Asim Smailagic
Attention Filtering for Multi-person Spatiotemporal Action Detection on Deep Two-Stream CNN Architectures
Jul 21, 2019
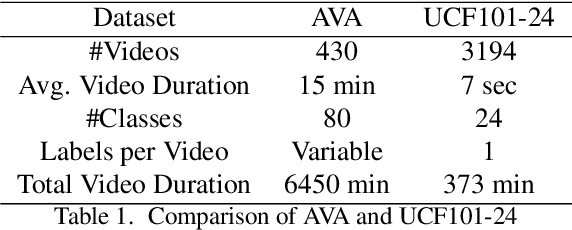


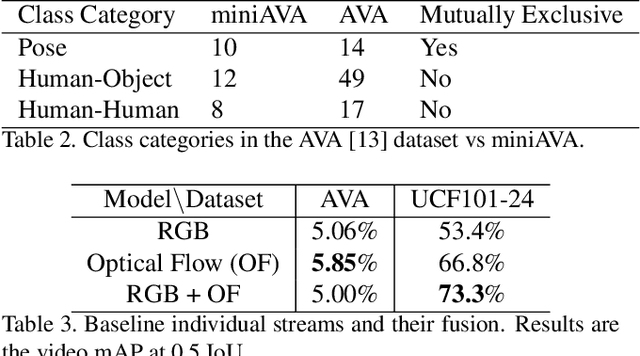
Abstract:Action detection and recognition tasks have been the target of much focus in the computer vision community due to their many applications, namely, security, robotics and recommendation systems. Recently, datasets like AVA, provide multi-person, multi-label, spatiotemporal action detection and recognition challenges. Being unable to discern which portions of the input to use for classification is a limitation of two-stream CNN approaches, once the vision task involves several people with several labels. We address this limitation and improve the state-of-the-art performance of two-stream CNNs. In this paper we present four contributions: our fovea attention filtering that highlights targets for classification without discarding background; a generalized binary loss function designed for the AVA dataset; miniAVA, a partition of AVA that maintains temporal continuity and class distribution with only one tenth of the dataset size; and ablation studies on alternative attention filters. Our method, using fovea attention filtering and our generalized binary loss, achieves a relative video mAP improvement of 20% over the two-stream baseline in AVA, and is competitive with the state-of-the-art in the UCF101-24. We also show a relative video mAP improvement of 12.6% when using our generalized binary loss over the standard sum-of-sigmoids.
Weighted Multisource Tradaboost
Mar 26, 2019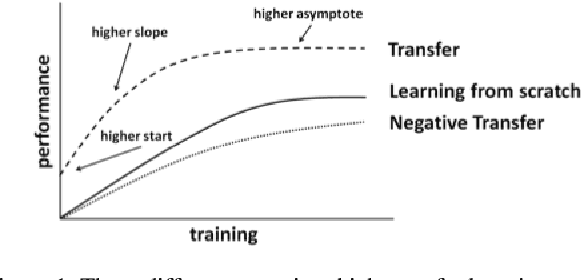
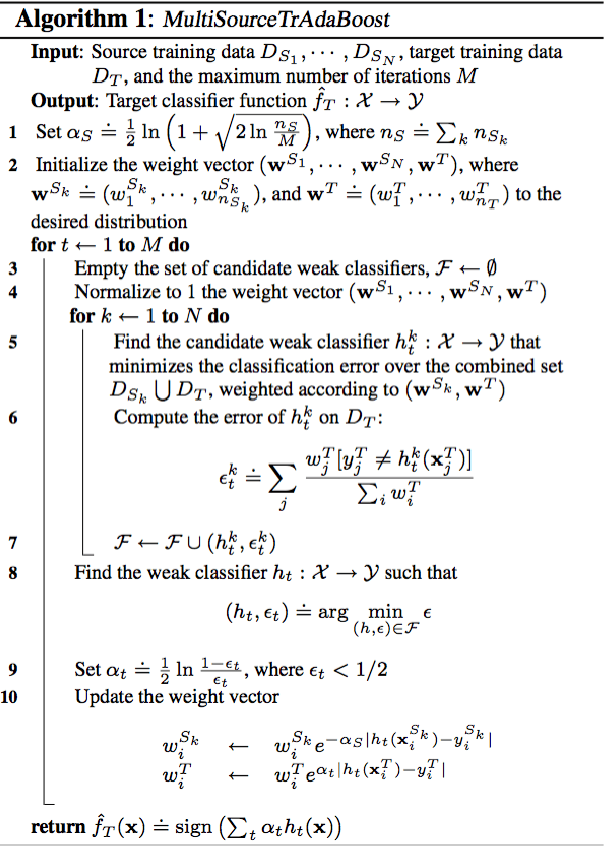

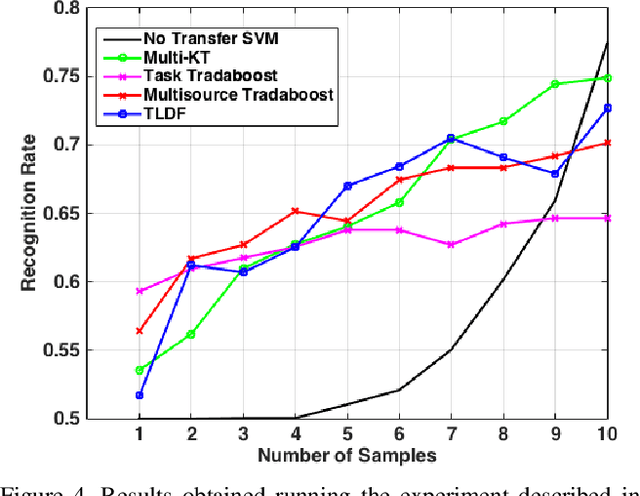
Abstract:In this paper we propose an improved method for transfer learning that takes into account the balance between target and source data. This method builds on the state-of-the-art Multisource Tradaboost, but weighs the importance of each datapoint taking into account the amount of target and source data available. A comparative study is then presented exposing the performance of four transfer learning methods as well as the proposed Weighted Multisource Tradaboost. The experimental results show that the proposed method is able to outperform the base method as the number of target samples increase. These results are promising in the sense that source-target ratio weighing may be a path to improve current methods of transfer learning. However, against the asymptotic conjecture, all transfer learning methods tested in this work get outperformed by a no-transfer SVM for large number on target samples.
MedAL: Deep Active Learning Sampling Method for Medical Image Analysis
Sep 28, 2018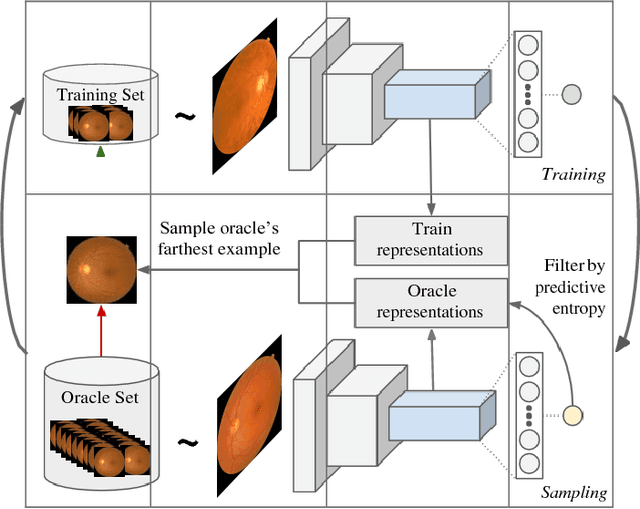
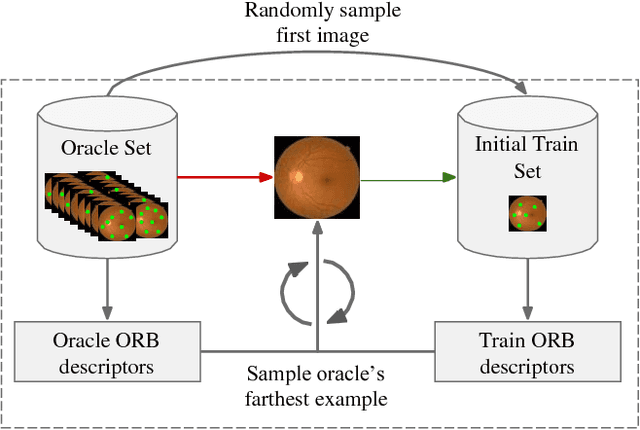
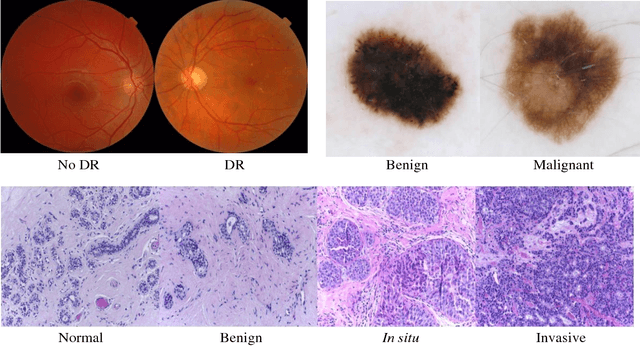
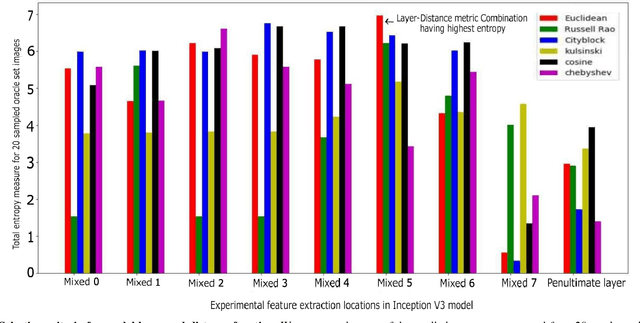
Abstract:Deep learning models have been successfully used in medical image analysis problems but they require a large amount of labeled images to obtain good performance.Deep learning models have been successfully used in medical image analysis problems but they require a large amount of labeled images to obtain good performance. However, such large labeled datasets are costly to acquire. Active learning techniques can be used to minimize the number of required training labels while maximizing the model's performance.In this work, we propose a novel sampling method that queries the unlabeled examples that maximize the average distance to all training set examples in a learned feature space. We then extend our sampling method to define a better initial training set, without the need for a trained model, by using ORB feature descriptors. We validate MedAL on 3 medical image datasets and show that our method is robust to different dataset properties. MedAL is also efficient, achieving 80% accuracy on the task of Diabetic Retinopathy detection using only 425 labeled images, corresponding to a 32% reduction in the number of required labeled examples compared to the standard uncertainty sampling technique, and a 40% reduction compared to random sampling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge