A. Webb
Benchmarking the performance of a low-cost Magnetic Resonance Control System at multiple sites in the open MaRCoS community
Mar 21, 2022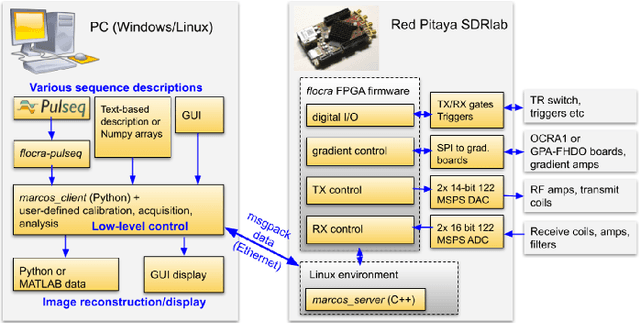
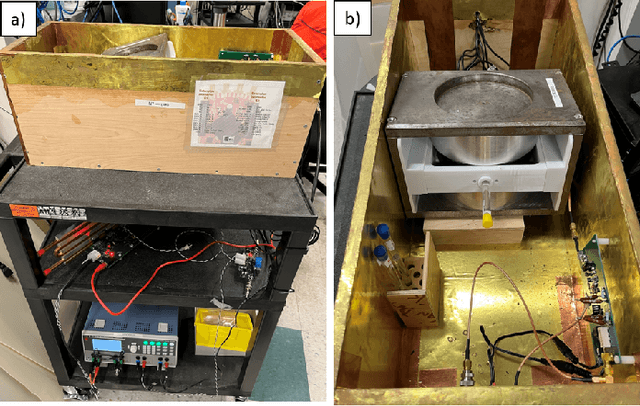
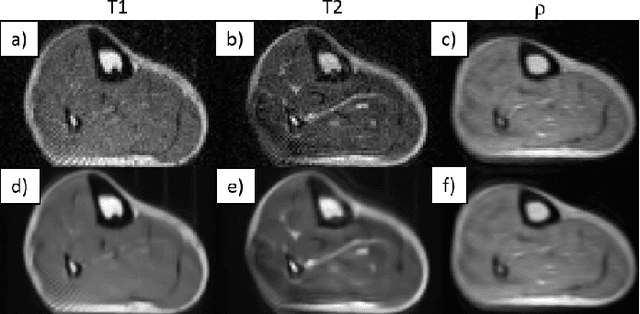

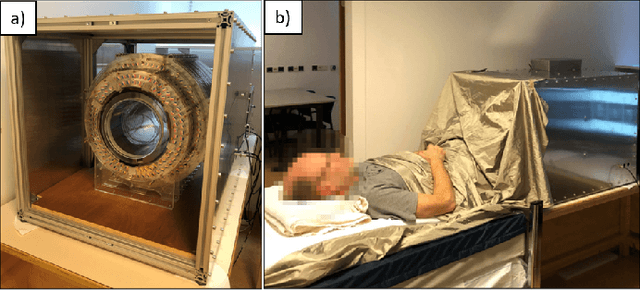
Abstract:Purpose: To describe the current properties and capabilities of an open-source hardware and software package that is being developed by many sites internationally with the aim of providing an inexpensive yet flexible platform for low-cost MRI. Methods: This paper describes three different setups from 50 to 360 mT in different settings, all of which used the MaRCoS console for acquiring data, and different types of software interfaces (custom-built GUI or PulSeq overlay) to acquire the data. Results: Images are presented from both phantoms and in vivo from healthy volunteers to demonstrate the image quality that can be obtained from the MaRCoS hardware/software interfaced to different low-field magnets. Conclusions: The results presented here show that a number of different sequences commonly used in the clinic can be programmed into an open-source system relatively quickly and easily, and can produce good quality images even at this early stage of development. Both the hardware and software will continue to develop, and it is an aim of this paper to encourage other groups to join this international consortium.
On Identification of Sparse Multivariable ARX Model: A Sparse Bayesian Learning Approach
Sep 30, 2016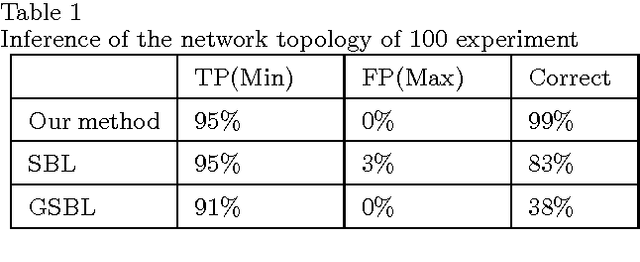
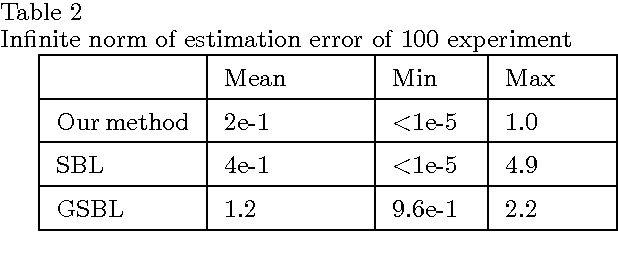
Abstract:This paper begins with considering the identification of sparse linear time-invariant networks described by multivariable ARX models. Such models possess relatively simple structure thus used as a benchmark to promote further research. With identifiability of the network guaranteed, this paper presents an identification method that infers both the Boolean structure of the network and the internal dynamics between nodes. Identification is performed directly from data without any prior knowledge of the system, including its order. The proposed method solves the identification problem using Maximum a posteriori estimation (MAP) but with inseparable penalties for complexity, both in terms of element (order of nonzero connections) and group sparsity (network topology). Such an approach is widely applied in Compressive Sensing (CS) and known as Sparse Bayesian Learning (SBL). We then propose a novel scheme that combines sparse Bayesian and group sparse Bayesian to efficiently solve the problem. The resulted algorithm has a similar form of the standard Sparse Group Lasso (SGL) while with known noise variance, it simplifies to exact re-weighted SGL. The method and the developed toolbox can be applied to infer networks from a wide range of fields, including systems biology applications such as signaling and genetic regulatory networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge