Andreas Kolb
Hand Tracking based on Hierarchical Clustering of Range Data
Oct 25, 2011
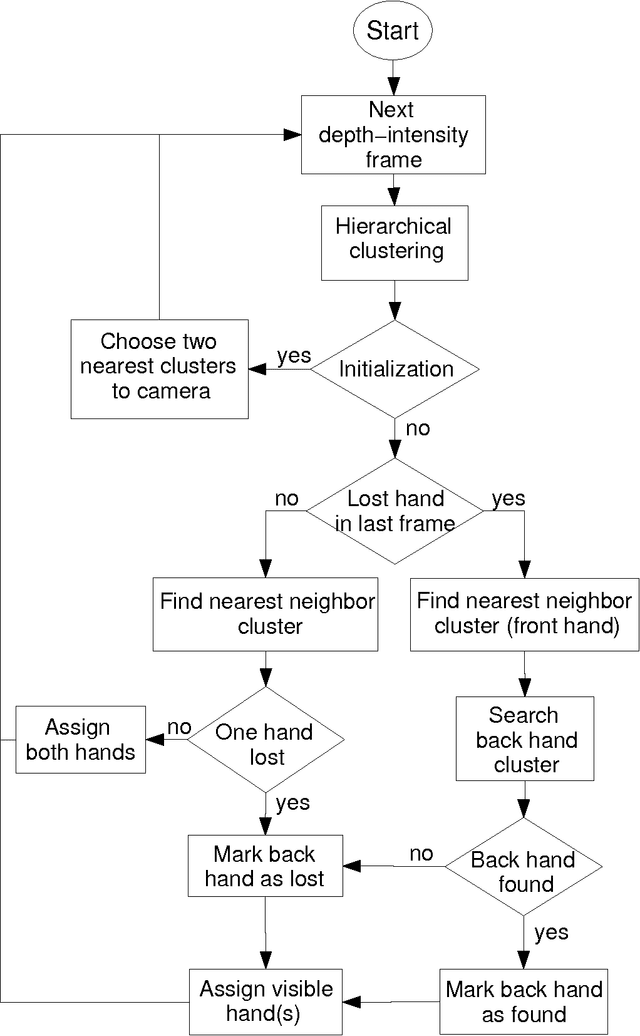
Abstract:Fast and robust hand segmentation and tracking is an essential basis for gesture recognition and thus an important component for contact-less human-computer interaction (HCI). Hand gesture recognition based on 2D video data has been intensively investigated. However, in practical scenarios purely intensity based approaches suffer from uncontrollable environmental conditions like cluttered background colors. In this paper we present a real-time hand segmentation and tracking algorithm using Time-of-Flight (ToF) range cameras and intensity data. The intensity and range information is fused into one pixel value, representing its combined intensity-depth homogeneity. The scene is hierarchically clustered using a GPU based parallel merging algorithm, allowing a robust identification of both hands even for inhomogeneous backgrounds. After the detection, both hands are tracked on the CPU. Our tracking algorithm can cope with the situation that one hand is temporarily covered by the other hand.
A Comparison of Two Human Brain Tumor Segmentation Methods for MRI Data
Mar 10, 2011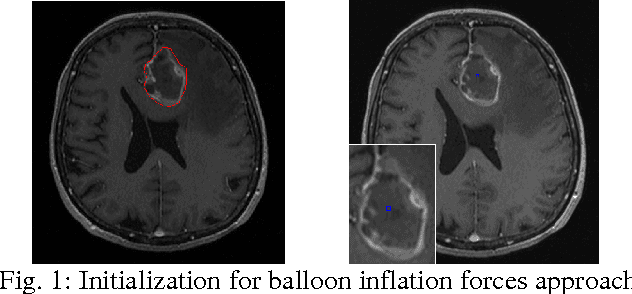
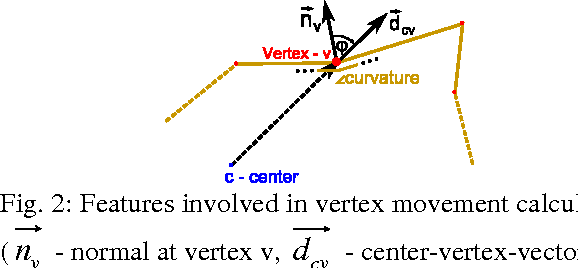
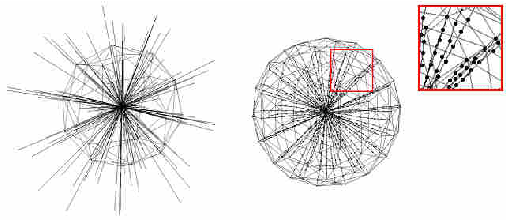
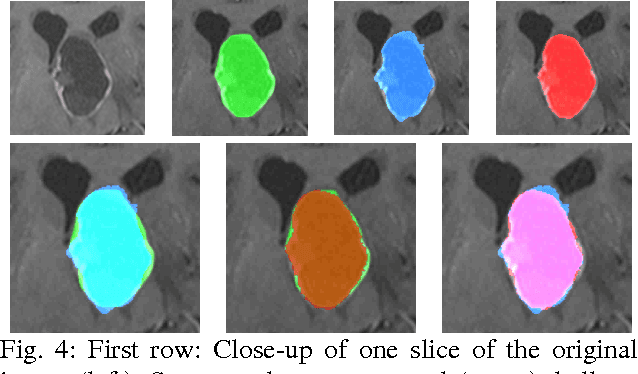
Abstract:The most common primary brain tumors are gliomas, evolving from the cerebral supportive cells. For clinical follow-up, the evaluation of the preoperative tumor volume is essential. Volumetric assessment of tumor volume with manual segmentation of its outlines is a time-consuming process that can be overcome with the help of computerized segmentation methods. In this contribution, two methods for World Health Organization (WHO) grade IV glioma segmentation in the human brain are compared using magnetic resonance imaging (MRI) patient data from the clinical routine. One method uses balloon inflation forces, and relies on detection of high intensity tumor boundaries that are coupled with the use of contrast agent gadolinium. The other method sets up a directed and weighted graph and performs a min-cut for optimal segmentation results. The ground truth of the tumor boundaries - for evaluating the methods on 27 cases - is manually extracted by neurosurgeons with several years of experience in the resection of gliomas. A comparison is performed using the Dice Similarity Coefficient (DSC), a measure for the spatial overlap of different segmentation results.
A Neural Network Classifier of Volume Datasets
Jun 12, 2009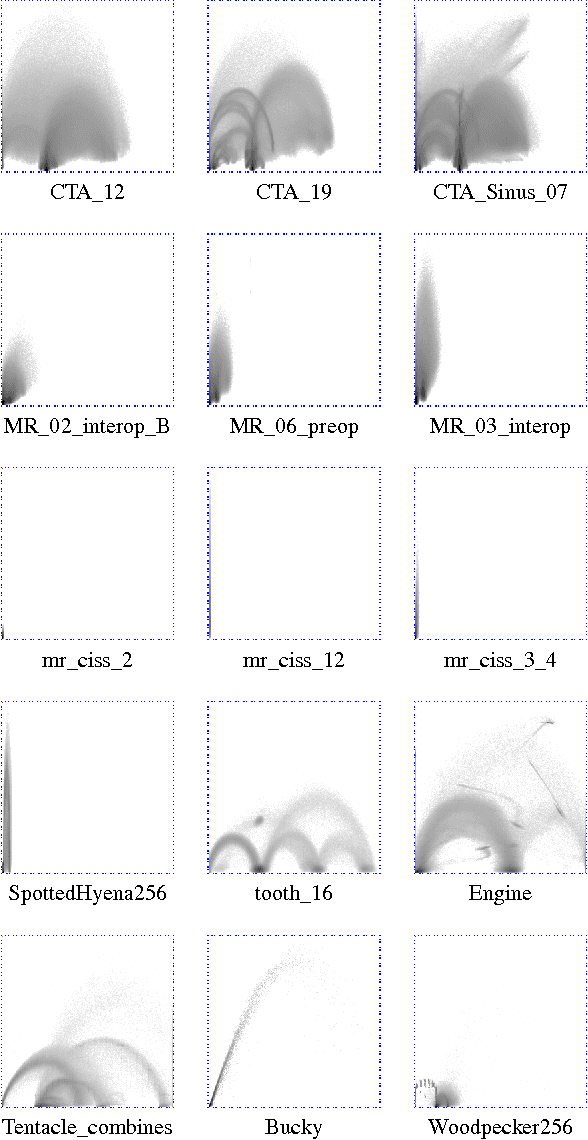
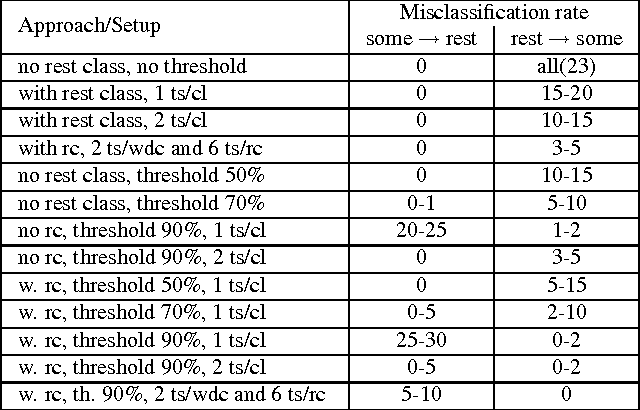
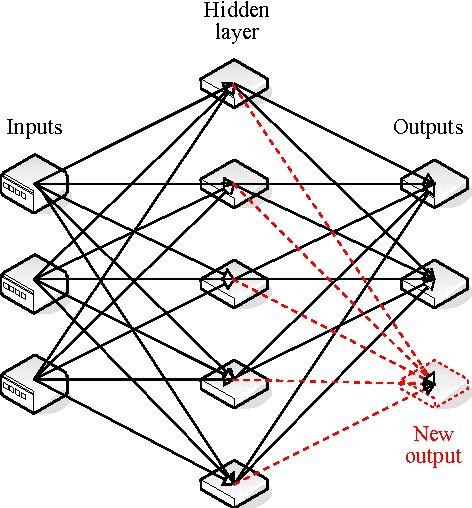
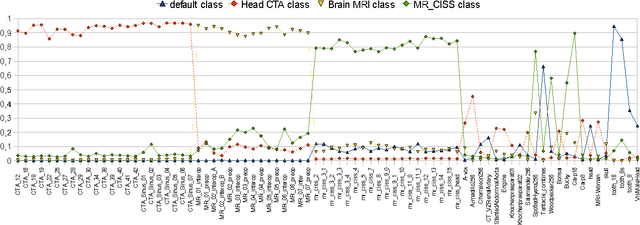
Abstract:Many state-of-the art visualization techniques must be tailored to the specific type of dataset, its modality (CT, MRI, etc.), the recorded object or anatomical region (head, spine, abdomen, etc.) and other parameters related to the data acquisition process. While parts of the information (imaging modality and acquisition sequence) may be obtained from the meta-data stored with the volume scan, there is important information which is not stored explicitly (anatomical region, tracing compound). Also, meta-data might be incomplete, inappropriate or simply missing. This paper presents a novel and simple method of determining the type of dataset from previously defined categories. 2D histograms based on intensity and gradient magnitude of datasets are used as input to a neural network, which classifies it into one of several categories it was trained with. The proposed method is an important building block for visualization systems to be used autonomously by non-experts. The method has been tested on 80 datasets, divided into 3 classes and a "rest" class. A significant result is the ability of the system to classify datasets into a specific class after being trained with only one dataset of that class. Other advantages of the method are its easy implementation and its high computational performance.
* 10 pages, 10 figures, 1 table, 3IA conference http://3ia.teiath.gr/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge