Anita Bandrowski
Use as Directed? A Comparison of Software Tools Intended to Check Rigor and Transparency of Published Work
Jul 23, 2025



Abstract:The causes of the reproducibility crisis include lack of standardization and transparency in scientific reporting. Checklists such as ARRIVE and CONSORT seek to improve transparency, but they are not always followed by authors and peer review often fails to identify missing items. To address these issues, there are several automated tools that have been designed to check different rigor criteria. We have conducted a broad comparison of 11 automated tools across 9 different rigor criteria from the ScreenIT group. We found some criteria, including detecting open data, where the combination of tools showed a clear winner, a tool which performed much better than other tools. In other cases, including detection of inclusion and exclusion criteria, the combination of tools exceeded the performance of any one tool. We also identified key areas where tool developers should focus their effort to make their tool maximally useful. We conclude with a set of insights and recommendations for stakeholders in the development of rigor and transparency detection tools. The code and data for the study is available at https://github.com/PeterEckmann1/tool-comparison.
Antibody Watch: Text Mining Antibody Specificity from the Literature
Aug 05, 2020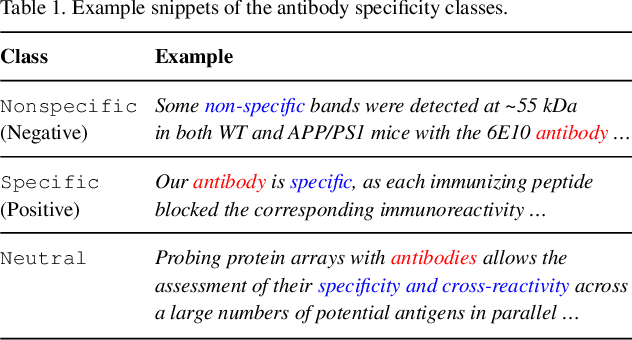
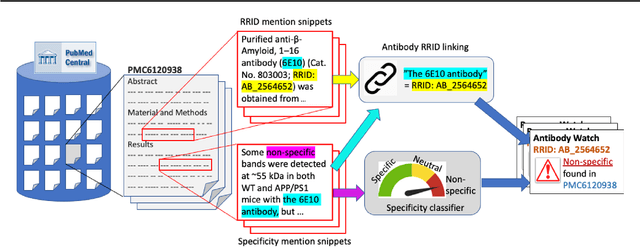
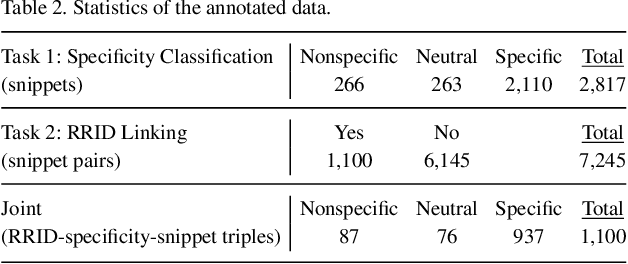
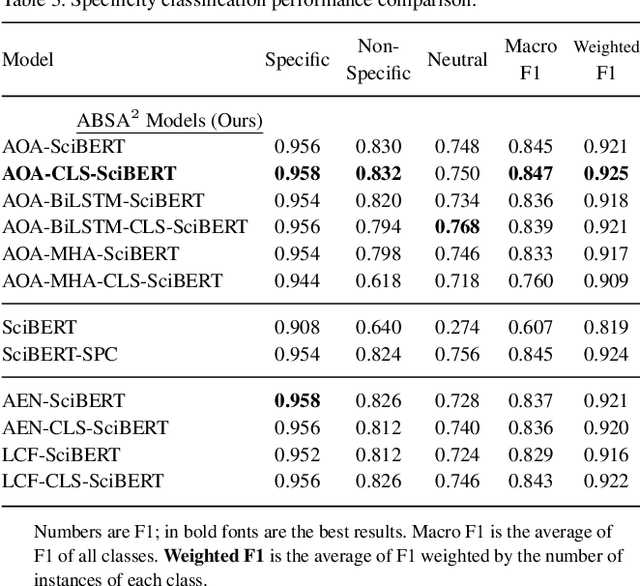
Abstract:Motivation: Antibodies are widely used reagents to test for expression of proteins. However, they might not always reliably produce results when they do not specifically bind to the target proteins that their providers designed them for, leading to unreliable research results. While many proposals have been developed to deal with the problem of antibody specificity, they may not scale well to deal with the millions of antibodies that are available to researchers. In this study, we investigate the feasibility of automatically generating a report to alert users of problematic antibodies by extracting statements about antibody specificity reported in the literature. Results: Our goal is to construct an "Antibody Watch" knowledge base containing supporting statements of problematic antibodies. We developed a deep neural network system and tested its performance with a corpus of more than two thousand articles that reported uses of antibodies. We divided the problem into two tasks. Given an input article, the first task is to identify snippets about antibody specificity and classify if the snippets report that any antibody exhibits nonspecificity, and thus is problematic. The second task is to link each of these snippets to one or more antibodies mentioned in the snippet. The experimental evaluation shows that our system can accurately perform both classification and linking tasks with weighted F-scores over 0.925 and 0.923, respectively, and 0.914 overall when combined to complete the joint task. We leveraged Research Resource Identifiers (RRID) to precisely identify antibodies linked to the extracted specificity snippets. The result shows that it is feasible to construct a reliable knowledge base about problematic antibodies by text mining.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge