Anthony Costa
Fold-CP: A Context Parallelism Framework for Biomolecular Modeling
Mar 16, 2026Abstract:Understanding cellular machinery requires atomic-scale reconstruction of large biomolecular assemblies. However, predicting the structures of these systems has been constrained by hardware memory requirements of models like AlphaFold 3, imposing a practical ceiling of a few thousand residues that can be processed on a single GPU. Here we present NVIDIA BioNeMo Fold-CP, a context parallelism framework that overcomes this barrier by distributing the inference and training pipelines of co-folding models across multiple GPUs. We use the Boltz models as open source reference architectures and implement custom multidimensional primitives that efficiently parallelize both the dense triangular updates and the irregular, data-dependent pattern of window-batched local attention. Our approach achieves efficient memory scaling; for an N-token input distributed across P GPUs, per-device memory scales as $O(N^2/P)$, enabling the structure prediction of assemblies exceeding 30,000 residues on 64 NVIDIA B300 GPUs. We demonstrate the scientific utility of this approach through successful developer use cases: Fold-CP enabled the scoring of over 90% of Comprehensive Resource of Mammalian protein complexes (CORUM) database, as well as folding of disease-relevant PI4KA lipid kinase complex bound to an intrinsically disordered region without cropping. By providing a scalable pathway for modeling massive systems with full global context, Fold-CP represents a significant step toward the realization of a virtual cell.
nach0: Multimodal Natural and Chemical Languages Foundation Model
Nov 21, 2023



Abstract:Large Language Models (LLMs) have substantially driven scientific progress in various domains, and many papers have demonstrated their ability to tackle complex problems with creative solutions. Our paper introduces a new foundation model, nach0, capable of solving various chemical and biological tasks: biomedical question answering, named entity recognition, molecular generation, molecular synthesis, attributes prediction, and others. nach0 is a multi-domain and multi-task encoder-decoder LLM pre-trained on unlabeled text from scientific literature, patents, and molecule strings to incorporate a range of chemical and linguistic knowledge. We employed instruction tuning, where specific task-related instructions are utilized to fine-tune nach0 for the final set of tasks. To train nach0 effectively, we leverage the NeMo framework, enabling efficient parallel optimization of both base and large model versions. Extensive experiments demonstrate that our model outperforms state-of-the-art baselines on single-domain and cross-domain tasks. Furthermore, it can generate high-quality outputs in molecular and textual formats, showcasing its effectiveness in multi-domain setups.
The Utility of General Domain Transfer Learning for Medical Language Tasks
Feb 16, 2020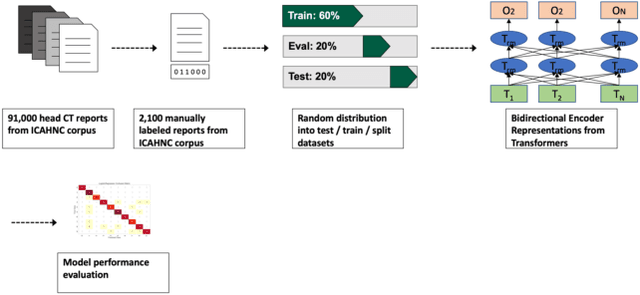
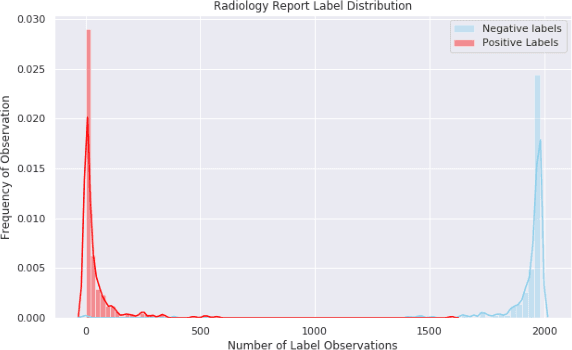
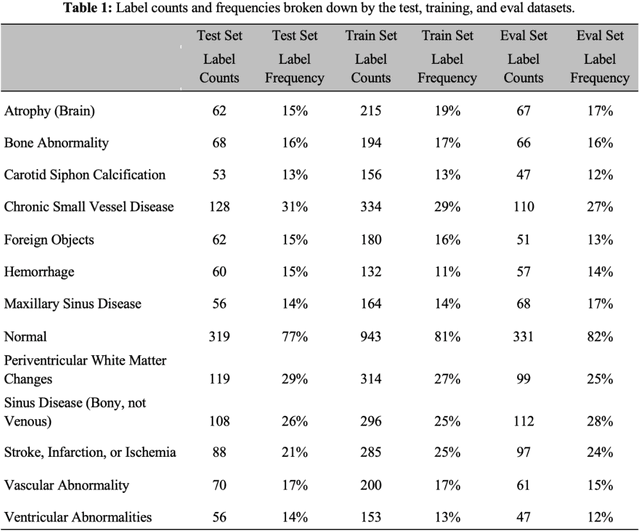
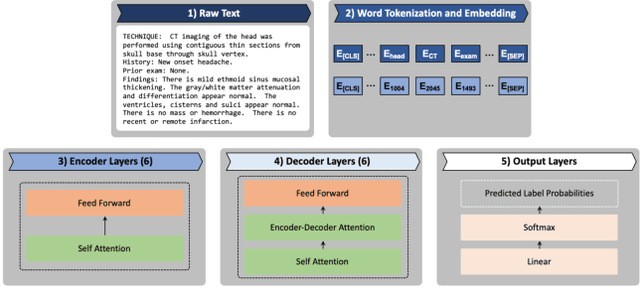
Abstract:The purpose of this study is to analyze the efficacy of transfer learning techniques and transformer-based models as applied to medical natural language processing (NLP) tasks, specifically radiological text classification. We used 1,977 labeled head CT reports, from a corpus of 96,303 total reports, to evaluate the efficacy of pretraining using general domain corpora and a combined general and medical domain corpus with a bidirectional representations from transformers (BERT) model for the purpose of radiological text classification. Model performance was benchmarked to a logistic regression using bag-of-words vectorization and a long short-term memory (LSTM) multi-label multi-class classification model, and compared to the published literature in medical text classification. The BERT models using either set of pretrained checkpoints outperformed the logistic regression model, achieving sample-weighted average F1-scores of 0.87 and 0.87 for the general domain model and the combined general and biomedical-domain model. General text transfer learning may be a viable technique to generate state-of-the-art results within medical NLP tasks on radiological corpora, outperforming other deep models such as LSTMs. The efficacy of pretraining and transformer-based models could serve to facilitate the creation of groundbreaking NLP models in the uniquely challenging data environment of medical text.
Wide and deep volumetric residual networks for volumetric image classification
Sep 18, 2017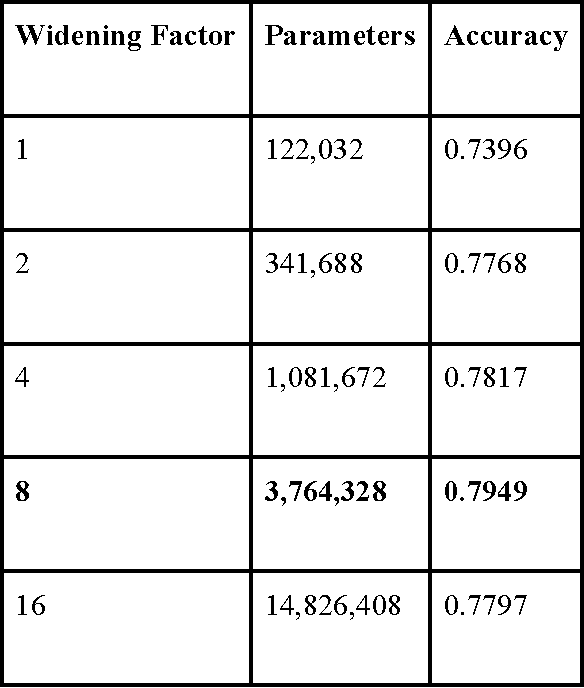
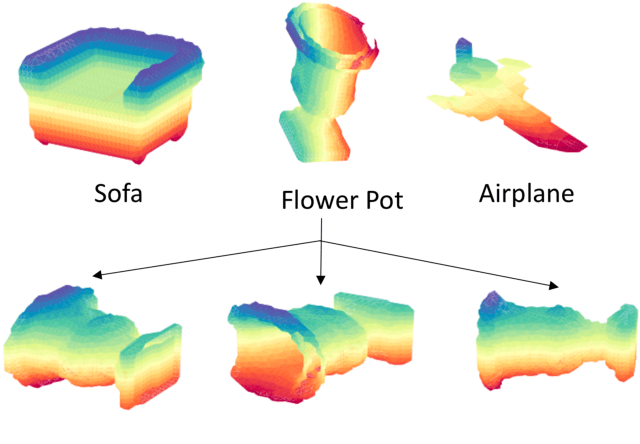
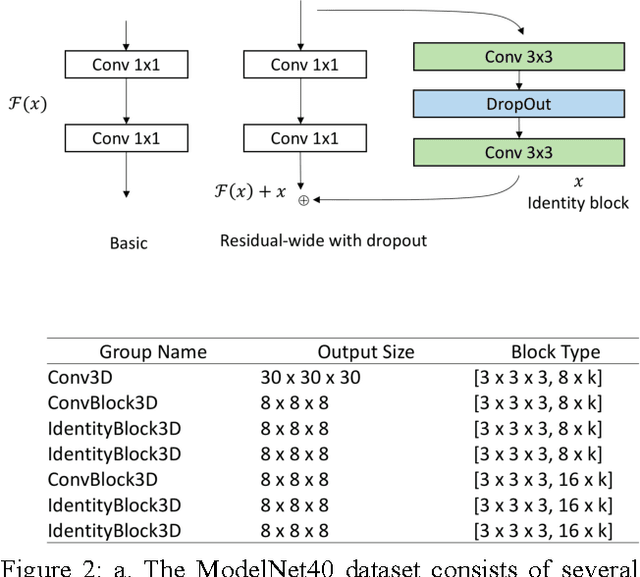
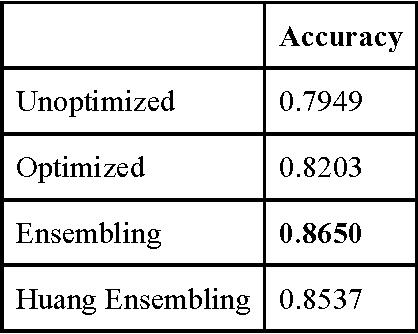
Abstract:3D shape models that directly classify objects from 3D information have become more widely implementable. Current state of the art models rely on deep convolutional and inception models that are resource intensive. Residual neural networks have been demonstrated to be easier to optimize and do not suffer from vanishing/exploding gradients observed in deep networks. Here we implement a residual neural network for 3D object classification of the 3D Princeton ModelNet dataset. Further, we show that widening network layers dramatically improves accuracy in shallow residual nets, and residual neural networks perform comparable to state-of-the-art 3D shape net models, and we show that widening network layers improves classification accuracy. We provide extensive training and architecture parameters providing a better understanding of available network architectures for use in 3D object classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge