Alexander Hudson
Transfer Learning for Protein Structure Classification at Low Resolution
Aug 31, 2020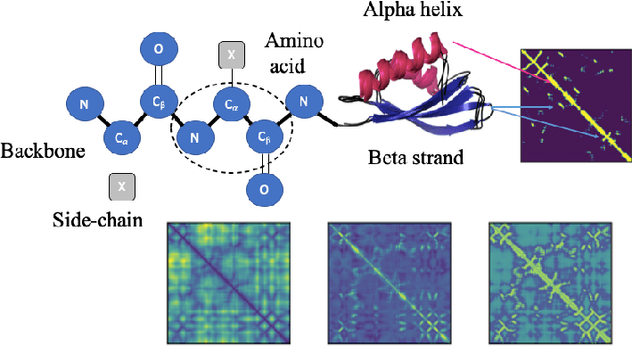
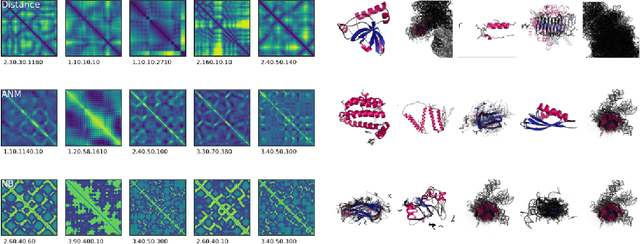
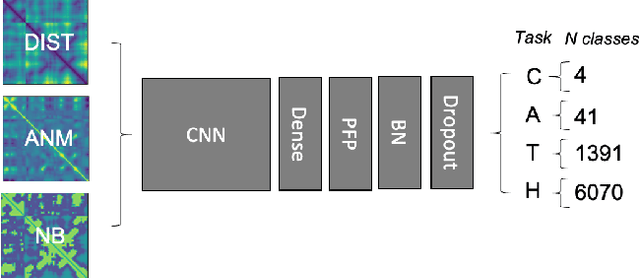
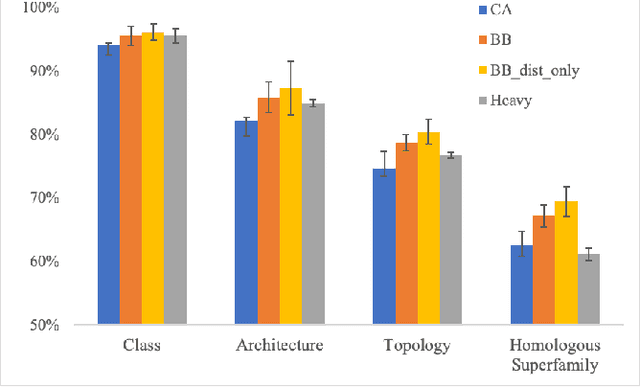

Abstract:Structure determination is key to understanding protein function at a molecular level. Whilst significant advances have been made in predicting structure and function from amino acid sequence, researchers must still rely on expensive, time-consuming analytical methods to visualise detailed protein conformation. In this study, we demonstrate that it is possible to make accurate ($\geq$80%) predictions of protein class and architecture from structures determined at low ($>$3A) resolution, using a deep convolutional neural network trained on high-resolution ($\leq$3A) structures represented as 2D matrices. Thus, we provide proof of concept for high-speed, low-cost protein structure classification at low resolution, and a basis for extension to prediction of function. We investigate the impact of the input representation on classification performance, showing that side-chain information may not be necessary for fine-grained structure predictions. Finally, we confirm that high-resolution, low-resolution and NMR-determined structures inhabit a common feature space, and thus provide a theoretical foundation for boosting with single-image super-resolution.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge