Brian Mac Namee
Extracting Pasture Phenotype and Biomass Percentages using Weakly Supervised Multi-target Deep Learning on a Small Dataset
Jan 08, 2021
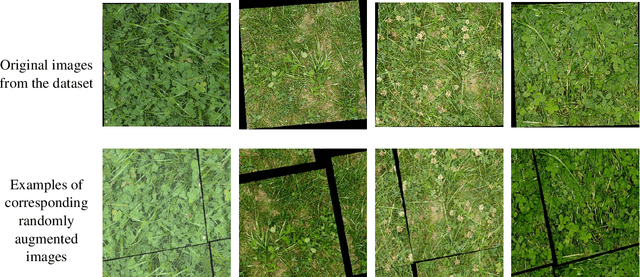

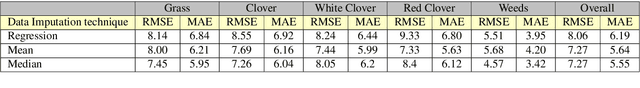
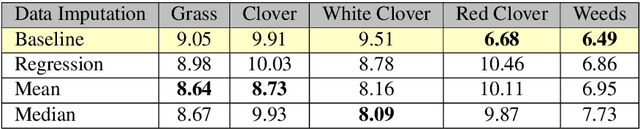
Abstract:The dairy industry uses clover and grass as fodder for cows. Accurate estimation of grass and clover biomass yield enables smart decisions in optimizing fertilization and seeding density, resulting in increased productivity and positive environmental impact. Grass and clover are usually planted together, since clover is a nitrogen-fixing plant that brings nutrients to the soil. Adjusting the right percentages of clover and grass in a field reduces the need for external fertilization. Existing approaches for estimating the grass-clover composition of a field are expensive and time consuming - random samples of the pasture are clipped and then the components are physically separated to weigh and calculate percentages of dry grass, clover and weeds in each sample. There is growing interest in developing novel deep learning based approaches to non-destructively extract pasture phenotype indicators and biomass yield predictions of different plant species from agricultural imagery collected from the field. Providing these indicators and predictions from images alone remains a significant challenge. Heavy occlusions in the dense mixture of grass, clover and weeds make it difficult to estimate each component accurately. Moreover, although supervised deep learning models perform well with large datasets, it is tedious to acquire large and diverse collections of field images with precise ground truth for different biomass yields. In this paper, we demonstrate that applying data augmentation and transfer learning is effective in predicting multi-target biomass percentages of different plant species, even with a small training dataset. The scheme proposed in this paper used a training set of only 261 images and provided predictions of biomass percentages of grass, clover, white clover, red clover, and weeds with mean absolute error of 6.77%, 6.92%, 6.21%, 6.89%, and 4.80% respectively.
Predicting Illness for a Sustainable Dairy Agriculture: Predicting and Explaining the Onset of Mastitis in Dairy Cows
Jan 07, 2021
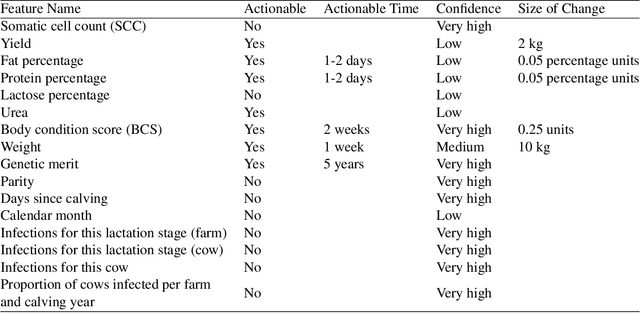
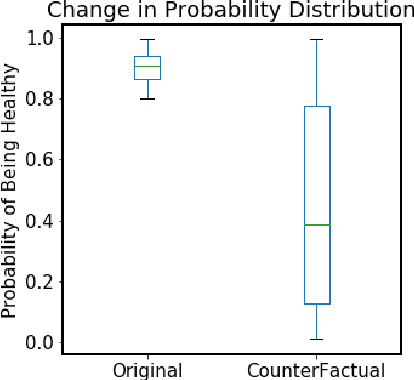

Abstract:Mastitis is a billion dollar health problem for the modern dairy industry, with implications for antibiotic resistance. The use of AI techniques to identify the early onset of this disease, thus has significant implications for the sustainability of this agricultural sector. Current approaches to treating mastitis involve antibiotics and this practice is coming under ever increasing scrutiny. Using machine learning models to identify cows at risk of developing mastitis and applying targeted treatment regimes to only those animals promotes a more sustainable approach. Incorrect predictions from such models, however, can lead to monetary losses, unnecessary use of antibiotics, and even the premature death of animals, so it is important to generate compelling explanations for predictions to build trust with users and to better support their decision making. In this paper we demonstrate a system developed to predict mastitis infections in cows and provide explanations of these predictions using counterfactuals. We demonstrate the system and describe the engagement with farmers undertaken to build it.
Can We Detect Mastitis earlier than Farmers?
Nov 05, 2020
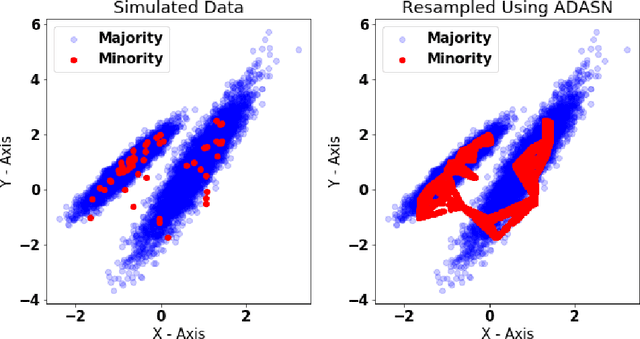
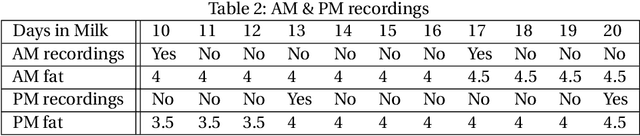
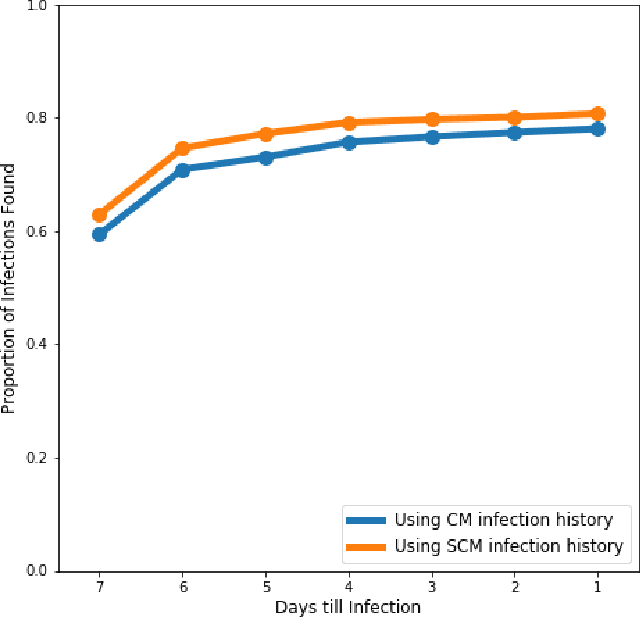
Abstract:The aim of this study was to build a modelling framework that would allow us to be able to detect mastitis infections before they would normally be found by farmers through the introduction of machine learning techniques. In the making of this we created two different modelling framework's, one that works on the premise of detecting Sub Clinical mastitis infections at one Somatic Cell Count recording in advance called SMA and the other tries to detect both Sub Clinical mastitis infections aswell as Clinical mastitis infections at any time the cow is milked called AMA. We also introduce the idea of two different feature sets for our study, these represent different characteristics that should be taken into account when detecting infections, these were the idea of a cow differing to a farm mean and also trends in the lactation. We reported that the results for SMA are better than those created by AMA for Sub Clinical infections yet it has the significant disadvantage of only being able to classify Sub Clinical infections due to how we recorded Sub Clinical infections as being any time a Somatic Cell Count measurement went above a certain threshold where as CM could appear at any stage of lactation. Thus in some cases the lower accuracy values for AMA might in fact be more beneficial to farmers.
Ramifications of Approximate Posterior Inference for Bayesian Deep Learning in Adversarial and Out-of-Distribution Settings
Oct 03, 2020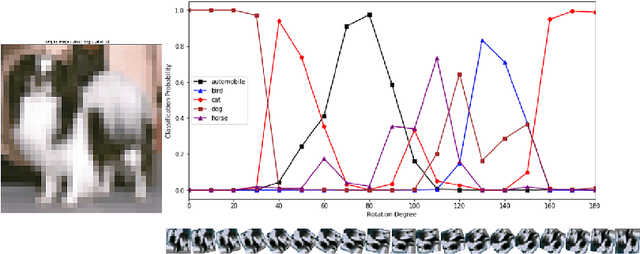
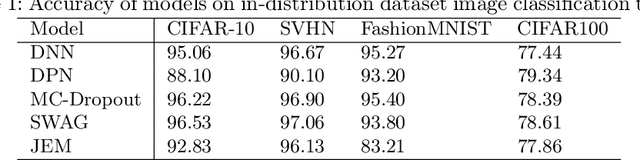
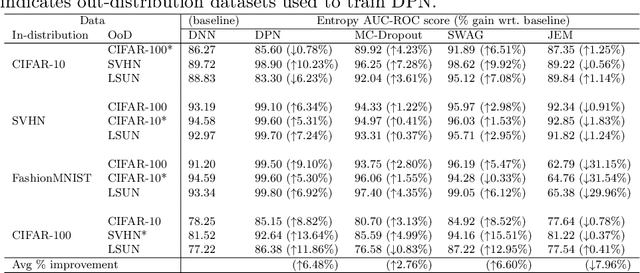
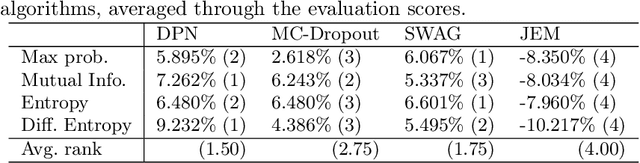
Abstract:Deep neural networks have been successful in diverse discriminative classification tasks, although, they are poorly calibrated often assigning high probability to misclassified predictions. Potential consequences could lead to trustworthiness and accountability of the models when deployed in real applications, where predictions are evaluated based on their confidence scores. Existing solutions suggest the benefits attained by combining deep neural networks and Bayesian inference to quantify uncertainty over the models' predictions for ambiguous datapoints. In this work we propose to validate and test the efficacy of likelihood based models in the task of out of distribution detection (OoD). Across different datasets and metrics we show that Bayesian deep learning models on certain occasions marginally outperform conventional neural networks and in the event of minimal overlap between in/out distribution classes, even the best models exhibit a reduction in AUC scores in detecting OoD data. Preliminary investigations indicate the potential inherent role of bias due to choices of initialisation, architecture or activation functions. We hypothesise that the sensitivity of neural networks to unseen inputs could be a multi-factor phenomenon arising from the different architectural design choices often amplified by the curse of dimensionality. Furthermore, we perform a study to find the effect of the adversarial noise resistance methods on in and out-of-distribution performance, as well as, also investigate adversarial noise robustness of Bayesian deep learners.
Deep Context-Aware Novelty Detection
Jun 01, 2020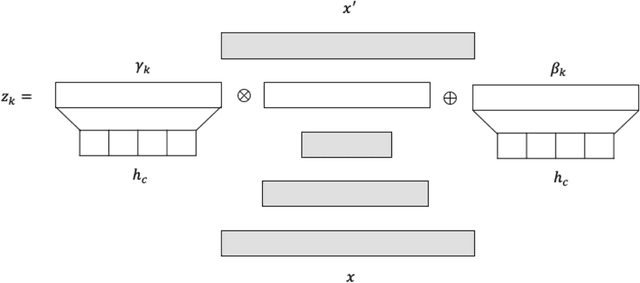
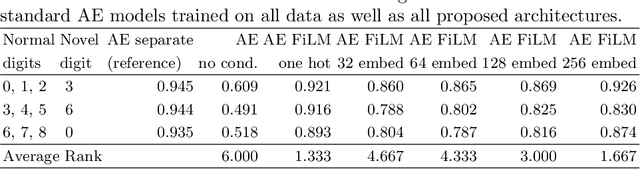
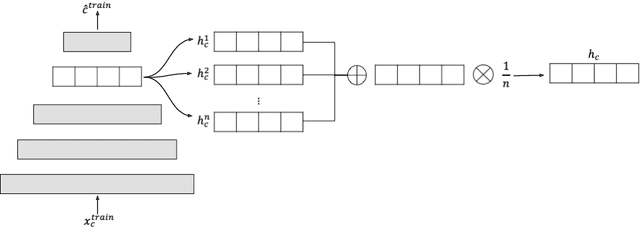
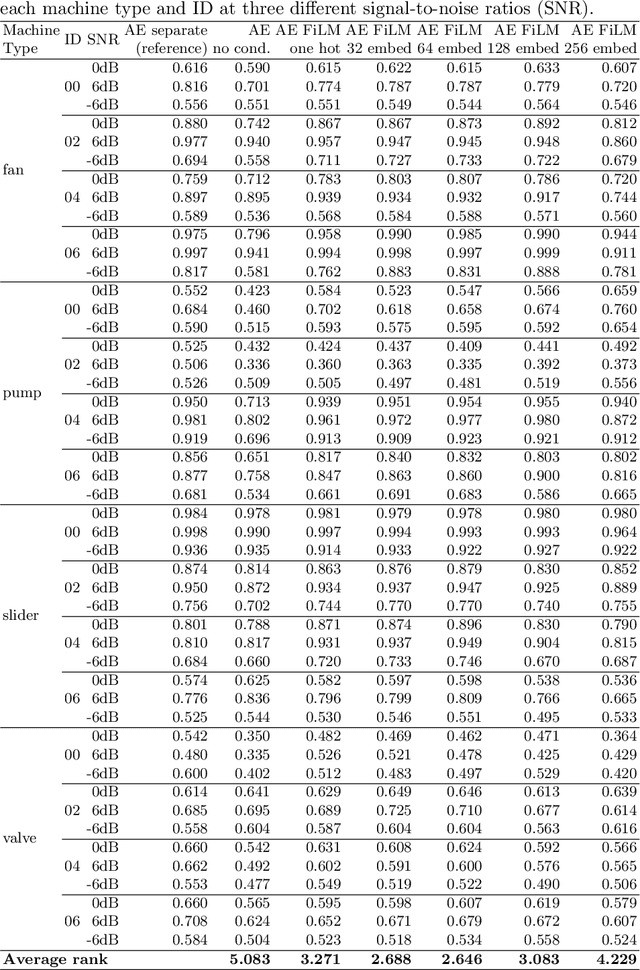
Abstract:A common assumption of novelty detection is that the distribution of both "normal" and "novel" data are static. However, this is often not the case in scenarios where data evolves over time, or when the definition of normal and novel depends on contextual information, leading to changes in these distributions. This can lead to significant difficulties when attempting to train a model on datasets where the distribution of normal data in one scenario is similar to that of novel data in another scenario. In this paper we propose a context-aware approach to novelty detection for deep autoencoders. We create a semi-supervised network architecture which utilises auxiliary labels in order to reveal contextual information and allows the model to adapt to a variety of normal and novel scenarios. We evaluate our approach on both synthetic image data and real world audio data displaying these characteristics.
On the Validity of Bayesian Neural Networks for Uncertainty Estimation
Dec 29, 2019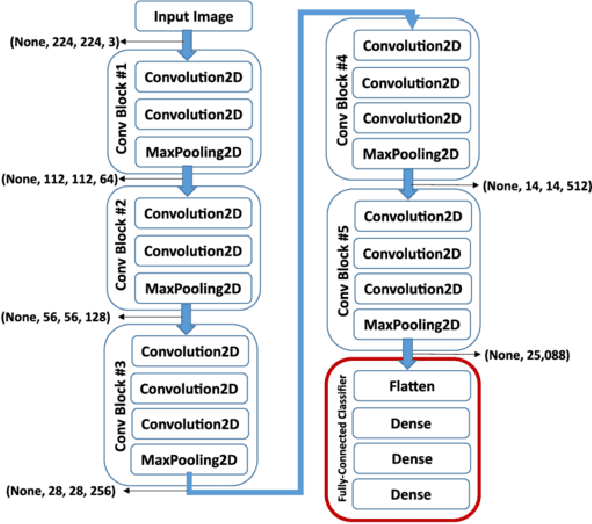
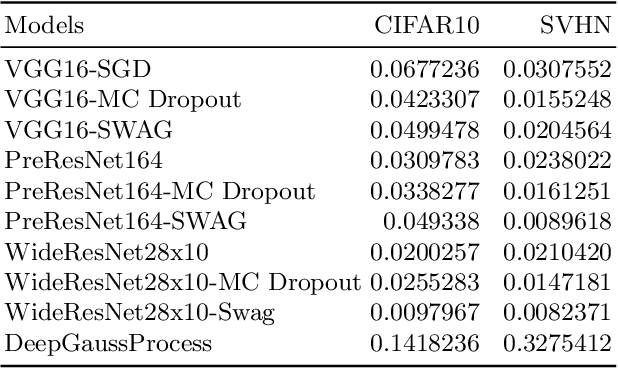
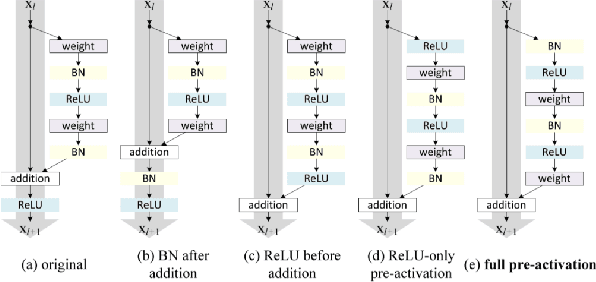
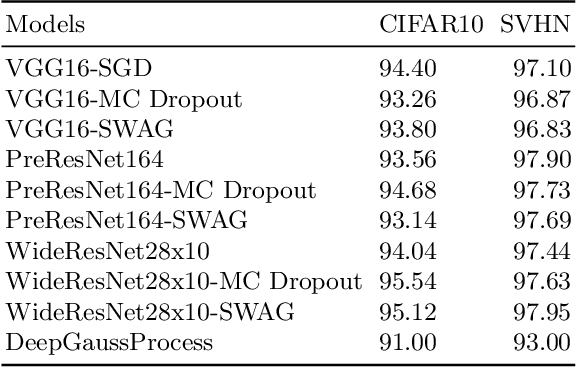
Abstract:Deep neural networks (DNN) are versatile parametric models utilised successfully in a diverse number of tasks and domains. However, they have limitations---particularly from their lack of robustness and over-sensitivity to out of distribution samples. Bayesian Neural Networks, due to their formulation under the Bayesian framework, provide a principled approach to building neural networks that address these limitations. This paper describes a study that empirically evaluates and compares Bayesian Neural Networks to their equivalent point estimate Deep Neural Networks to quantify the predictive uncertainty induced by their parameters, as well as their performance in view of this uncertainty. In this study, we evaluated and compared three point estimate deep neural networks against comparable Bayesian neural network alternatives using two well-known benchmark image classification datasets (CIFAR-10 and SVHN).
Real-time Bidding campaigns optimization using attribute selection
Oct 29, 2019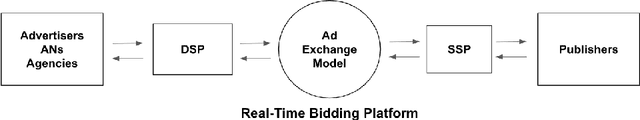
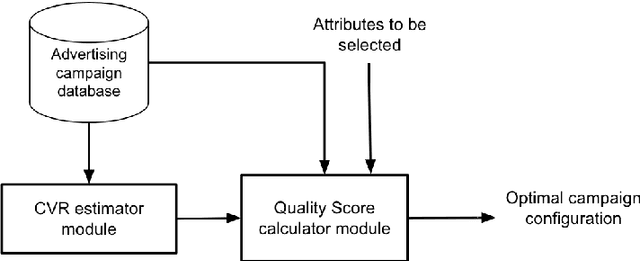
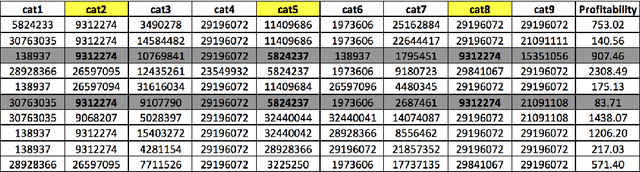
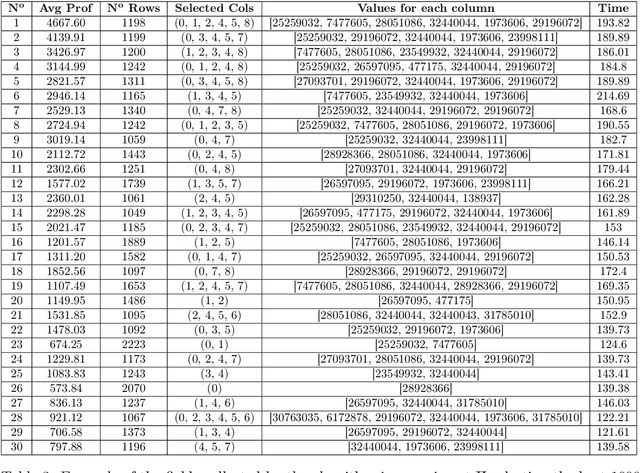
Abstract:Real-Time Bidding is nowadays one of the most promising systems in the online advertising ecosystem. In the presented study, the performance of RTB campaigns is improved by optimising the parameters of the users' profiles and the publishers' websites. Most studies about optimising RTB campaigns are focused on the bidding strategy. In contrast, the objective of our research consists of optimising RTB campaigns by finding out configurations that maximise both the number of impressions and their average profitability. The experiments demonstrate that, when the number of required visits by advertisers is low, it is easy to find configurations with high average profitability, but as the required number of visits increases, the average profitability tends to go down. Additionally, configuration optimisation has been combined with other interesting strategies to increase, even more, the campaigns' profitability. Along with parameter configuration the study considers the following complementary strategies to increase profitability: i) selecting multiple configurations with a small number of visits instead of a unique configuration with a large number, ii) discarding visits according to the thresholds of cost and profitability, iii) analysing a reduced space of the dataset and extrapolating the solution, and iv) increasing the search space by including solutions below the required number of visits. The developed campaign optimisation methodology could be offered by RTB platforms to advertisers to make their campaigns more profitable.
Investigating the Effectiveness of Representations Based on Word-Embeddings in Active Learning for Labelling Text Datasets
Oct 10, 2019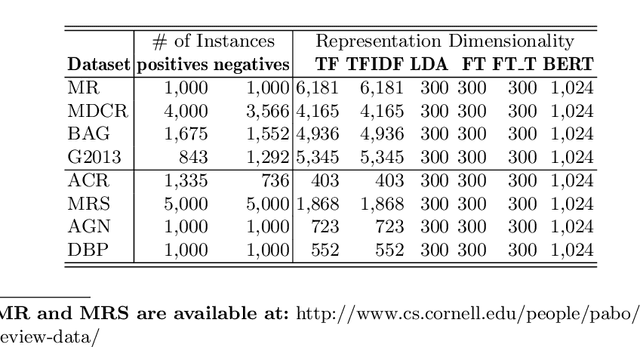
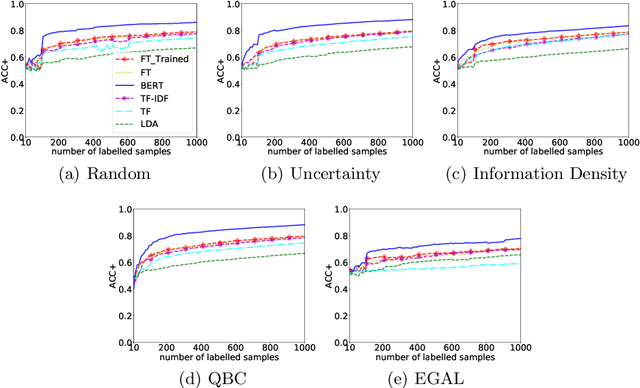
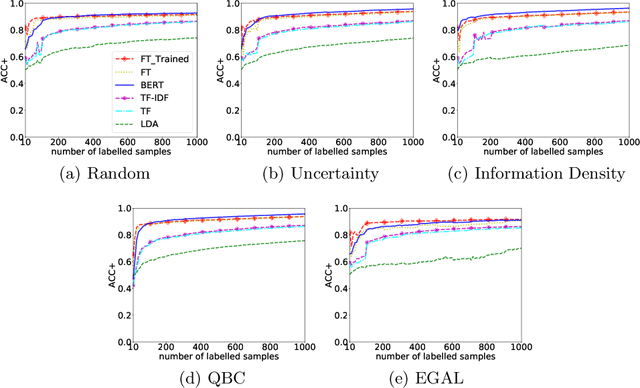
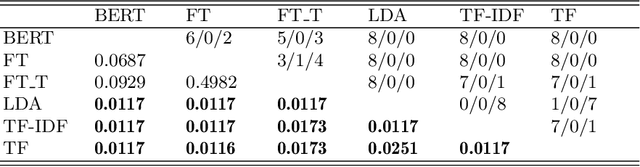
Abstract:Manually labelling large collections of text data is a time-consuming, expensive, and laborious task, but one that is necessary to support machine learning based on text datasets. Active learning has been shown to be an effective way to alleviate some of the effort required in utilising large collections of unlabelled data for machine learning tasks without needing to fully label them. The representation mechanism used to represent text documents when performing active learning, however, has a significant influence on how effective the process will be. While simple vector representations such as bag of words have been shown to be an effective way to represent documents during active learning, the emergence of representation mechanisms based on the word embeddings prevalent in neural network research (e.g. word2vec and transformer-based models like BERT) offer a promising, and as yet not fully explored, alternative. This paper describes a large-scale evaluation of the effectiveness of different text representation mechanisms for active learning across 8 datasets from varied domains. This evaluation shows that using representations based on modern word embeddings---especially BERT---, which have not yet been widely used in active learning, achieves a significant improvement over more commonly used vector-based methods like bag of words.
ZeLiC and ZeChipC: Time Series Interpolation Methods for Lebesgue or Event-based Sampling
Jun 06, 2019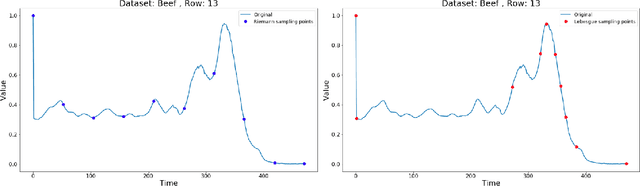
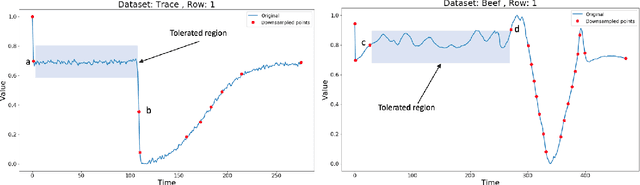
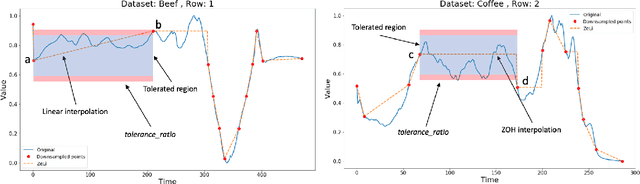
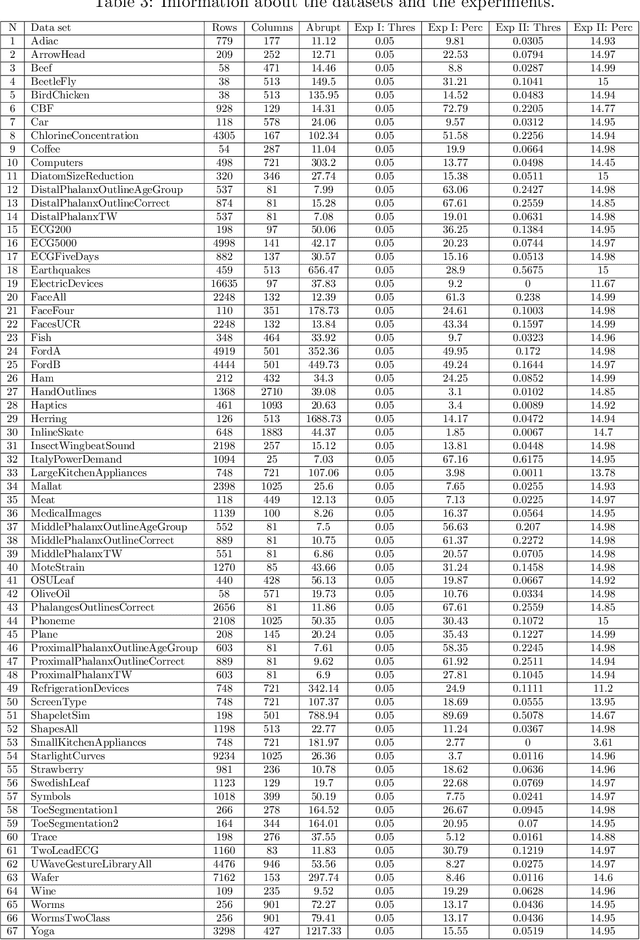
Abstract:Lebesgue sampling is based on collecting information depending on the values of the signal. Although the interpolation methods for periodic sampling have been a topic of research for a long time, there is a lack of study in methods capable of taking advantage of the Lebesgue sampling characteristics to reconstruct time series more accurately. Indeed, Lebesgue sampling contains additional information about the shape of the signal in-between two sampled points. Using this information would allow us to generate an interpolated signal closer to the original one. That is to say, the average distance between the interpolated signal and the original signal will be smaller than a signal interpolated with other interpolation methods. In this paper, we propose two novel time series interpolation methods specifically designed for Lebesgue sampling called ZeLiC and ZeChipC. ZeLiC is an algorithm that combines both Zero-order hold interpolation and Linear interpolation to reconstruct time series. ZeChipC is a similar idea, it is a combination of Zero-order hold and PCHIP interpolation. Zero-order hold interpolation is favourable for interpolating abrupt changes while Linear and PCHIP interpolation are more suitable for smooth transitions. In order to apply one method or the other, we have introduced a new concept called tolerated region. ZeLiC and ZeChipC include a new functionality to adapt the reconstructed signal to concave/convex regions. The proposed methods have been compared with the state-of-the-art interpolation methods using Lebesgue sampling and have offered higher average performance. Additionally, we have compared the performance of the methods using both Riemann and Lebesgue sampling using an approximate number of sampled points. The performance of the combination "Lebesgue sampling with ZeChipC interpolation method" is clearly much better than any other combination.
KFHE-HOMER: Kalman Filter-based Heuristic Ensemble of HOMER for Multi-Label Classification
Apr 23, 2019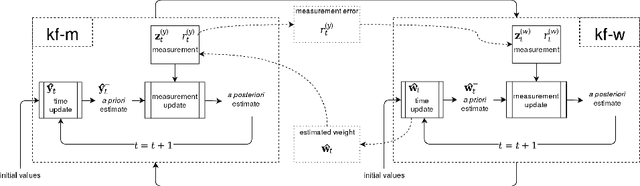
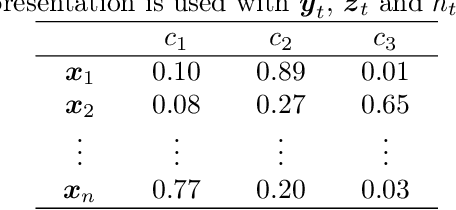
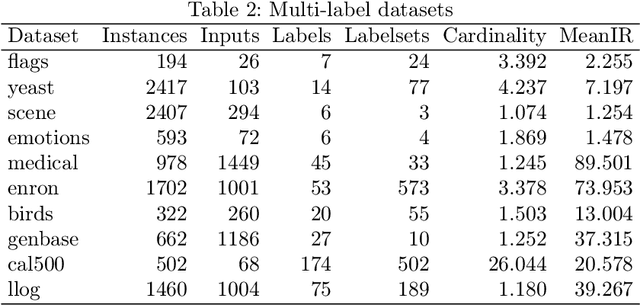
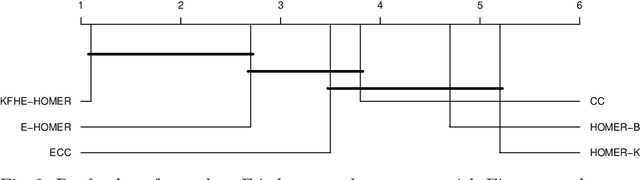
Abstract:Multi-label classification allows a datapoint to be labelled with more than one class at the same time. Ensemble methods generally perform much better than single classifiers. Except bagging style ensembles like ECC, RAkEL, in multi-label classification, other ensemble methods have not been explored much. KFHE (Kalman Filter-based Heuristic Ensemble), is a recent ensemble method which uses the Kalman filter to combine several models. KFHE views the final ensemble to be learned as a state to be estimated which it estimates using multiple noisy "measurements". These "measurements" are essentially component classifiers trained under different settings. This work extends KFHE to multi-label domain by proposing KFHE-HOMER which enhances the performance of HOMER using the KFHE framework. KFHE-HOMER sequentially trains multiple HOMER classifiers using weighted training datapoints and random hyperparameters. These models are considered as measurements and their related error as the uncertainty of the measurements. Then the Kalman filter framework is used to combine these measurements to get a more accurate estimate. The method was tested on 10 multi-label datasets and compared with other multi-label classification algorithms. Results show that KFHE-HOMER performs consistently better than similar multi-label ensemble methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge