Cencheng Shen
University of Delaware
Discovering Communication Pattern Shifts in Large-Scale Networks using Encoder Embedding and Vertex Dynamics
May 03, 2023Abstract:The analysis of large-scale time-series network data, such as social media and email communications, remains a significant challenge for graph analysis methodology. In particular, the scalability of graph analysis is a critical issue hindering further progress in large-scale downstream inference. In this paper, we introduce a novel approach called "temporal encoder embedding" that can efficiently embed large amounts of graph data with linear complexity. We apply this method to an anonymized time-series communication network from a large organization spanning 2019-2020, consisting of over 100 thousand vertices and 80 million edges. Our method embeds the data within 10 seconds on a standard computer and enables the detection of communication pattern shifts for individual vertices, vertex communities, and the overall graph structure. Through supporting theory and synthesis studies, we demonstrate the theoretical soundness of our approach under random graph models and its numerical effectiveness through simulation studies.
Synergistic Graph Fusion via Encoder Embedding
Mar 31, 2023Abstract:In this paper, we introduce a novel approach to multi-graph embedding called graph fusion encoder embedding. The method is designed to work with multiple graphs that share a common vertex set. Under the supervised learning setting, we show that the resulting embedding exhibits a surprising yet highly desirable "synergistic effect": for sufficiently large vertex size, the vertex classification accuracy always benefits from additional graphs. We provide a mathematical proof of this effect under the stochastic block model, and identify the necessary and sufficient condition for asymptotically perfect classification. The simulations and real data experiments confirm the superiority of the proposed method, which consistently outperforms recent benchmark methods in classification.
Graph Encoder Ensemble for Simultaneous Vertex Embedding and Community Detection
Jan 18, 2023Abstract:In this paper we propose a novel and computationally efficient method to simultaneously achieve vertex embedding, community detection, and community size determination. By utilizing a normalized one-hot graph encoder and a new rank-based cluster size measure, the proposed graph encoder ensemble algorithm achieves excellent numerical performance throughout a variety of simulations and real data experiments.
Graph Encoder Embedding
Sep 27, 2021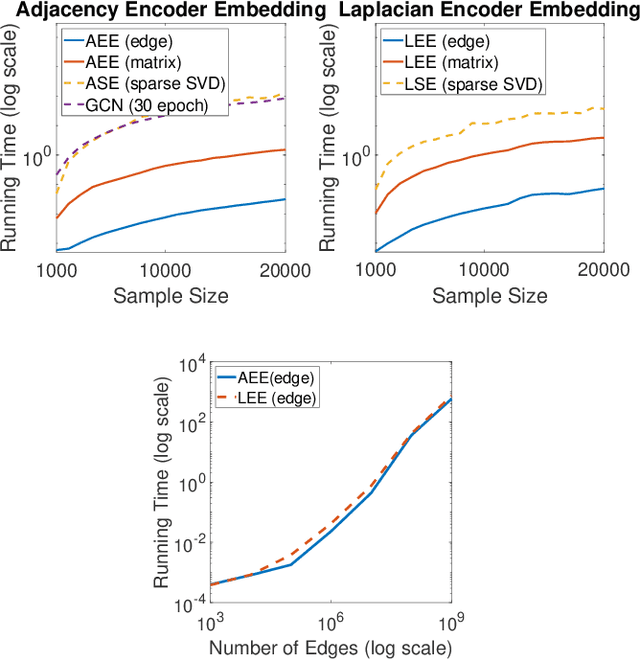
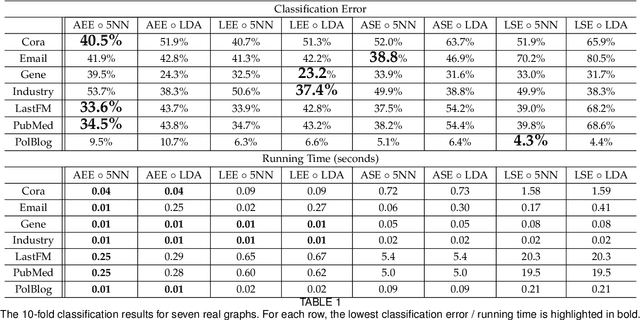
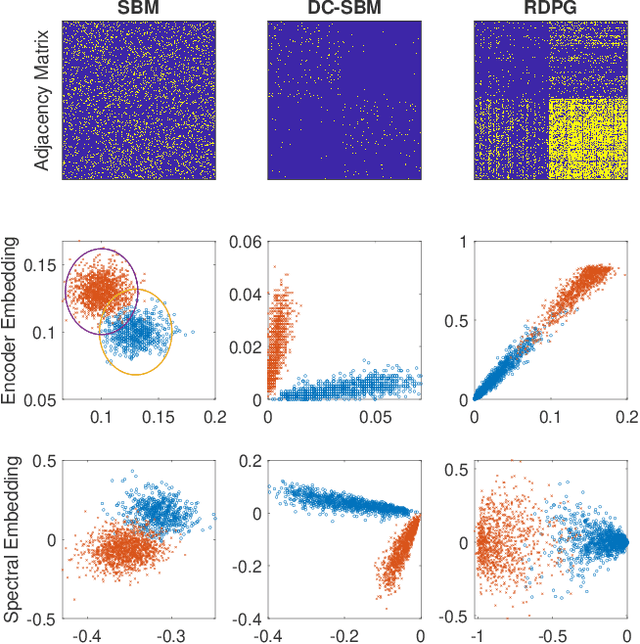
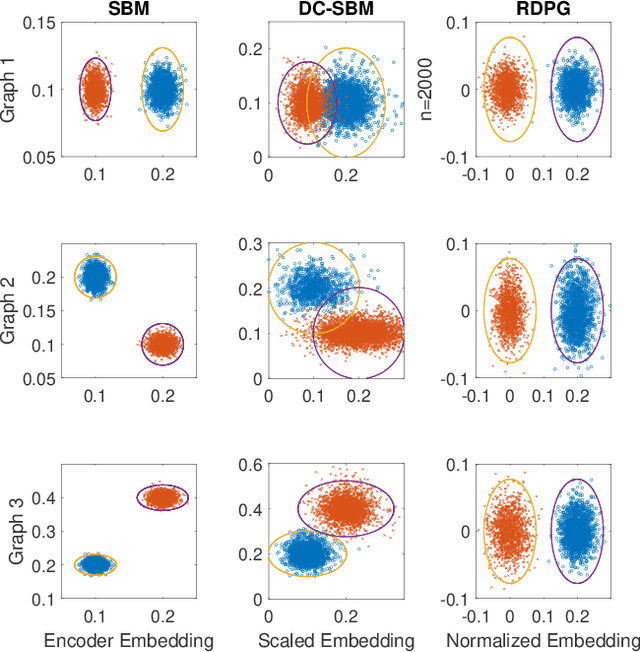
Abstract:In this paper we propose a lightning fast graph embedding method called graph encoder embedding. The proposed method has a linear computational complexity and the capacity to process billions of edges within minutes on standard PC -- an unattainable feat for any existing graph embedding method. The speedup is achieved without sacrificing embedding performance: the encoder embedding performs as good as, and can be viewed as a transformation of the more costly spectral embedding. The encoder embedding is applicable to either adjacency matrix or graph Laplacian, and is theoretically sound, i.e., under stochastic block model or random dot product graph, the graph encoder embedding asymptotically converges to the block probability or latent positions, and is approximately normally distributed. We showcase three important applications: vertex classification, vertex clustering, and graph bootstrap; and the embedding performance is evaluated via a comprehensive set of synthetic and real data. In every case, the graph encoder embedding exhibits unrivalled computational advantages while delivering excellent numerical performance.
A Simple Spectral Failure Mode for Graph Convolutional Networks
Oct 27, 2020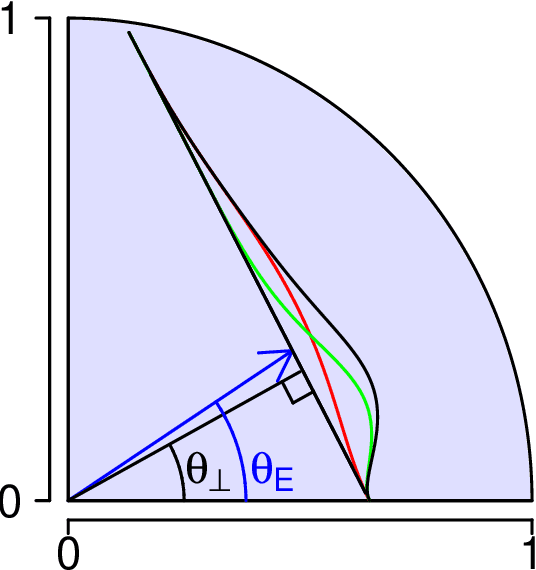
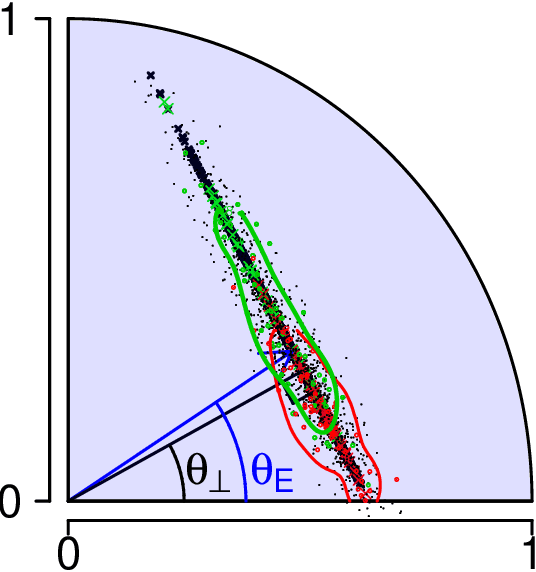
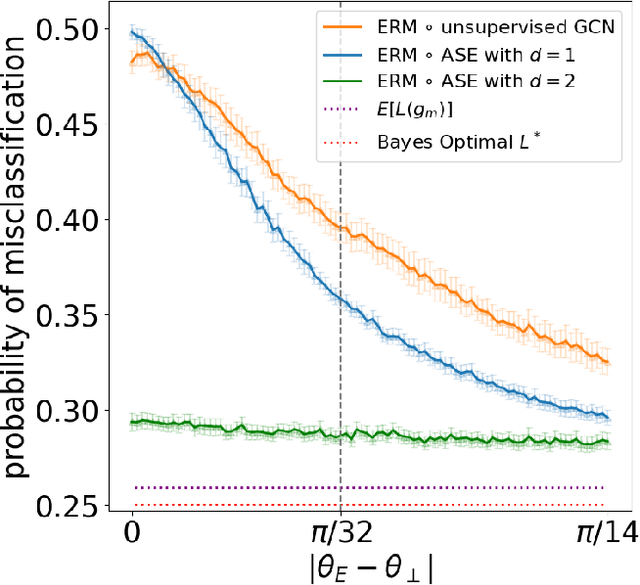
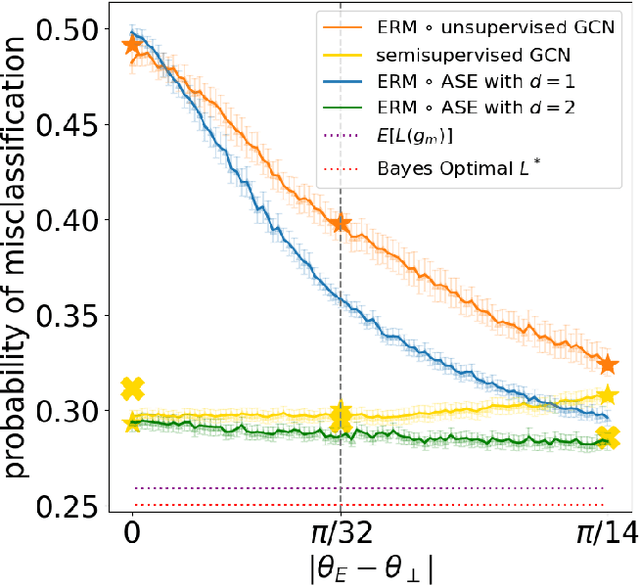
Abstract:We present a simple generative model in which spectral graph embedding for subsequent inference succeeds whereas unsupervised graph convolutional networks (GCN) fail. The geometrical insight is that the GCN is unable to look beyond the first non-informative spectral dimension.
The Chi-Square Test of Distance Correlation
Jan 22, 2020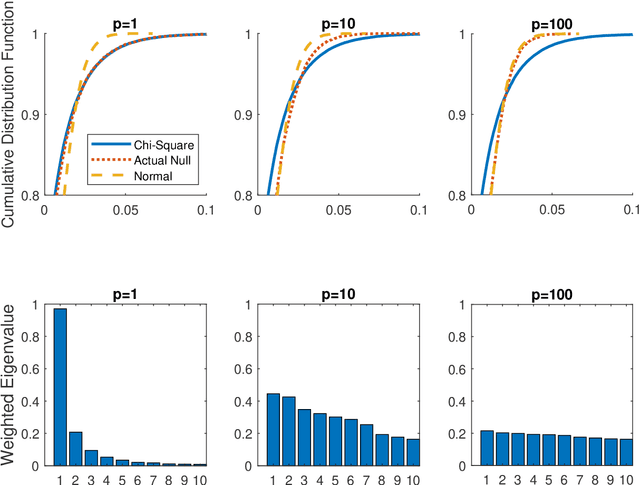
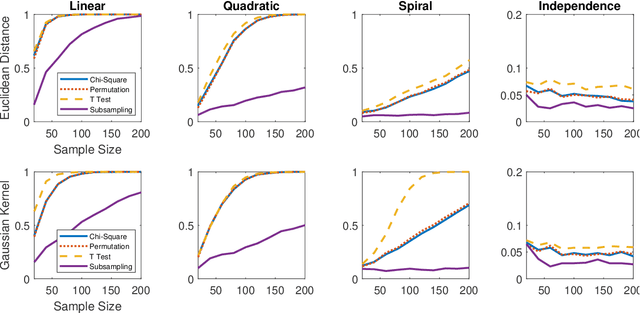
Abstract:Distance correlation has gained much recent attention in the data science community: the sample statistic is straightforward to compute and asymptotically equals zero if and only if independence, making it an ideal choice to test any type of dependency structure given sufficient sample size. One major bottleneck is the testing process: because the null distribution of distance correlation depends on the underlying random variables and metric choice, it typically requires a permutation test to estimate the null and compute the p-value, which is very costly for large amount of data. To overcome the difficulty, we propose a centered chi-square distribution, demonstrate it well-approximates the null distribution of unbiased distance correlation, and prove upper tail dominance and distribution bound between them. The resulting distance correlation chi-square test is a nonparametric test for independence, is valid and universally consistent using any strong negative type metric or characteristic kernel, enjoys a similar finite-sample testing power as the standard permutation test, and is provably the most powerful test of distance correlation among all valid tests with known distribution.
High-Dimensional Independence Testing and Maximum Marginal Correlation
Jan 04, 2020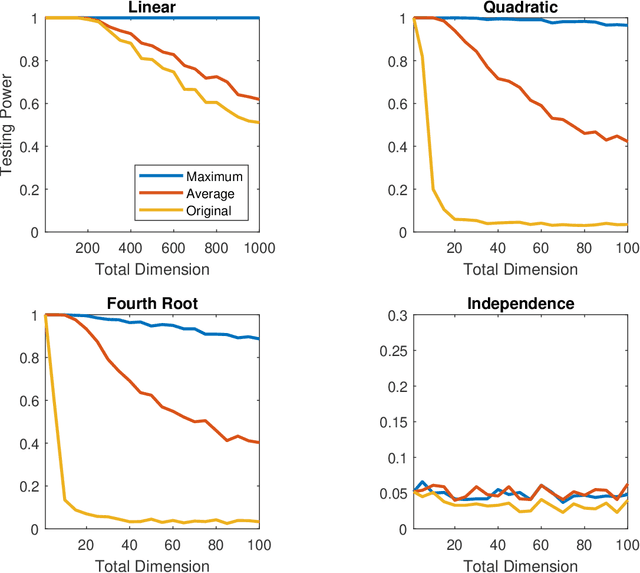
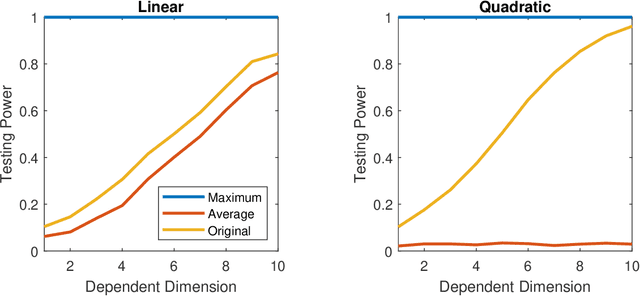
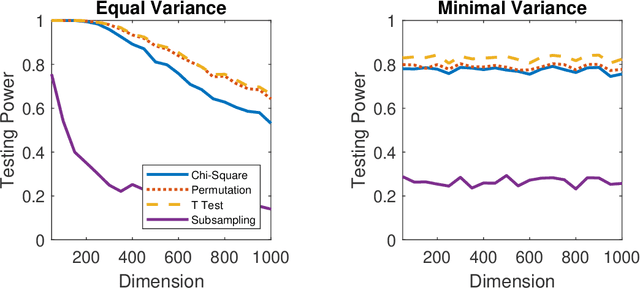
Abstract:A number of universally consistent dependence measures have been recently proposed for testing independence, such as distance correlation, kernel correlation, multiscale graph correlation, etc. They provide a satisfactory solution for dependence testing in low-dimensions, but often exhibit decreasing power for high-dimensional data, a phenomenon that has been recognized but remains mostly unchartered. In this paper, we aim to better understand the high-dimensional testing scenarios and explore a procedure that is robust against increasing dimension. To that end, we propose the maximum marginal correlation method and characterize high-dimensional dependence structures via the notion of dependent dimensions. We prove that the maximum method can be valid and universally consistent for testing high-dimensional dependence under regularity conditions, and demonstrate when and how the maximum method may outperform other methods. The methodology can be implemented by most existing dependence measures, has a superior testing power in a variety of common high-dimensional settings, and is computationally efficient for big data analysis when using the distance correlation chi-square test.
The Exact Equivalence of Independence Testing and Two-Sample Testing
Oct 20, 2019
Abstract:Testing independence and testing equality of distributions are two tightly related statistical hypotheses. Several distance and kernel-based statistics are recently proposed to achieve universally consistent testing for either hypothesis. On the distance side, the distance correlation is proposed for independence testing, and the energy statistic is proposed for two-sample testing. On the kernel side, the Hilbert-Schmidt independence criterion is proposed for independence testing and the maximum mean discrepancy is proposed for two-sample testing. In this paper, we show that two-sample testing are special cases of independence testing via an auxiliary label vector, and prove that distance correlation is exactly equivalent to the energy statistic in terms of the population statistic, the sample statistic, and the testing p-value via permutation test. The equivalence can be further generalized to K-sample testing and extended to the kernel regime. As a consequence, it suffices to always use an independence statistic to test equality of distributions, which enables better interpretability of the test statistic and more efficient testing.
A Consistent Independence Test for Multivariate Time-Series
Aug 18, 2019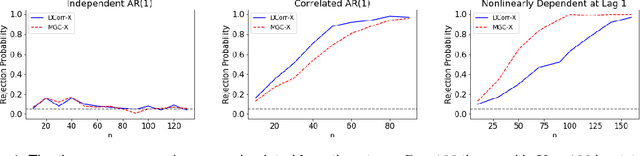
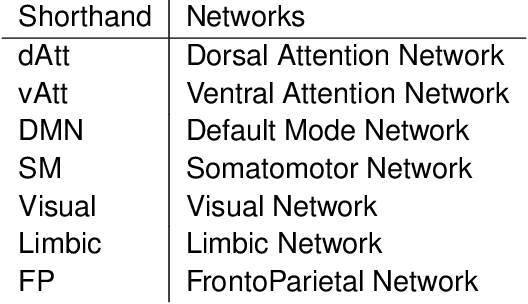
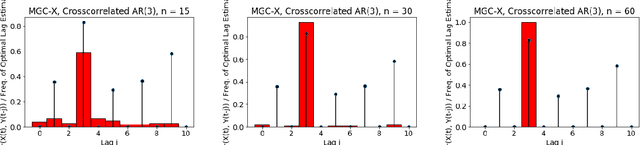
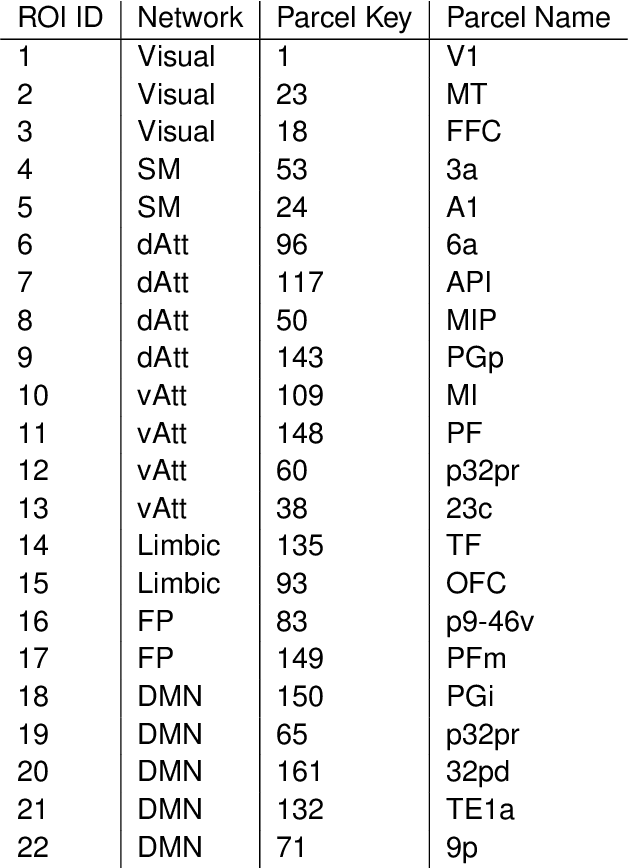
Abstract:A fundamental problem in statistical data analysis is testing whether two phenomena are related. When the phenomena in question are time series, many challenges emerge. The first is defining a dependence measure between time series at the population level, as well as a sample level test statistic. The second is computing or estimating the distribution of this test statistic under the null, as the permutation test procedure is invalid for most time series structures. This work aims to address these challenges by combining distance correlation and multiscale graph correlation (MGC) from independence testing literature and block permutation testing from time series analysis. Two hypothesis tests for testing the independence of time series are proposed. These procedures also characterize whether the dependence relationship between the series is linear or nonlinear, and the time lag at which this dependence is maximized. For strictly stationary auto-regressive moving average (ARMA) processes, the proposed independence tests are proven valid and consistent. Finally, neural connectivity in the brain is analyzed using fMRI data, revealing linear dependence of signals within the visual network and default mode network, and nonlinear relationships in other regions. This work opens up new theoretical and practical directions for many modern time series analysis problems.
mgcpy: A Comprehensive High Dimensional Independence Testing Python Package
Jul 18, 2019
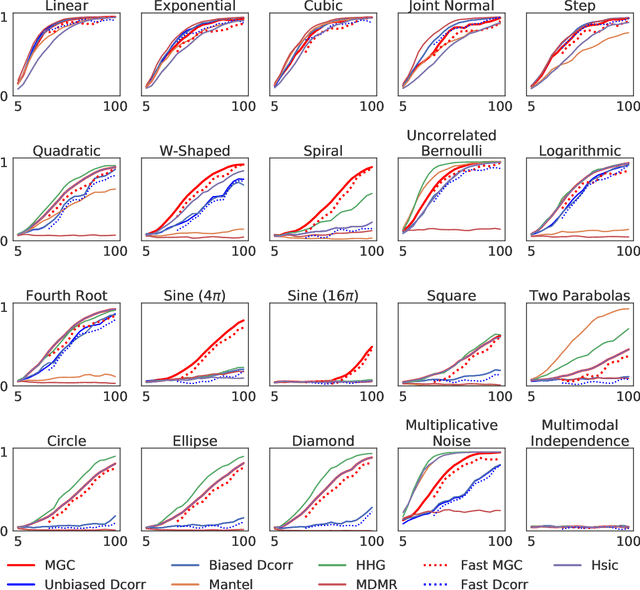
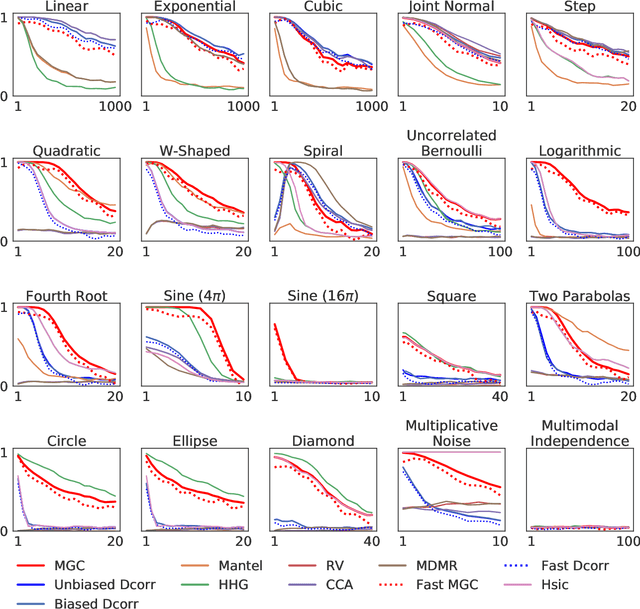
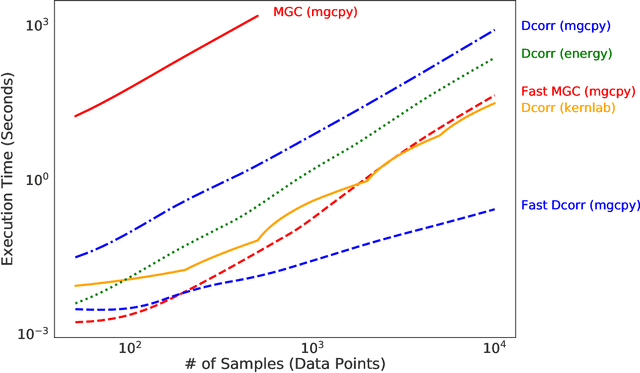
Abstract:With the increase in the amount of data in many fields, a method to consistently and efficiently decipher relationships within high dimensional data sets is important. Because many modern datasets are high-dimensional, univariate independence tests are not applicable. While many multivariate independence tests have R packages available, the interfaces are inconsistent, most are not available in Python. mgcpy is an extensive Python library that includes many state of the art high-dimensional independence testing procedures using a common interface. The package is easy-to-use and is flexible enough to enable future extensions. This manuscript provides details for each of the tests as well as extensive power and run-time benchmarks on a suite of high-dimensional simulations previously used in different publications. The appendix includes demonstrations of how the user can interact with the package, as well as links and documentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge