Andrew W Dowsey
Automated Re-Identification of Holstein-Friesian Cattle in Dense Crowds
Feb 17, 2026Abstract:Holstein-Friesian detection and re-identification (Re-ID) methods capture individuals well when targets are spatially separate. However, existing approaches, including YOLO-based species detection, break down when cows group closely together. This is particularly prevalent for species which have outline-breaking coat patterns. To boost both effectiveness and transferability in this setting, we propose a new detect-segment-identify pipeline that leverages the Open-Vocabulary Weight-free Localisation and the Segment Anything models as pre-processing stages alongside Re-ID networks. To evaluate our approach, we publish a collection of nine days CCTV data filmed on a working dairy farm. Our methodology overcomes detection breakdown in dense animal groupings, resulting in a 98.93% accuracy. This significantly outperforms current oriented bounding box-driven, as well as SAM species detection baselines with accuracy improvements of 47.52% and 27.13%, respectively. We show that unsupervised contrastive learning can build on this to yield 94.82% Re-ID accuracy on our test data. Our work demonstrates that Re-ID in crowded scenarios is both practical as well as reliable in working farm settings with no manual intervention. Code and dataset are provided for reproducibility.
MultiCamCows2024 -- A Multi-view Image Dataset for AI-driven Holstein-Friesian Cattle Re-Identification on a Working Farm
Oct 16, 2024



Abstract:We present MultiCamCows2024, a farm-scale image dataset filmed across multiple cameras for the biometric identification of individual Holstein-Friesian cattle exploiting their unique black and white coat-patterns. Captured by three ceiling-mounted visual sensors covering adjacent barn areas over seven days on a working dairy farm, the dataset comprises 101, 329 images of 90 cows, plus the underlying original CCTV footage. The dataset is provided alongside full computer vision recognition baselines, that is both a supervised and self-supervised learning framework for individual cow identification trained on cattle tracklets. We report a performance above 96% single image identification accuracy from the dataset and demonstrate that combining data from multiple cameras during learning enhances self-supervised identification. We show that our framework enables fully automatic cattle identification, barring only the simple human verification of tracklet integrity during data collection. Crucially, our study highlights that multi-camera, supervised and self-supervised components in tandem not only deliver highly accurate individual cow identification but also achieve this efficiently with no labelling of cattle identities by humans at all. We argue that this improvement in efficacy has practical implications for livestock management, behaviour analysis, and agricultural monitoring. For full reproducibility and practical ease of use, we publish all key software and code including re-identification components and the species detector with this paper.
Visual Identification of Individual Holstein-Friesian Cattle via Deep Metric Learning
Jul 04, 2020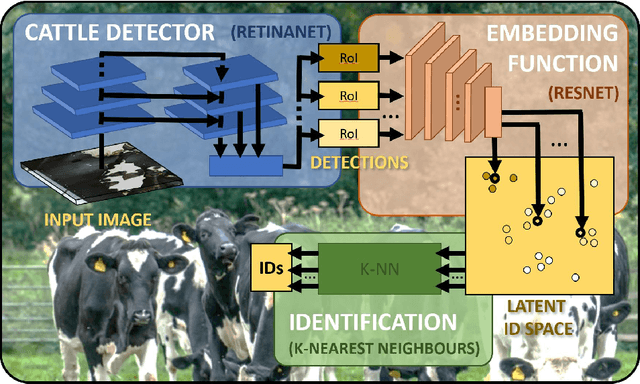
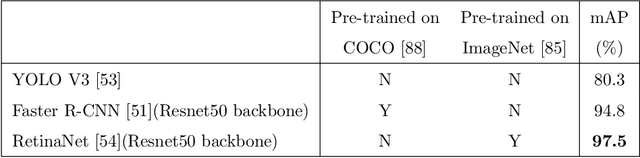
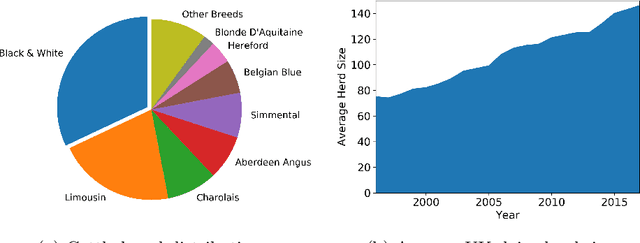
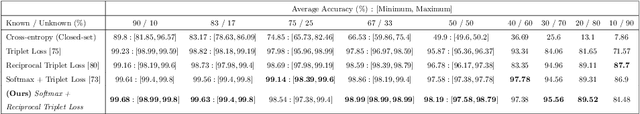
Abstract:Holstein-Friesian cattle exhibit individually-characteristic black and white coat patterns visually akin to those arising from Turing's reaction-diffusion systems. This work takes advantage of these natural markings in order to automate visual detection and biometric identification of individual Holstein-Friesians via convolutional neural networks and deep metric learning techniques. Existing approaches rely on markings, tags or wearables with a variety of maintenance requirements, whereas we present a totally hands-off method for the automated detection, localisation, and identification of individual animals from overhead imaging in an open herd setting, i.e. where new additions to the herd are identified without re-training. We propose the use of SoftMax-based reciprocal triplet loss to address the identification problem and evaluate the techniques in detail against fixed herd paradigms. We find that deep metric learning systems show strong performance even when many cattle unseen during system training are to be identified and re-identified - achieving 98.2% accuracy when trained on just half of the population. This work paves the way for facilitating the non-intrusive monitoring of cattle applicable to precision farming and surveillance for automated productivity, health and welfare monitoring, and to veterinary research such as behavioural analysis, disease outbreak tracing, and more. Key parts of the source code, network weights and underpinning datasets are available publicly.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge