Anirban Roychowdhury
Adaptive Bayesian Sampling with Monte Carlo EM
Nov 06, 2017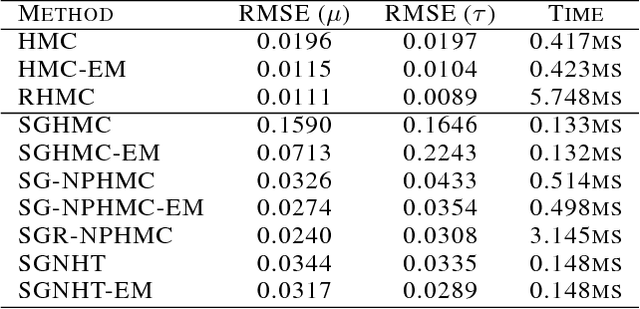
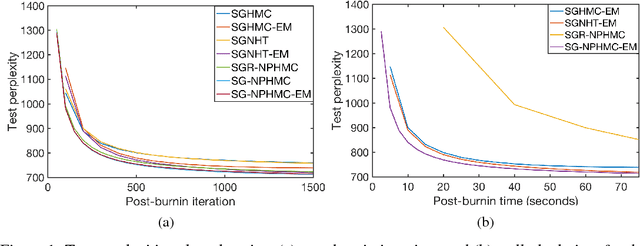
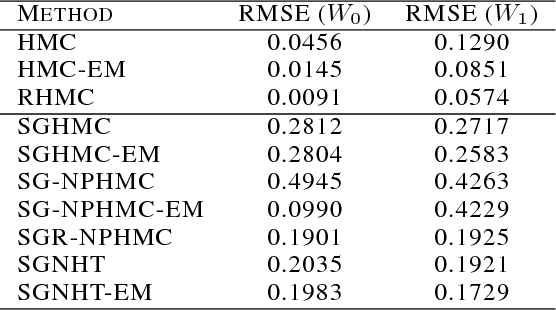
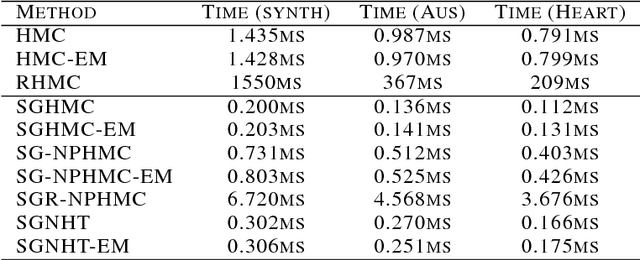
Abstract:We present a novel technique for learning the mass matrices in samplers obtained from discretized dynamics that preserve some energy function. Existing adaptive samplers use Riemannian preconditioning techniques, where the mass matrices are functions of the parameters being sampled. This leads to significant complexities in the energy reformulations and resultant dynamics, often leading to implicit systems of equations and requiring inversion of high-dimensional matrices in the leapfrog steps. Our approach provides a simpler alternative, by using existing dynamics in the sampling step of a Monte Carlo EM framework, and learning the mass matrices in the M step with a novel online technique. We also propose a way to adaptively set the number of samples gathered in the E step, using sampling error estimates from the leapfrog dynamics. Along with a novel stochastic sampler based on Nos\'{e}-Poincar\'{e} dynamics, we use this framework with standard Hamiltonian Monte Carlo (HMC) as well as newer stochastic algorithms such as SGHMC and SGNHT, and show strong performance on synthetic and real high-dimensional sampling scenarios; we achieve sampling accuracies comparable to Riemannian samplers while being significantly faster.
Accelerated Stochastic Quasi-Newton Optimization on Riemann Manifolds
May 22, 2017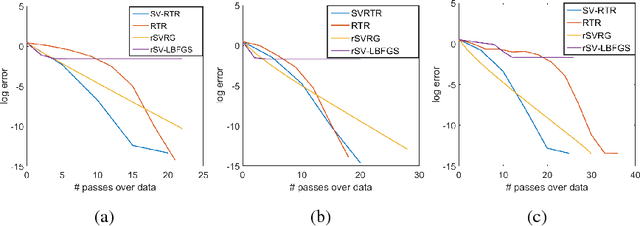
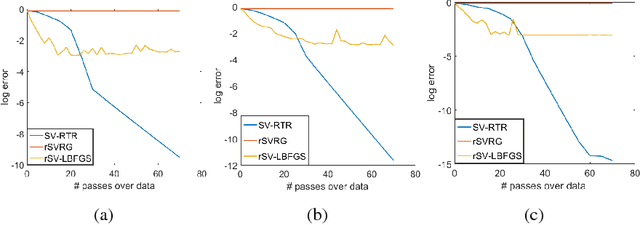
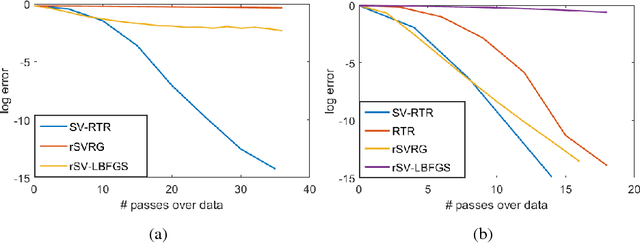
Abstract:We propose an L-BFGS optimization algorithm on Riemannian manifolds using minibatched stochastic variance reduction techniques for fast convergence with constant step sizes, without resorting to linesearch methods designed to satisfy Wolfe conditions. We provide a new convergence proof for strongly convex functions without using curvature conditions on the manifold, as well as a convergence discussion for nonconvex functions. We discuss a couple of ways to obtain the correction pairs used to calculate the product of the gradient with the inverse Hessian, and empirically demonstrate their use in synthetic experiments on computation of Karcher means for symmetric positive definite matrices and leading eigenvalues of large scale data matrices. We compare our method to VR-PCA for the latter experiment, along with Riemannian SVRG for both cases, and show strong convergence results for a range of datasets.
Gamma Processes, Stick-Breaking, and Variational Inference
Oct 04, 2014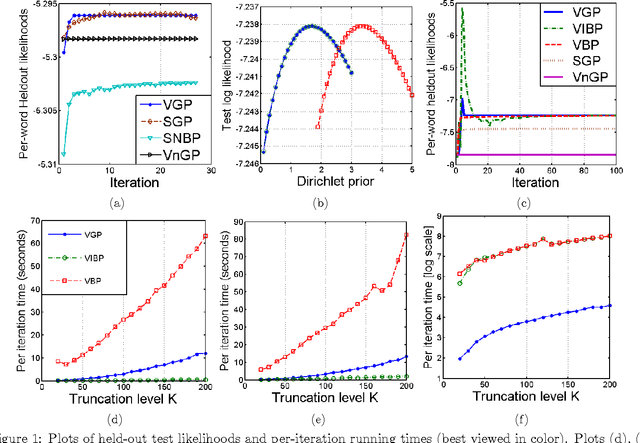
Abstract:While most Bayesian nonparametric models in machine learning have focused on the Dirichlet process, the beta process, or their variants, the gamma process has recently emerged as a useful nonparametric prior in its own right. Current inference schemes for models involving the gamma process are restricted to MCMC-based methods, which limits their scalability. In this paper, we present a variational inference framework for models involving gamma process priors. Our approach is based on a novel stick-breaking constructive definition of the gamma process. We prove correctness of this stick-breaking process by using the characterization of the gamma process as a completely random measure (CRM), and we explicitly derive the rate measure of our construction using Poisson process machinery. We also derive error bounds on the truncation of the infinite process required for variational inference, similar to the truncation analyses for other nonparametric models based on the Dirichlet and beta processes. Our representation is then used to derive a variational inference algorithm for a particular Bayesian nonparametric latent structure formulation known as the infinite Gamma-Poisson model, where the latent variables are drawn from a gamma process prior with Poisson likelihoods. Finally, we present results for our algorithms on nonnegative matrix factorization tasks on document corpora, and show that we compare favorably to both sampling-based techniques and variational approaches based on beta-Bernoulli priors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge