techniques that show success in improving accuracy. However, they overlook the non-linear nature, dynamics characteristics, and non-Euclidean dependencies between complex process variables. To tackle these challenges, we present a framework known as a Knowledge discovery graph Attention Network for effective Soft sensing (KANS). Unlike the existing deep learning soft sensor models, KANS can discover the intrinsic correlations and irregular relationships between the multivariate industrial processes without a predefined topology. First, an unsupervised graph structure learning method is introduced, incorporating the cosine similarity between different sensor embedding to capture the correlations between sensors. Next, we present a graph attention-based representation learning that can compute the multivariate data parallelly to enhance the model in learning complex sensor nodes and edges. To fully explore KANS, knowledge discovery analysis has also been conducted to demonstrate the interpretability of the model. Experimental results demonstrate that KANS significantly outperforms all the baselines and state-of-the-art methods in soft sensing performance. Furthermore, the analysis shows that KANS can find sensors closely related to different process variables without domain knowledge, significantly improving soft sensing accuracy.
ST-HCSS: Deep Spatio-Temporal Hypergraph Convolutional Neural Network for Soft Sensing
Jan 02, 2025



Abstract:Higher-order sensor networks are more accurate in characterizing the nonlinear dynamics of sensory time-series data in modern industrial settings by allowing multi-node connections beyond simple pairwise graph edges. In light of this, we propose a deep spatio-temporal hypergraph convolutional neural network for soft sensing (ST-HCSS). In particular, our proposed framework is able to construct and leverage a higher-order graph (hypergraph) to model the complex multi-interactions between sensor nodes in the absence of prior structural knowledge. To capture rich spatio-temporal relationships underlying sensor data, our proposed ST-HCSS incorporates stacked gated temporal and hypergraph convolution layers to effectively aggregate and update hypergraph information across time and nodes. Our results validate the superiority of ST-HCSS compared to existing state-of-the-art soft sensors, and demonstrates that the learned hypergraph feature representations aligns well with the sensor data correlations. The code is available at https://github.com/htew0001/ST-HCSS.git
PK-YOLO: Pretrained Knowledge Guided YOLO for Brain Tumor Detection in Multiplanar MRI Slices
Oct 29, 2024Abstract:Brain tumor detection in multiplane Magnetic Resonance Imaging (MRI) slices is a challenging task due to the various appearances and relationships in the structure of the multiplane images. In this paper, we propose a new You Only Look Once (YOLO)-based detection model that incorporates Pretrained Knowledge (PK), called PK-YOLO, to improve the performance for brain tumor detection in multiplane MRI slices. To our best knowledge, PK-YOLO is the first pretrained knowledge guided YOLO-based object detector. The main components of the new method are a pretrained pure lightweight convolutional neural network-based backbone via sparse masked modeling, a YOLO architecture with the pretrained backbone, and a regression loss function for improving small object detection. The pretrained backbone allows for feature transferability of object queries on individual plane MRI slices into the model encoders, and the learned domain knowledge base can improve in-domain detection. The improved loss function can further boost detection performance on small-size brain tumors in multiplanar two-dimensional MRI slices. Experimental results show that the proposed PK-YOLO achieves competitive performance on the multiplanar MRI brain tumor detection datasets compared to state-of-the-art YOLO-like and DETR-like object detectors. The code is available at https://github.com/mkang315/PK-YOLO.
Variational Potential Flow: A Novel Probabilistic Framework for Energy-Based Generative Modelling
Jul 21, 2024



Abstract:Energy based models (EBMs) are appealing for their generality and simplicity in data likelihood modeling, but have conventionally been difficult to train due to the unstable and time-consuming implicit MCMC sampling during contrastive divergence training. In this paper, we present a novel energy-based generative framework, Variational Potential Flow (VAPO), that entirely dispenses with implicit MCMC sampling and does not rely on complementary latent models or cooperative training. The VAPO framework aims to learn a potential energy function whose gradient (flow) guides the prior samples, so that their density evolution closely follows an approximate data likelihood homotopy. An energy loss function is then formulated to minimize the Kullback-Leibler divergence between density evolution of the flow-driven prior and the data likelihood homotopy. Images can be generated after training the potential energy, by initializing the samples from Gaussian prior and solving the ODE governing the potential flow on a fixed time interval using generic ODE solvers. Experiment results show that the proposed VAPO framework is capable of generating realistic images on various image datasets. In particular, our proposed framework achieves competitive FID scores for unconditional image generation on the CIFAR-10 and CelebA datasets.
A Multimodal Feature Distillation with CNN-Transformer Network for Brain Tumor Segmentation with Incomplete Modalities
Apr 22, 2024



Abstract:Existing brain tumor segmentation methods usually utilize multiple Magnetic Resonance Imaging (MRI) modalities in brain tumor images for segmentation, which can achieve better segmentation performance. However, in clinical applications, some modalities are missing due to resource constraints, leading to severe degradation in the performance of methods applying complete modality segmentation. In this paper, we propose a Multimodal feature distillation with Convolutional Neural Network (CNN)-Transformer hybrid network (MCTSeg) for accurate brain tumor segmentation with missing modalities. We first design a Multimodal Feature Distillation (MFD) module to distill feature-level multimodal knowledge into different unimodality to extract complete modality information. We further develop a Unimodal Feature Enhancement (UFE) module to model the relationship between global and local information semantically. Finally, we build a Cross-Modal Fusion (CMF) module to explicitly align the global correlations among different modalities even when some modalities are missing. Complementary features within and across different modalities are refined via the CNN-Transformer hybrid architectures in both the UFE and CMF modules, where local and global dependencies are both captured. Our ablation study demonstrates the importance of the proposed modules with CNN-Transformer networks and the convolutional blocks in Transformer for improving the performance of brain tumor segmentation with missing modalities. Extensive experiments on the BraTS2018 and BraTS2020 datasets show that the proposed MCTSeg framework outperforms the state-of-the-art methods in missing modalities cases. Our code is available at: https://github.com/mkang315/MCTSeg.
Dynamic MRI reconstruction using low-rank plus sparse decomposition with smoothness regularization
Jan 30, 2024



Abstract:The low-rank plus sparse (L+S) decomposition model has enabled better reconstruction of dynamic magnetic resonance imaging (dMRI) with separation into background (L) and dynamic (S) component. However, use of low-rank prior alone may not fully explain the slow variations or smoothness of the background part at the local scale. In this paper, we propose a smoothness-regularized L+S (SR-L+S) model for dMRI reconstruction from highly undersampled k-t-space data. We exploit joint low-rank and smooth priors on the background component of dMRI to better capture both its global and local temporal correlated structures. Extending the L+S formulation, the low-rank property is encoded by the nuclear norm, while the smoothness by a general \ell_{p}-norm penalty on the local differences of the columns of L. The additional smoothness regularizer can promote piecewise local consistency between neighboring frames. By smoothing out the noise and dynamic activities, it allows accurate recovery of the background part, and subsequently more robust dMRI reconstruction. Extensive experiments on multi-coil cardiac and synthetic data shows that the SR-L+S model outp
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge